Abstract
Protein synthesis in eukaryotes is regulated by diverse reprogramming mechanisms that expand the coding capacity of individual genes. Here, we exploit one such mechanism, termed −1 programmed ribosomal frameshifting (−1 PRF), to engineer ligand-responsive RNA switches that regulate protein expression. First, efficient −1 PRF stimulatory RNA elements were discovered by in vitro selection; then, ligand-responsive switches were constructed by coupling −1 PRF stimulatory elements to RNA aptamers using rational design and directed evolution in Saccharomyces cerevisiae. We demonstrate that −1 PRF switches tightly control the relative stoichiometry of two distinct protein outputs from a single mRNA, exhibiting consistent ligand response across whole populations of cells. Furthermore, −1 PRF switches were applied to build single-mRNA logic gates and an apoptosis module in yeast. Together, these results showcase the potential for harnessing translation-reprogramming mechanisms for synthetic biology, and they establish −1 PRF switches as powerful RNA tools for controlling protein synthesis in eukaryotes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Zaher, H.S. & Green, R. Fidelity at the molecular level: lessons from protein synthesis. Cell 136, 746–762 (2009).
Gesteland, R.F. & Atkins, J.F. Recoding: dynamic reprogramming of translation. Annu. Rev. Biochem. 65, 741–768 (1996).
Firth, A.E. & Brierley, I. Non-canonical translation in RNA viruses. J. Gen. Virol. 93, 1385–1409 (2012).
Atkins, J.F. & Gesteland, R.F. (eds.). Recoding: Expansion of Decoding Rules Enriches Gene Expression (Springer, New York, 2010).
Mountford, P.S. & Smith, A.G. Internal ribosome entry sites and dicistronic RNAs in mammalian transgenesis. Trends Genet. 11, 179–184 (1995).
de Felipe, P., Hughes, L.E., Ryan, M.D. & Brown, J.D. Co-translational, intraribosomal cleavage of polypeptides by the foot-and-mouth disease virus 2A peptide. J. Biol. Chem. 278, 11441–11448 (2003).
Liu, C.C. & Schultz, P.G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Isaacs, F.J., Dwyer, D.J. & Collins, J.J. RNA synthetic biology. Nat. Biotechnol. 24, 545–554 (2006).
Ellington, A.D. & Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Robertson, D.L. & Joyce, G.F. Selection in vitro of an RNA enzyme that specifically cleaves single-stranded DNA. Nature 344, 467–468 (1990).
Jenison, R.D., Gill, S.C., Pardi, A. & Polisky, B. High-resolution molecular discrimination by RNA. Science 263, 1425–1429 (1994).
Berens, C., Thain, A. & Schroeder, R. A tetracycline-binding RNA aptamer. Bioorg. Med. Chem. 9, 2549–2556 (2001).
Tang, J. & Breaker, R.R. Rational design of allosteric ribozymes. Chem. Biol. 4, 453–459 (1997).
Klauser, B., Atanasov, J., Siewert, L.K. & Hartig, J.S. Ribozyme-based aminoglycoside switches of gene expression engineered by genetic selection in S. cerevisiae. ACS Synth. Biol. 4, 516–525 (2015).
Townshend, B., Kennedy, A.B., Xiang, J.S. & Smolke, C.D. High-throughput cellular RNA device engineering. Nat. Methods 12, 989–994 (2015).
Koizumi, M., Soukup, G.A., Kerr, J.N. & Breaker, R.R. Allosteric selection of ribozymes that respond to the second messengers cGMP and cAMP. Nat. Struct. Biol. 6, 1062–1071 (1999).
Link, K.H. et al. Engineering high-speed allosteric hammerhead ribozymes. Biol. Chem. 388, 779–786 (2007).
Win, M.N. & Smolke, C.D. Higher-order cellular information processing with synthetic RNA devices. Science 322, 456–460 (2008).
Ausländer, S. et al. A general design strategy for protein-responsive riboswitches in mammalian cells. Nat. Methods 11, 1154–1160 (2014).
Chen, Y.Y., Jensen, M.C. & Smolke, C.D. Genetic control of mammalian T-cell proliferation with synthetic RNA regulatory systems. Proc. Natl. Acad. Sci. USA 107, 8531–8536 (2010).
Galloway, K.E., Franco, E. & Smolke, C.D. Dynamically reshaping signaling networks to program cell fate via genetic controllers. Science 341, 1235005 (2013).
Brierley, I. Ribosomal frameshifting on viral RNAs. J. Gen. Virol. 76, 1885–1892 (1995).
Jacks, T., Madhani, H.D., Masiarz, F.R. & Varmus, H.E. Signals for ribosomal frameshifting in the Rous sarcoma virus gag-pol region. Cell 55, 447–458 (1988).
Ivanov, I.P., Gesteland, R.F. & Atkins, J.F. Antizyme expression: a subversion of triplet decoding, which is remarkably conserved by evolution, is a sensor for an autoregulatory circuit. Nucleic Acids Res. 28, 3185–3196 (2000).
Yu, C.H., Luo, J., Iwata-Reuyl, D. & Olsthoorn, R.C.L. Exploiting preQ1 riboswitches to regulate ribosomal frameshifting. ACS Chem. Biol. 8, 733–740 (2013).
Hsu, H.T., Lin, Y.H. & Chang, K.Y. Synergetic regulation of translational reading-frame switch by ligand-responsive RNAs in mammalian cells. Nucleic Acids Res. 42, 14070–14082 (2014).
Roberts, R.W. & Szostak, J.W. RNA-peptide fusions for the in vitro selection of peptides and proteins. Proc. Natl. Acad. Sci. USA 94, 12297–12302 (1997).
Frankel, A. & Roberts, R.W. In vitro selection for sense codon suppression. RNA 9, 780–786 (2003).
Liu, R., Barrick, J.E., Szostak, J.W. & Roberts, R.W. Optimized synthesis of RNA-protein fusions for in vitro protein selection. Methods Enzymol. 318, 268–293 (2000).
Cho, G., Keefe, A.D., Liu, R., Wilson, D.S. & Szostak, J.W. Constructing high complexity synthetic libraries of long ORFs using in vitro selection. J. Mol. Biol. 297, 309–319 (2000).
Weigand, J.E. et al. Screening for engineered neomycin riboswitches that control translation initiation. RNA 14, 89–97 (2008).
Fields, S. & Song, O. A novel genetic system to detect protein-protein interactions. Nature 340, 245–246 (1989).
Vidal, M., Brachmann, R.K., Fattaey, A., Harlow, E. & Boeke, J.D. Reverse two-hybrid and one-hybrid systems to detect dissociation of protein-protein and DNA-protein interactions. Proc. Natl. Acad. Sci. USA 93, 10315–10320 (1996).
Zadeh, J.N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput. Chem. 32, 170–173 (2011).
Bonnet, J., Yin, P., Ortiz, M.E., Subsoontorn, P. & Endy, D. Amplifying genetic logic gates. Science 340, 599–603 (2013).
Moon, T.S., Lou, C., Tamsir, A., Stanton, B.C. & Voigt, C.A. Genetic programs constructed from layered logic gates in single cells. Nature 491, 249–253 (2012).
Youle, R.J. & Strasser, A. The BCL-2 protein family: opposing activities that mediate cell death. Nat. Rev. Mol. Cell Biol. 9, 47–59 (2008).
Sato, T. et al. Interactions among members of the Bcl-2 protein family analyzed with a yeast two-hybrid system. Proc. Natl. Acad. Sci. USA 91, 9238–9242 (1994).
Priault, M., Camougrand, N., Kinnally, K.W., Vallette, F.M. & Manon, S. Yeast as a tool to study Bax/mitochondrial interactions in cell death. FEMS Yeast Res. 4, 15–27 (2003).
Gallenne, T. et al. Bax activation by the BH3-only protein Puma promotes cell dependence on antiapoptotic Bcl-2 family members. J. Cell Biol. 185, 279–290 (2009).
Czabotar, P.E., Lessene, G., Strasser, A. & Adams, J.M. Control of apoptosis by the BCL-2 protein family: implications for physiology and therapy. Nat. Rev. Mol. Cell Biol. 15, 49–63 (2014).
Lee, R.E.C., Walker, S.R., Savery, K., Frank, D.A. & Gaudet, S. Fold change of nuclear NF-κB determines TNF-induced transcription in single cells. Mol. Cell 53, 867–879 (2014).
Kang, M., Peterson, R. & Feigon, J. Structural insights into riboswitch control of the biosynthesis of queuosine, a modified nucleotide found in the anticodon of tRNA. Mol. Cell 33, 784–790 (2009).
Seelig, B. mRNA display for the selection and evolution of enzymes from in vitro-translated protein libraries. Nat. Protoc. 6, 540–552 (2011).
Keppler-Ross, S., Noffz, C. & Dean, N. A new purple fluorescent color marker for genetic studies in Saccharomyces cerevisiae and candida albicans. Genetics 179, 705–710 (2008).
Pirakitikulr, N., Ostrov, N., Peralta-Yahya, P. & Cornish, V.W. PCRless library mutagenesis via oligonucleotide recombination in yeast. Protein Sci. 19, 2336–2346 (2010).
Gross, A. et al. Biochemical and genetic analysis of the mitochondrial response of yeast to BAX and BCL-X(L). Mol. Cell. Biol. 20, 3125–3136 (2000).
Acknowledgements
We thank N. Dean (Stony Brook University, USA) for providing the yEmRFP plasmid encoding yeast-optimized mCherry. We thank R.L. Gonzalez for valuable discussions and feedback and N. Ostrov for critical comments on the manuscript. This work was supported by funds from the US National Institutes of Health (NIH) (grants R01 GM090126 and R01 AI110794 to V.W.C.). S.Z. and R.R. were supported by the Center for Topology of Cancer Evolution and Heterogeneity (NIH U54 CA193313). A.V.A. was supported by a US National Institutes of Health F30 fellowship (F30 CA174357). S.Z. was supported by a TL1 Precision Medicine fellowship (5TL1 TR000082). Research reported in this publication was performed in the CCTI Flow Cytometry Core, supported in part by US National Institutes of Health award S10RR027050.
Author information
Authors and Affiliations
Contributions
A.V.A. conceived the project, designed and performed experiments, analyzed data, and wrote the manuscript. A.J.L. performed experiments and analyzed data. S.Z. and R.R. developed custom software to analyze the NGS data. V.W.C. supervised the research and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 In vitro frameshift selection construct
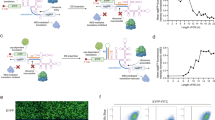
(a) The library was constructed based on a previously characterized prokaryotic PreQ1 riboswitch scaffold. Nucleotides surrounding the ligand binding pocket were randomized, generating a library of ~2.68×108 variants. (b) Annotated sequence of the selection construct in DNA form. (c) The translation product resulting from maintenance of the original reading frame (upper) or -1 frameshift at the heptanucleotide slippery site (lower).
5. Yu, C.-H., Luo, J., Iwata-Reuyl, D. & Olsthoorn, R. C. L. Exploiting preQ1 Riboswitches To Regulate Ribosomal Frameshifting. ACS Chem. Biol. 8, 733–740 (2013).
Supplementary Figure 2 Screening and characterization of in vitro selection products in the dual-FP reporter assay.
(a) Sample of randomly selected individual clones quantified for frameshift activity (not representative of the full library distribution). (b) Dot plot and (c) histogram of bulk in vitro selection products after three rounds of selection cloned into the dual-FP reporter in yeast and analyzed by flow cytometry.
Supplementary Figure 3 Pseudoknot feature space construction.
H-type pseudoknots can be defined by the lengths of their segments, specifically their loops (L1, L2, and L3) and stems (S1 and S2). To fix the position of the pseudoknot within the library scaffold, an upstream (Up) and downstream (Dn) segment is also defined. The sum of these strand lengths (stem segments are double stranded) must equal the scaffold’s total length of 44 nucleotides. All combinations of stand lengths that obey the above constraints were generated, and then assessed for base pairing compatibility using the scaffold sequence (where N can take on any nucleotide identity). The resulting PK features that satisfy these requirements define the set of all possible PKs. For this set of constraints and scaffold there are 2,068 PK features.
Supplementary Figure 4 PK feature enrichment.
The PK feature probability was calculated from the theoretical starting library (assuming equal distribution of sequences) by dividing the number of sequences within a PK feature by the starting library size (approx. 2.68×108). Post selection probability was computed by dividing the number of sequencing reads contained within the PK feature by the total sequencing reads (approx. 6×107).
Supplementary Figure 5 NGS analysis and the FS-2 sequence.
The analysis pipeline outputs various statistics for a motif, as well as a sequence logo (underlined segments represent regions of randomized nucleotides in the starting library). Demonstrated above for FS-2, individual sequences can be analyzed for single or pairwise nucleotide variants to identify mutation tolerant or intolerant positions, and reveal potential mutual information.
Supplementary Figure 6 Design and characterization of −1 PRF OFF switches.
(a) General scheme depicting the intended structural rearrangements. A representative theophylline responsive OFF switch sequence is shown. (b) Optimization of a theophylline responsive OFF-switch. Increasing the length of the theophylline aptamer stem leads to increasing disruption of the stimulatory pseudoknot. Frameshift levels are inversely correlated with aptamer thermodynamic stability, both in the presence and absence of ligand. (c) Efforts to optimize a neomycin responsive OFF-switch. As with the theophylline responsive switches, in general, frameshift levels are inversely correlated to the thermodynamic stability of the aptamer. However, this relationship is not as direct, and this method was unable to provide a switch that possessed ideal -1 PRF levels and ligand responsiveness. Therefore, the neomycin OFF-switch was optimized by in vivo directed evolution (see Supplementary Fig. 9).
Supplementary Figure 7 Directed evolution of −1 PRF switches in S. cerevisiae.
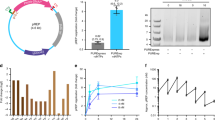
(a) -1 PRF switches are encoded between a Gal4 minimal DNA binding domain (Gal4-BD) and a Gal4 minimal activation domain (Gal4-AD). Failure to frameshift at the switch terminates translation and releases a Gal4-BD, which can bind to Gal4-UAS but does not activate transcription. -1 frameshifting produces the full minimal Gal4 transcription factor. Activation of Gal4 responsive genes is a function of the Gal4-BD/Gal4-TF ratio, which is controlled by -1 PRF efficiency (see Supplementary Figure 8). (b). Selection scheme. A library of -1 PRF devices is cloned into the Gal4 selection vector by gap repair and tested in the yeast 2-hybrid selection strain MAV203 (MATα; leu2-3,112; trp1-901; his3Δ200; ade2-101; cyh2R; can1R; gal4Δ; gal80 D; GAL1::lacZ; HIS3UASGAL1::HIS3@LYS2; SPAL10::URA3). The transformants are subjected to positive selection in media lacking histidine supplemented with the His3 inhibitor 3-aminotriazole (3-AT). If selecting for ON switches, media is also supplemented with the small molecule inducer. Counterselection is performed in minimal media containing 5-fluoroorotic acid (5-FOA) and the small molecule ligand for OFF switch selections. After selection, DNA can be extracted from the culture and cloned into the dual-FP reporter for characterization, or mutagenized and retransformed for further rounds of selection.
Supplementary Figure 8 Growth assays of −1 PRF devices in the Gal4 selection construct.
-1 PRF stimulators of varying efficiency were cloned into the Gal4 selection plasmid and assayed for growth in (a) positive selection conditions in minimal media lacking histidine and containing varying amounts of 3-AT or (b) counterselection conditions in minimal media containing varying concentrations of 5-FOA. Lower efficiency frameshift stimulators are more sensitive to 3-AT in the positive selection and are more easily growth inhibited. Higher efficiency frameshift stimulators are more sensitive to 5-FOA in the counterselection and are more easily growth inhibited. (c) A theophylline responsive ON-switch in the Gal4 selection. The construct shows theophylline-assisted growth in the positive selection conditions and growth inhibition by theophylline in the counterselection conditions.
Supplementary Figure 9 Optimization of a neomycin responsive OFF-switch by in vivo directed evolution.
(a) Scheme of the neomycin OFF switch library targeting the aptamer stem. Two libraries, each containing 5 randomized nucleotides in the aptamer, were, cloned into the Gal4 selection vector and transformed into the yeast two-hybrid selection strain (see Supplementary Fig. 7). (b) After one round of counterselection in the presence of neomycin and one round of positive selection in the absence of neomycin, selection products were cloned into the dual-FP reporter. Randomly sampled colonies were assayed for ligand responsiveness. One variant (aptamer structure depicted on the right) demonstrated significantly improved switch behavior.
Supplementary Figure 10 Design and characterization of theophylline responsive ON switches.
(a) General scheme depicting the intended structural rearrangements induced by ligand binding. A representative theophylline responsive ON switch sequence is shown. (b) First, basal frameshifting levels were tuned down by inserting a switching hairpin to compete with FS-2 pseudoknot folding. Basal frameshifting and ligand induced frameshifting is inversely correlated with switching hairpin length and thermodynamic stability. (c) Activation was optimized by adjusting the length of the aptamer stem. Frameshifting activity, in general, correlated with the stem length and thermodynamic stability of the aptamer. Incorporation of other thermodynamic parameters partially accounts for the discrepancy in this trend (see Supplementary Table 1).
Supplementary Figure 11 Design and characterization of neomycin responsive ON switches.
(a) General scheme depicting the intended structural rearrangements induced by ligand binding. A representative neomycin responsive ON switch sequence is shown. (b) Frameshift levels were tuned by adjusting the length and loop composition of the switching hairpin. In general, basal frameshifting and ligand induced frameshifting is inversely correlated with switching hairpin thermodynamic stability (Supplementary Table 1). By comparison to the theophylline ON switches, neomycin ON switches, the hairpin stability was tuned by modifying the tetraloop nucleotides.
Supplementary Figure 12 Hypothetical logic gates composed of layered −1 PRF switches.
(a) AND gate, (b) NOR gate, (c) OR gate, (d) NAND gate, (e) XOR gate, (f) XNOR gate.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1–6, and Supplementary Notes 1 and 2 (PDF 6935 kb)
Supplementary Data 1
-1 PRF motif table describing various enriched pseudoknot structures from the in vitro mRNA display selection (XLS 55 kb)
Supplementary Data 2
Thermodynamic calculations for designing -1 PRF ON switches (XLSX 44 kb)
Rights and permissions
About this article
Cite this article
Anzalone, A., Lin, A., Zairis, S. et al. Reprogramming eukaryotic translation with ligand-responsive synthetic RNA switches. Nat Methods 13, 453–458 (2016). https://doi.org/10.1038/nmeth.3807
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3807
This article is cited by
-
Engineered pegRNAs improve prime editing efficiency
Nature Biotechnology (2022)
-
Engineering synthetic RNA devices for cell control
Nature Reviews Genetics (2022)
-
Translational control of enzyme scavenger expression with toxin-induced micro RNA switches
Scientific Reports (2021)
-
The T-cell leukemia-associated ribosomal RPL10 R98S mutation enhances JAK-STAT signaling
Leukemia (2018)