Abstract
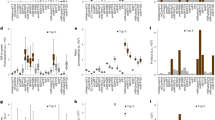
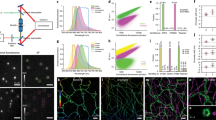
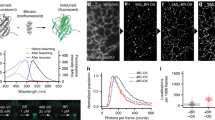
Photoswitchable fluorescent probes are central to localization-based super-resolution microscopy. Among these probes, fluorescent proteins are appealing because they are genetically encoded. Moreover, the ability to achieve a 1:1 labeling ratio between the fluorescent protein and the protein of interest makes these probes attractive for quantitative single-molecule counting. The percentage of fluorescent protein that is photoactivated into a fluorescently detectable form (i.e., the photoactivation efficiency) plays a crucial part in properly interpreting the quantitative information. It is important to characterize the photoactivation efficiency at the single-molecule level under the conditions used in super-resolution imaging. Here, we used the human glycine receptor expressed in Xenopus oocytes and stepwise photobleaching or single-molecule counting photoactivated localization microcopy (PALM) to determine the photoactivation efficiency of fluorescent proteins mEos2, mEos3.1, mEos3.2, Dendra2, mClavGR2, mMaple, PA-GFP and PA-mCherry. This analysis provides important information that must be considered when using these fluorescent proteins in quantitative super-resolution microscopy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
13 January 2014
In the version of this article initially published online, the institution name "ICFO" was incorrectly written as "IFCO" in the author affiliations. The error has been corrected for the print, PDF and HTML versions of this article.
References
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S.T., Girirajan, T.P. & Mason, M.D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Rust, M.J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–796 (2006).
Huang, B., Babcock, H. & Zhuang, X. Breaking the diffraction barrier: super-resolution imaging of cells. Cell 143, 1047–1058 (2010).
Huang, B., Jones, S.A., Brandenburg, B. & Zhuang, X. Whole-cell 3D STORM reveals interactions between cellular structures with nanometer-scale resolution. Nat. Methods 5, 1047–1052 (2008).
Lakadamyali, M., Babcock, H., Bates, M., Zhuang, X. & Lichtman, J. 3D multicolor super-resolution imaging offers improved accuracy in neuron tracing. PLoS ONE 7, e30826 (2012).
Balint, S., Verdeny Vilanova, I., Sandoval Alvarez, A. & Lakadamyali, M. Correlative live-cell and superresolution microscopy reveals cargo transport dynamics at microtubule intersections. Proc. Natl. Acad. Sci. USA 110, 3375–3380 (2013).
Kanchanawong, P. et al. Nanoscale architecture of integrin-based cell adhesions. Nature 468, 580–584 (2010).
Shroff, H. et al. Dual-color superresolution imaging of genetically expressed probes within individual adhesion complexes. Proc. Natl. Acad. Sci. USA 104, 20308–20313 (2007).
Xu, K., Zhong, G. & Zhuang, X. Actin, spectrin, and associated proteins form a periodic cytoskeletal structure in axons. Science 339, 452–456 (2013).
Gunzenhäuser, J., Olivier, N., Pengo, T. & Manley, S. Quantitative super-resolution imaging reveals protein stoichiometry and nanoscale morphology of assembling HIV-Gag virions. Nano Lett. 12, 4705–4710 (2012).
Lando, D. et al. Quantitative single-molecule microscopy reveals that CENP-A(Cnp1) deposition occurs during G2 in fission yeast. Open Biol. 2, 120078 (2012).
Lee, S.H., Shin, J.Y., Lee, A. & Bustamante, C. Counting single photoactivatable fluorescent molecules by photoactivated localization microscopy (PALM). Proc. Natl. Acad. Sci. USA 109, 17436–17441 (2012).
Renz, M., Daniels, B.R., Vamosi, G., Arias, I.M. & Lippincott-Schwartz, J. Plasticity of the asialoglycoprotein receptor deciphered by ensemble FRET imaging and single-molecule counting PALM imaging. Proc. Natl. Acad. Sci. USA 109, E2989–E2997 (2012).
Sengupta, P. et al. Probing protein heterogeneity in the plasma membrane using PALM and pair correlation analysis. Nat. Methods 8, 969–975 (2011).
Fernández-Suárez, M. & Ting, A.Y. Fluorescent probes for super-resolution imaging in living cells. Nat. Rev. Mol. Cell Biol. 9, 929–943 (2008).
Lippincott-Schwartz, J. & Patterson, G.H. Photoactivatable fluorescent proteins for diffraction-limited and super-resolution imaging. Trends Cell Biol. 19, 555–565 (2009).
Brakemann, T. et al. A reversibly photoswitchable GFP-like protein with fluorescence excitation decoupled from switching. Nat. Biotechnol. 29, 942–947 (2011).
Grotjohann, T. et al. Diffraction-unlimited all-optical imaging and writing with a photochromic GFP. Nature 478, 204–208 (2011).
Habuchi, S. et al. Reversible single-molecule photoswitching in the GFP-like fluorescent protein Dronpa. Proc. Natl. Acad. Sci. USA 102, 9511–9516 (2005).
Patterson, G.H. & Lippincott-Schwartz, J. A photoactivatable GFP for selective photolabeling of proteins and cells. Science 297, 1873–1877 (2002).
Subach, F.V. et al. Photoactivatable mCherry for high-resolution two-color fluorescence microscopy. Nat. Methods 6, 153–159 (2009).
McKinney, S.A., Murphy, C.S., Hazelwood, K.L., Davidson, M.W. & Looger, L.L. A bright and photostable photoconvertible fluorescent protein. Nat. Methods 6, 131–133 (2009).
Gurskaya, N.G. et al. Engineering of a monomeric green-to-red photoactivatable fluorescent protein induced by blue light. Nat. Biotechnol. 24, 461–465 (2006).
Zhang, M. et al. Rational design of true monomeric and bright photoactivatable fluorescent proteins. Nat. Methods 9, 727–729 (2012).
Hoi, H. et al. A monomeric photoconvertible fluorescent protein for imaging of dynamic protein localization. J. Mol. Biol. 401, 776–791 (2010).
McEvoy, A.L. et al. mMaple: a photoconvertible fluorescent protein for use in multiple imaging modalities. PLoS ONE 7, e51314 (2012).
Dempsey, G.T., Vaughan, J.C., Chen, K.H., Bates, M. & Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 8, 1027–1036 (2011).
Annibale, P., Scarselli, M., Kodiyan, A. & Radenovic, A. Photoactivatable fluorescent protein mEos2 displays repeated photoactivation after a long-lived dark state in the red photoconverted form. J. Phys. Chem. Lett. 1, 1506–1510 (2010).
Annibale, P., Vanni, S., Scarselli, M., Rothlisberger, U. & Radenovic, A. Quantitative photo activated localization microscopy: unraveling the effects of photoblinking. PLoS ONE 6, e22678 (2011).
Sengupta, P., Jovanovic-Talisman, T. & Lippincott-Schwartz, J. Quantifying spatial organization in point-localization superresolution images using pair correlation analysis. Nat. Protoc. 8, 345–354 (2013).
Adam, V., Nienhaus, K., Bourgeois, D. & Nienhaus, G.U. Structural basis of enhanced photoconversion yield in green fluorescent protein-like protein Dendra2. Biochemistry 48, 4905–4915 (2009).
Wiedenmann, J. et al. EosFP, a fluorescent marker protein with UV-inducible green-to-red fluorescence conversion. Proc. Natl. Acad. Sci. USA 101, 15905–15910 (2004).
Annibale, P., Scarselli, M., Greco, M. & Radenovic, A. Identification of the factors affecting co-localization precision for quantitative multicolor localization microscopy. Opt. Nanoscopy 1, 9 (2012).
Habuchi, S., Tsutsui, H., Kochaniak, A.B., Miyawaki, A. & van Oijen, A.M. mKikGR, a monomeric photoswitchable fluorescent protein. PLoS ONE 3, e3944 (2008).
Durisic, N. et al. Stoichiometry of the human glycine receptor revealed by direct subunit counting. J. Neurosci. 32, 12915–12920 (2012).
Lynch, J.W. Native glycine receptor subtypes and their physiological roles. Neuropharmacology 56, 303–309 (2009).
Simonson, P.D. et al. Counting bungarotoxin binding sites of nicotinic acetylcholine receptors in mammalian cells with high signal/noise ratios. Biophys. J. 99, L81–L83 (2010).
Ulbrich, M.H. & Isacoff, E.Y. Subunit counting in membrane-bound proteins. Nat. Methods 4, 319–321 (2007).
Puchner, E.M., Walter, J.M., Kasper, R., Huang, B. & Lim, W.A. Counting molecules in single organelles with superresolution microscopy allows tracking of the endosome maturation trajectory. Proc. Natl. Acad. Sci. USA 110, 16015–16020 (2013).
Goldin, A.L. Maintenance of Xenopus laevis and oocyte injection. Methods Enzymol. 207, 266–279 (1992).
Wang, M.H. A technical consideration concerning the removal of oocyte vitelline membranes for patch clamp recording. Biochem. Biophys. Res. Commun. 324, 971–972 (2004).
Heyes, C.D., Kobitski, A.Y., Breus, V.V. & Nienhaus, G.U. Effect of the shell on the blinking statistics of core-shell quantum dots: a single-particle fluorescence study. Phys. Rev. B 75, 125431 (2007).
Acknowledgements
We thank J. Dent (McGill University) for the GlyR-α–VFP and GlyR-β–VFP; X. Zhuang (Harvard University), for mEos2; T. Xu and P. Xu (Chinese Academy of Sciences) for mEos3.1 and mEos3.2; A. McEvoy (University of California, Berkeley) for Dendra2, mClavGR2, mMaple and PA-GFP; T. Misgeld (Technical University of Munich) for PAmCherry; and M. Ulbrich (University Freiburg) for pGEMHE-α1E-Ca2+-mEGFP. We thank C.D. Heyes (University of Arkansas) for critical reading of the manuscript, and I. Vernos and J. Cela-Gallego (Center for Genomic Regulation) for the oocytes. This work was supported in part by Fundació Cellex Barcelona and in part by a Marie Curie International Reintegration grant to M.L. (FP7-PEOPLE-2010-RG). N.D. is a NEST postdoctoral fellow partially supported by the Marie Curie Co-funding of Regional, National and International Programs (COFUND) action of the European Commission.
Author information
Authors and Affiliations
Contributions
M.L. and N.D. designed the experiments. N.D. and L.L.-C. performed the experiments and the data analysis. Á.S.-Á. provided reagents and performed the cloning of the plasmids. J.S.B. carried out simulations and wrote software. M.L. and N.D. wrote the manuscript. M.L. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1 and 2, and Supplementary Notes 1–3 (PDF 2835 kb)
Rights and permissions
About this article
Cite this article
Durisic, N., Laparra-Cuervo, L., Sandoval-Álvarez, Á. et al. Single-molecule evaluation of fluorescent protein photoactivation efficiency using an in vivo nanotemplate. Nat Methods 11, 156–162 (2014). https://doi.org/10.1038/nmeth.2784
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2784
This article is cited by
-
Ligand functionalization of titanium nanopattern enables the analysis of cell–ligand interactions by super-resolution microscopy
Nature Protocols (2022)
-
Correction of multiple-blinking artifacts in photoactivated localization microscopy
Nature Methods (2022)
-
Single-molecule localization microscopy
Nature Reviews Methods Primers (2021)
-
Cellular dynamics of the SecA ATPase at the single molecule level
Scientific Reports (2021)
-
Single-particle tracking photoactivated localization microscopy of membrane proteins in living plant tissues
Nature Protocols (2021)