Abstract
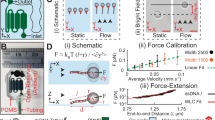
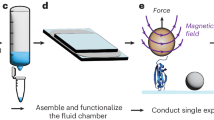
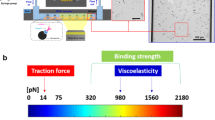
Single-molecule force spectroscopy enables mechanical testing of individual proteins, but low experimental throughput limits the ability to screen constructs in parallel. We describe a microfluidic platform for on-chip expression, covalent surface attachment and measurement of single-molecule protein mechanical properties. A dockerin tag on each protein molecule allowed us to perform thousands of pulling cycles using a single cohesin-modified cantilever. The ability to synthesize and mechanically probe protein libraries enables high-throughput mechanical phenotyping.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
05 November 2014
In the version of this article initially published, the grant "European Research Council Grant Cellufuel (Advanced Grant 294438)" was mistakenly left out of the Acknowledgements. The error has been corrected in the HTML and PDF versions of the article.
References
Thomas, W.E., Trintchina, E., Forero, M., Vogel, V. & Sokurenko, E.V. Cell 109, 913–923 (2002).
Li, C. & Xu, Q. Cell. Signal. 12, 435–445 (2000).
Engler, A.J., Sen, S., Sweeney, H.L. & Discher, D.E. Cell 126, 677–689 (2006).
Müller, D., Helenius, J., Alsteens, D. & Dufrêne, Y.F. Nat. Chem. Biol. 5, 383–390 (2009).
Florin, E.-L., Moy, V.T. & Gaub, H.E. Science 264, 415–417 (1994).
Rief, M., Gautel, M., Oesterhelt, F., Fernandez, J. & Gaub, H. Science 276, 1109–1112 (1997).
Fernandez, J.M. & Li, H. Science 303, 1674–1678 (2004).
Oesterhelt, F. et al. Science 288, 143–146 (2000).
Abbondanzieri, E.A., Greenleaf, W.J., Shaevitz, J.W., Landick, R. & Block, S.M. Nature 438, 460–465 (2005).
Bryant, Z. et al. Nature 424, 338–341 (2003).
Linke, W.A. Cardiovasc. Res. 77, 637–648 (2008).
Yin, J. et al. Proc. Natl. Acad. Sci. USA 102, 15815–15820 (2005).
Bayer, E.A., Belaich, J.-P., Shoham, Y. & Lamed, R. Annu. Rev. Microbiol. 58, 521–554 (2004).
Maerkl, S.J. & Quake, S.R. Science 315, 233–237 (2007).
Rockel, S., Geertz, M., Hens, K., Deplancke, B. & Maerkl, S.J. Nucleic Acids Res. 41, e52 (2013).
Gerber, D., Maerkl, S.J. & Quake, S.R. Nat. Methods 6, 71–74 (2009).
Otten, M., Wolf, P. & Gaub, H.E. Lab Chip 13, 4198–4204 (2013).
Thorsen, T., Maerkl, S.J. & Quake, S.R. Science 298, 580–584 (2002).
Garcia-Cordero, J.L. & Maerkl, S.J. Chem. Commun. 49, 1264–1266 (2013).
Gumpp, H., Stahl, S.W., Strackharn, M., Puchner, E.M. & Gaub, H.E. Rev. Sci. Instrum. 80, 063704 (2009).
Stahl, S.W. et al. Proc. Natl. Acad. Sci. USA 109, 20431–20436 (2012).
Li, L., Huang, H.H.-L., Badilla, C.L. & Fernandez, J.M. J. Mol. Biol. 345, 817–826 (2005).
Rief, M., Pascual, J., Saraste, M. & Gaub, H.E. J. Mol. Biol. 286, 553–561 (1999).
Dietz, H. & Rief, M. Proc. Natl. Acad. Sci. USA 103, 1244–1247 (2006).
Greene, D.N. et al. Biophys. J. 95, 1360–1370 (2008).
Puchner, E.M., Franzen, G., Gautel, M. & Gaub, H.E. Biophys. J. 95, 426–434 (2008).
Jobst, M.A., Schoeler, C., Malinowska, K. & Nash, M.A. J. Vis. Exp. 82, e50950 (2013).
Gibson, C.T., Smith, D.A. & Roberts, C.J. Nanotechnology 16, 234–238 (2005).
Gibson, D.G. et al. Nat. Methods 6, 343–345 (2009).
Zimmermann, J.L., Nicolaus, T., Neuert, G. & Blank, K. Nat. Protoc. 5, 975–985 (2010).
Huang, B., Wu, H., Kim, S. & Zare, R.N. Lab Chip 5, 1005–1007 (2005).
Hutter, J.L. & Bechhoefer, J. Rev. Sci. Instrum. 64, 1868 (1993).
Cook, S.M. et al. Nanotechnology 17, 2135–2145 (2006).
Proksch, R., Schäffer, T.E., Cleveland, J.P., Callahan, R.C. & Viani, M.B. Nanotechnology 15, 1344–1350 (2004).
Acknowledgements
M.O. is grateful to the Elite Network of Bavaria (IDK-NBT) for a doctoral fellowship. M.A.N. acknowledges support from Society in Science—The Branco Weiss Fellowship administered by the ETH Zürich. The authors acknowledge support from the DFG Sonderforschungsbereich 1032 and the European Research Council Grant Cellufuel (Advanced Grant 294438). The authors thank E. Bayer (Weizmann Institute) for starting genetic materials used for Doc and Coh modules.
Author information
Authors and Affiliations
Contributions
M.O., M.A.N. and H.E.G. designed the research; M.O., W.O., M.A.J. and T.V. performed experiments; D.A.P. helped with immobilization strategies; M.O., W.O., M.A.J., L.F.M. and M.A.N. performed data analysis; M.O., W.O., M.A.J., M.A.N. and H.E.G. cowrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Microfluidic chip overview.
(a) Photograph of a microfluidic chip bonded to a glass slide with a US dime for scale. Control channels are filled with food dye for better visualization. (b) Pattern of a typical DNA array, consisting of repeats of rows with four different genes and one row with nothing spotted as negative control. (c) Photograph of a bonded PDMS chip onto the glass slide with DNA spots in the back chamber. The orange highlighted frame shows a zoom in of the bottom left corner. (d) Typical fluorescence collage assembled from 640 single fluorescence micrographs of each protein chamber on one single chip shows pattern of expressed protein (assembly not to scale). Fluorescence signal of TagRFP reveals expression levels and Dockerin specificity. Here, low passivation of the protein chamber facilitates visualization. (e) Three of 640 adjacent dumbbell-shaped chambers, one with sfGFP DNA spotted (left), one with Xylanase DNA (center) and one negative control without DNA (right). Control channels are visualized with food dye: neck valve (green), sandwich valve (red), and button valve (blue). (f) Fluorescence images showing GFP signal (top) from expressed and immobilized ybbR-sfGFP-Dockerin (left), ybbR-Xylanase-Dockerin (center) with negative control lacking the spotted DNA (right). The bottom row shows the signal from the TagRFP detection construct, which specifically bound to the Dockerin tag via the Cohesin domain.
Supplementary Figure 7 Exemplary force traces
Example curves showing (a) uninterpretable interaction, (b) non-specific interaction of cantilever with surface, (c) no interaction, and (d) a specific Xylanase-Dockerin unfolding and unbinding trace. Curves similar to those shown in a-c were excluded from the analysis.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–3 and Supplementary Discussion (PDF 6736 kb)
Source data
Rights and permissions
About this article
Cite this article
Otten, M., Ott, W., Jobst, M. et al. From genes to protein mechanics on a chip. Nat Methods 11, 1127–1130 (2014). https://doi.org/10.1038/nmeth.3099
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3099
This article is cited by
-
Low cost and massively parallel force spectroscopy with fluid loading on a chip
Nature Communications (2022)
-
Protein nanomechanics in biological context
Biophysical Reviews (2021)
-
Multiplex flow magnetic tweezers reveal rare enzymatic events with single molecule precision
Nature Communications (2020)
-
Concurrent atomic force spectroscopy
Communications Physics (2019)
-
High-speed force spectroscopy: microsecond force measurements using ultrashort cantilevers
Biophysical Reviews (2019)