Abstract
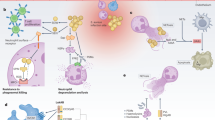
The molecular processes underlying epidemic waves of methicillin-resistant Staphylococcus aureus (MRSA) infection are poorly understood1. Although a major role has been attributed to the acquisition of virulence determinants by horizontal gene transfer2, there are insufficient epidemiological and functional data supporting that concept. We here report the spread of clones containing a previously extremely rare3,4 mobile genetic element–encoded gene, sasX. We demonstrate that sasX has a key role in MRSA colonization and pathogenesis, substantially enhancing nasal colonization, lung disease and abscess formation and promoting mechanisms of immune evasion. Moreover, we observed the recent spread of sasX from sequence type 239 (ST239) to invasive clones belonging to other sequence types. Our study identifies sasX as a quickly spreading crucial determinant of MRSA pathogenic success and a promising target for therapeutic interference. Our results provide proof of principle that horizontal gene transfer of key virulence determinants drives MRSA epidemic waves.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chambers, H.F. & DeLeo, F.R. Waves of resistance: Staphylococcus aureus in the antibiotic era. Nat. Rev. Microbiol. 7, 629–641 (2009).
Diep, B.A., Carleton, H.A., Chang, R.F., Sensabaugh, G.F. & Perdreau-Remington, F. Roles of 34 virulence genes in the evolution of hospital- and community-associated strains of methicillin-resistant Staphylococcus aureus. J. Infect. Dis. 193, 1495–1503 (2006).
Holden, M.T. et al. Genome sequence of a recently emerged, highly transmissible, multi-antibiotic- and antiseptic-resistant variant of methicillin-resistant Staphylococcus aureus, sequence type 239 (TW). J. Bacteriol. 192, 888–892 (2010).
Harris, S.R. et al. Evolution of MRSA during hospital transmission and intercontinental spread. Science 327, 469–474 (2010).
Lowy, F.D. Antimicrobial resistance: the example of Staphylococcus aureus. J. Clin. Invest. 111, 1265–1273 (2003).
DeLeo, F.R. & Chambers, H.F. Reemergence of antibiotic-resistant Staphylococcus aureus in the genomics era. J. Clin. Invest. 119, 2464–2474 (2009).
Klevens, R.M. et al. Invasive methicillin-resistant Staphylococcus aureus infections in the United States. J. Am. Med. Assoc. 298, 1763–1771 (2007).
Hiramatsu, K., Cui, L., Kuroda, M. & Ito, T. The emergence and evolution of methicillin-resistant Staphylococcus aureus. Trends Microbiol. 9, 486–493 (2001).
Liu, Y. et al. Molecular evidence for spread of two major methicillin-resistant Staphylococcus aureus clones with a unique geographic distribution in Chinese hospitals. Antimicrob. Agents Chemother. 53, 512–518 (2009).
Li, Y. et al. Complete genome sequence of Staphylococcus aureus T0131, an ST239-MRSA-SCCmec type III clone isolated in china. J. Bacteriol. 193, 3411–3412 (2011).
Söderquist, B. et al. Staphylococcus epidermidis surface protein I (SesI): a marker of the invasive capacity of S. epidermidis? J. Med. Microbiol. 58, 1395–1397 (2009).
Aires de Sousa, M. et al. Frequent recovery of a single clonal type of multidrug-resistant Staphylococcus aureus from patients in two hospitals in Taiwan and China. J. Clin. Microbiol. 41, 159–163 (2003).
Arakere, G. et al. Genotyping of methicillin-resistant Staphylococcus aureus strains from two hospitals in Bangalore, South India. J. Clin. Microbiol. 43, 3198–3202 (2005).
Ip, M., Lyon, D.J., Chio, F., Enright, M.C. & Cheng, A.F. Characterization of isolates of methicillin-resistant Staphylococcus aureus from Hong Kong by phage typing, pulsed-field gel electrophoresis, and fluorescent amplified-fragment length polymorphism analysis. J. Clin. Microbiol. 41, 4980–4985 (2003).
Ko, K.S. et al. Distribution of major genotypes among methicillin-resistant Staphylococcus aureus clones in Asian countries. J. Clin. Microbiol. 43, 421–426 (2005).
Foster, T.J. & Hook, M. Surface protein adhesins of Staphylococcus aureus. Trends Microbiol. 6, 484–488 (1998).
Wertheim, H.F. et al. The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect. Dis. 5, 751–762 (2005).
von Eiff, C., Becker, K., Machka, K., Stammer, H. & Peters, G. Nasal carriage as a source of Staphylococcus aureus bacteremia. Study Group. N. Engl. J. Med. 344, 11–16 (2001).
Weidenmaier, C. et al. Role of teichoic acids in Staphylococcus aureus nasal colonization, a major risk factor in nosocomial infections. Nat. Med. 10, 243–245 (2004).
O'Brien, L.M., Walsh, E.J., Massey, R.C., Peacock, S.J. & Foster, T.J. Staphylococcus aureus clumping factor B (ClfB) promotes adherence to human type I cytokeratin 10: implications for nasal colonization. Cell. Microbiol. 4, 759–770 (2002).
Wertheim, H.F. et al. Key role for clumping factor B in Staphylococcus aureus nasal colonization of humans. PLoS Med. 5, e17 (2008).
Corrigan, R.M., Miajlovic, H. & Foster, T.J. Surface proteins that promote adherence of Staphylococcus aureus to human desquamated nasal epithelial cells. BMC Microbiol. 9, 22 (2009).
Foster, T.J. Immune evasion by staphylococci. Nat. Rev. Microbiol. 3, 948–958 (2005).
Otto, M. Staphylococcal biofilms. Curr. Top. Microbiol. Immunol. 322, 207–228 (2008).
Otto, M. Bacterial evasion of antimicrobial peptides by biofilm formation. Curr. Top. Microbiol. Immunol. 306, 251–258 (2006).
Vuong, C., Saenz, H.L., Gotz, F. & Otto, M. Impact of the agr quorum-sensing system on adherence to polystyrene in Staphylococcus aureus. J. Infect. Dis. 182, 1688–1693 (2000).
Moran, G.J. et al. Methicillin-resistant S. aureus infections among patients in the emergency department. N. Engl. J. Med. 355, 666–674 (2006).
Kennedy, A.D. et al. Targeting of α-hemolysin by active or passive immunization decreases severity of USA300 skin infection in a mouse model. J. Infect. Dis. 202, 1050–1058 (2010).
Wang, R. et al. Identification of novel cytolytic peptides as key virulence determinants for community-associated MRSA. Nat. Med. 13, 1510–1514 (2007).
Koh, S.S. et al. Molecular classification of melanomas and nevi using gene expression microarray signatures and formalin-fixed and paraffin-embedded tissue. Mod. Pathol. 22, 538–546 (2009).
Acknowledgements
This study was supported by a grant from the China National Clinical Key Subject to Y.L., the National Natural Science Foundation of China (grants 30900026 and 81171623) and the Shanghai Pujiang Program (grant 09PJ1402300) to M.L., and the Intramural Research Program of the National Institute of Allergy and Infectious Diseases, US National Institutes of Health (to M.O.).
Author information
Authors and Affiliations
Contributions
M.L., X.D., A.E.V., D.W., Y.S., Y.T., J.H. and F.Y. conducted experiments. M.L., B.A.D., Y.L. and M.O. planned and supervised experiments. M.O. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–3 and Supplementary Methods (PDF 1158 kb)
Supplementary Data
Characteristics of isolates investigated in this study (XLS 195 kb)
Rights and permissions
About this article
Cite this article
Li, M., Du, X., Villaruz, A. et al. MRSA epidemic linked to a quickly spreading colonization and virulence determinant. Nat Med 18, 816–819 (2012). https://doi.org/10.1038/nm.2692
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2692
This article is cited by
-
Isolation and characterization of novel Staphylococcus aureus bacteriophage Hesat from dairy origin
Applied Microbiology and Biotechnology (2024)
-
Molecular characterisation of Staphylococcus aureus in school-age children in Guangzhou: associations among agr types, virulence genes, sequence types, and antibiotic resistant phenotypes
BMC Microbiology (2023)
-
Prevalence of S. aureus and/or MRSA from seafood products from Indian seafood products
BMC Microbiology (2022)
-
Staphylococcus epidermidis clones express Staphylococcus aureus-type wall teichoic acid to shift from a commensal to pathogen lifestyle
Nature Microbiology (2021)
-
The protective roles of tea tree oil extracts in bovine mammary epithelial cells and polymorphonuclear leukocytes
Journal of Animal Science and Biotechnology (2020)