Abstract
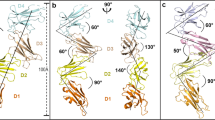
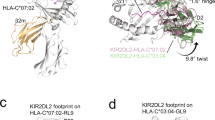
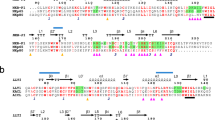
Leukocyte immunoglobulin-like receptor 1 (LIR-1), an inhibitory receptor expressed on monocytes, dendritic cells and lymphocytes, regulates cellular function by binding a broad range of classical and nonclassical major histocompatibility complex (MHC) class I molecules, and the human cytomegalovirus MHC class I homolog UL18. Here we describe the 3.4-Å crystal structure of a complex between the LIR-1 D1D2 domains and the MHC class I molecule HLA-A2. LIR-1 contacts the mostly conserved β2-microglobulin and α3 domains of HLA-A2. The LIR-1 binding site comprises residues at the interdomain hinge, and a patch at the D1 tip. The structure shows how LIR-1 recognizes UL18 and diverse MHC class I molecules, and indicates that a similar mode of MHC class I recognition is used by other LIR family members.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Borges, L. & Cosman, D. LIRs/ILTs/MIRs, inhibitory and stimulatory Ig-superfamily receptors expressed in myeloid and lymphoid cells. Cytokine Growth Factor Rev. 11, 209–217 (2000).
Cella, M., Nakajima, H., Facchetti, F., Hoffmann, T. & Colonna, M. ILT receptors at the interface between lymphoid and myeloid cells. Curr. Top. Microbiol. Immunol. 251, 161–166 (2000).
Colonna, M. et al. A common inhibitory receptor for major histocompatibility complex class I molecules on human lymphoid and myelomonocytic cells. J. Exp. Med. 186, 1809–1818 (1997).
Cosman, D. et al. A novel immunoglobulin superfamily receptor for cellular and viral MHC class I molecules. Immunity 7, 273–282 (1997).
Beck, S. & Barrell, B.G. Human cytomegalovirus encodes a glycoprotein homologous to MHC class-I antigens. Nature 331, 269–272 (1988).
Browne, H., Smith, G., Beck, S. & Minson, T. A complex between the MHC class I homologue encoded by human cytomegalovirus and β2 microglobulin. Nature 347, 770–772 (1990).
Fahnestock, M.L. et al. The MHC class I homolog encoded by human cytomegalovirus binds endogenous peptides. Immunity 3, 583–590 (1995).
Chapman, T.L., Heikema, A.P. & Bjorkman, P.J. The inhibitory receptor LIR-1 uses a common binding interaction to recognize class I MHC molecules and the viral MHC homolog UL18. Immunity 11, 603–611 (1999).
Shiroishi, M. et al. Human inhibitory receptors ILT2 and ILT4 compete with CD8 for MHC class I binding and bind preferentially to HLA-G. Proc. Natl. Acad. Sci. USA 100, 8856–8861 (2003).
Natarajan, K., Dimasi, N., Wang, J., Mariuzza, R.A. & Margulies, D.H. Structure and function of natural killer cell receptors: multiple molecular solutions to self, nonself discrimination. Annu. Rev. Immunol. 20, 853–885 (2002).
Chapman, T.L., Heikema, A.P., West, A.P., Jr. & Bjorkman, P.J. Crystal structure and ligand binding properties of the D1D2 region of the inhibitory receptor LIR-1 (ILT2). Immunity 13, 727–736 (2000).
Madden, D.R., Garboczi, D.N. & Wiley, D.C. The antigenic identity of peptide-MHC complexes: a comparison of the conformations of five viral peptides presented by HLA-A2. Cell 75, 693–708 (1993).
Rudolph, M.G. & Wilson, I.A. The specificity of TCR/pMHC interaction. Curr. Opin. Immunol. 14, 52–65 (2002).
Fan, Q.R., Long, E.O. & Wiley, D.C. Crystal structure of the human natural killer cell inhibitory receptor KIR2DL1-HLA-Cw4 complex. Nat. Immunol. 2, 452–460 (2001).
Gao, G.F. et al. Crystal structure of the complex between human CD8α(α) and HLA-A2. Nature 387, 630–634 (1997).
Cosman, D., Fanger, N. & Borges, L. Human cytomegalovirus, MHC class I and inhibitory signalling receptors: more questions than answers. Immunol. Rev. 168, 177–185 (1999).
Willcox, B.E. et al. Crystal structure of LIR-2 (ILT4) at 1.8 Å: differences from LIR-1 (ILT2) in regions implicated in the binding of the human cytomegalovirus class I MHC homolog UL18. BMC. Struct. Biol. 2, 6 (2002).
Allen, R.L., Raine, T., Haude, A., Trowsdale, J. & Wilson, M.J. Leukocyte receptor complex-encoded immunomodulatory receptors show differing specificity for alternative HLA-B27 structures. J. Immunol. 167, 5543–5547 (2001).
van der Merwe, P.A., Davis, S.J., Shaw, A.S. & Dustin, M.L. Cytoskeletal polarization and redistribution of cell-surface molecules during T cell antigen recognition. Semin. Immunol. 12, 5–21 (2000).
Dietrich, J., Cella, M. & Colonna, M. Ig-like transcript 2 (ILT2)/leukocyte Ig-like receptor 1 (LIR1) inhibits TCR signaling and actin cytoskeleton reorganization. J. Immunol. 166, 2514–2521 (2001).
Young, N.T., Uhrberg, M., Phillips, J.H., Lanier, L.L. & Parham, P. Differential expression of leukocyte receptor complex-encoded Ig-like receptors correlates with the transition from effector to memory CTL. J. Immunol. 166, 3933–3941 (2001).
Wende, H., Colonna, M., Ziegler, A. & Volz, A. Organization of the leukocyte receptor cluster (LRC) on human chromosome 19q13.4. Mamm. Genome 10, 154–160 (1999).
Herr, A.B., Ballister, E.R. & Bjorkman, P.J. Insights into mucosal immunity from the structures of human FcaRI and its complex with IgA1-Fc. Nature 423, 614–620 (2003).
Kubagawa, H., Burrows, P.D. & Cooper, M.D. A novel pair of immunoglobulin-like receptors expressed by B cells and myeloid cells. Proc. Natl. Acad. Sci. USA 94, 5261–5266 (1997).
Martin, A.M., Kulski, J.K., Witt, C., Pontarotti, P. & Christiansen, F.T. Leukocyte Ig-like receptor complex (LRC) in mice and men. Trends Immunol. 23, 81–88 (2002).
Barten, R., Torkar, M., Haude, A., Trowsdale, J. & Wilson, M.J. Divergent and convergent evolution of NK-cell receptors. Trends Immunol. 22, 52–57 (2001).
Trowsdale, J. Genetic and functional relationships between MHC and NK receptor genes. Immunity 15, 363–374 (2001).
Garboczi, D.N., Hung, D.T. & Wiley, D.C. HLA-A2-peptide complexes: refolding and crystallization of molecules expressed in Escherichia coli and complexed with single antigenic peptides. Proc. Natl. Acad. Sci. USA 89, 3429–3433 (1992).
Pace, C.N., Vajdos, F., Fee, L., Grimsley, G. & Gray, T. How to measure and predict the molar absorption coefficient of a protein. Protein Sci. 4, 2411–2423 (1995).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Navaza, J. AMORE—an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
CCP4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brunger, A.T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D. Biol. Crystallogr. 54, 905–921 (1998).
Su, X.D. et al. Crystal structure of hemolin: a horseshoe shape with implications for homophilic adhesion. Science 281, 991–995 (1998).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Murphy, M.E.P. Raster3D Version 2.0, a program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 (1994).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
DeLano, W.L. The PyMOL Molecular Graphics System. (DeLano Scientific, San Carlos, California, 2002).
Ding, Y.H. et al. Two human T cell receptors bind in a similar diagonal mode to the HLA-A2/Tax peptide complex using different TCR amino acids. Immunity 8, 403–411 (1998).
Acknowledgements
We thank members of the Bjorkman laboratory for technical assistance, and C. O'Callaghan and A. van der Merwe for critical reading of the manuscript. B.E.W. was supported by a Wellcome Trust Travelling Fellowship and is now funded by a Medical Research Council Career Development Award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1.
Assessment of agreement between the atomic model and the x-ray data. From the top left and working from left to right and top to bottom, the graphs indicate the following: the Wilson plot; optical resolution as a function of the crystallographic resolution; data completeness and structure factor error as a function of the d spacing; maximal and minimal coordinate error dependence on d spacing; a stereographic projection of the averaged radial data structure-factor data completeness; and the R-factor dependence and Luzzati plot with an atomic error = 0.532. All graphs are taken directly from the program SFcheck output1. (PDF 273 kb)
1. Vaguine, A. A., Richelle, J. & Wodak, S. J. SFCHECK: a unified set of procedures for evaluating the quality of macromolecular structure-factor data and their agreement with the atomic model. Acta Crystallogr D Biol Crystallogr 55, 191-205. (1999).
Rights and permissions
About this article
Cite this article
Willcox, B., Thomas, L. & Bjorkman, P. Crystal structure of HLA-A2 bound to LIR-1, a host and viral major histocompatibility complex receptor. Nat Immunol 4, 913–919 (2003). https://doi.org/10.1038/ni961
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni961
This article is cited by
-
Distinct frequency patterns of LILRB3 and LILRA6 allelic variants in Europeans
Immunogenetics (2023)
-
Epithelial cells remove precancerous cells by cell competition via MHC class I–LILRB3 interaction
Nature Immunology (2021)
-
Structural basis for RIFIN-mediated activation of LILRB1 in malaria
Nature (2020)
-
Structures of the four Ig-like domain LILRB2 and the four-domain LILRB1 and HLA-G1 complex
Cellular & Molecular Immunology (2020)
-
Engagement of MHC class I by the inhibitory receptor LILRB1 suppresses macrophages and is a target of cancer immunotherapy
Nature Immunology (2018)