Abstract
Natural killer (NK) cells are innate lymphocytes that exhibit many features of adaptive immunity, including clonal proliferation and long-lived memory. Here we demonstrate that the BTB-ZF transcription factor Zbtb32 (also known as ROG, FAZF, TZFP and PLZP) was essential for the proliferative burst and protective capacity of virus-specific NK cells. Signals from proinflammatory cytokines were both necessary and sufficient to induce high expression of Zbtb32 in NK cells. Zbtb32 facilitated NK cell proliferation during infection by antagonizing the anti-proliferative factor Blimp-1 (Prdm1). Our data support a model in which Zbtb32 acts as a cellular 'hub' through which proinflammatory signals instruct a 'proliferation-permissive' state in NK cells, thereby allowing their prolific expansion in response to viral infection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Sun, J.C. & Lanier, L.L. NK cell development, homeostasis and function: parallels with CD8(+) T cells. Nat. Rev. Immunol. 11, 645–657 (2011).
Smith, H.R. et al. Recognition of a virus-encoded ligand by a natural killer cell activation receptor. Proc. Natl. Acad. Sci. USA 99, 8826–8831 (2002).
Arase, H., Mocarski, E.S., Campbell, A.E., Hill, A.B. & Lanier, L.L. Direct recognition of cytomegalovirus by activating and inhibitory NK cell receptors. Science 296, 1323–1326 (2002).
Daniels, K.A. et al. Murine cytomegalovirus is regulated by a discrete subset of natural killer cells reactive with monoclonal antibody to Ly49H. J. Exp. Med. 194, 29–44 (2001).
Brown, M.G. et al. Vital involvement of a natural killer cell activation receptor in resistance to viral infection. Science 292, 934–937 (2001).
Bubic, I. et al. Gain of virulence caused by loss of a gene in murine cytomegalovirus. J. Virol. 78, 7536–7544 (2004).
Dokun, A.O. et al. Specific and nonspecific NK cell activation during virus infection. Nat. Immunol. 2, 951–956 (2001).
Sun, J.C., Beilke, J.N. & Lanier, L.L. Adaptive immune features of natural killer cells. Nature 457, 557–561 (2009).
Kelly, K.F. & Daniel, J.M. POZ for effect–POZ-ZF transcription factors in cancer and development. Trends Cell Biol. 16, 578–587 (2006).
Beaulieu, A.M. & Sant'Angelo, D.B. The BTB-ZF family of transcription factors: key regulators of lineage commitment and effector function development in the immune system. J. Immunol. 187, 2841–2847 (2011).
Maeda, T. et al. Regulation of B versus T lymphoid lineage fate decision by the proto-oncogene LRF. Science 316, 860–866 (2007).
Bilic, I. et al. Negative regulation of CD8 expression via Cd8 enhancer-mediated recruitment of the zinc finger protein MAZR. Nat. Immunol. 7, 392–400 (2006).
Sakaguchi, S. et al. The zinc-finger protein MAZR is part of the transcription factor network that controls the CD4 versus CD8 lineage fate of double-positive thymocytes. Nat. Immunol. 11, 442–448 (2010).
Sun, G. et al. The zinc finger protein cKrox directs CD4 lineage differentiation during intrathymic T cell positive selection. Nat. Immunol. 6, 373–381 (2005).
He, X. et al. The zinc finger transcription factor Th-POK regulates CD4 versus CD8 T-cell lineage commitment. Nature 433, 826–833 (2005).
Muroi, S. et al. Cascading suppression of transcriptional silencers by ThPOK seals helper T cell fate. Nat. Immunol. 9, 1113–1121 (2008).
Kovalovsky, D. et al. The BTB-zinc finger transcriptional regulator PLZF controls the development of invariant natural killer T cell effector functions. Nat. Immunol. 9, 1055–1064 (2008).
Alonzo, E.S. et al. Development of promyelocytic zinc finger and ThPOK-expressing innate gamma delta T cells is controlled by strength of TCR signaling and Id3. J. Immunol. 184, 1268–1279 (2010).
Savage, A.K. et al. The transcription factor PLZF directs the effector program of the NKT cell lineage. Immunity 29, 391–403 (2008).
Kreslavsky, T. et al. TCR-inducible PLZF transcription factor required for innate phenotype of a subset of gammadelta T cells with restricted TCR diversity. Proc. Natl. Acad. Sci. USA 106, 12453–12458 (2009).
Dent, A.L., Shaffer, A.L., Yu, X., Allman, D. & Staudt, L.M. Control of inflammation, cytokine expression, and germinal center formation by BCL-6. Science 276, 589–592 (1997).
Ye, B.H. et al. The BCL-6 proto-oncogene controls germinal-centre formation and Th2-type inflammation. Nat. Genet. 16, 161–170 (1997).
Yu, D. et al. The transcriptional repressor Bcl-6 directs T follicular helper cell lineage commitment. Immunity 31, 457–468 (2009).
Johnston, R.J. et al. Bcl6 and Blimp-1 are reciprocal and antagonistic regulators of T follicular helper cell differentiation. Science 325, 1006–1010 (2009).
Nurieva, R.I. et al. Bcl6 mediates the development of T follicular helper cells. Science 325, 1001–1005 (2009).
Bezman, N.A. et al. Molecular definition of the identity and activation of natural killer cells. Nat. Immunol. 13, 1000–1009 (2012).
Bukowski, J.F., Warner, J.F., Dennert, G. & Welsh, R.M. Adoptive transfer studies demonstrating the antiviral effect of natural killer cells in vivo. J. Exp. Med. 161, 40–52 (1985).
Omori, M. et al. CD8 T cell-specific downregulation of histone hyperacetylation and gene activation of the IL-4 gene locus by ROG, repressor of GATA. Immunity 19, 281–294 (2003).
Piazza, F., Costoya, J.A., Merghoub, T., Hobbs, R.M. & Pandolfi, P.P. Disruption of PLZP in mice leads to increased T-lymphocyte proliferation, cytokine production, and altered hematopoietic stem cell homeostasis. Mol. Cell. Biol. 24, 10456–10469 (2004).
Miaw, S.C., Choi, A., Yu, E., Kishikawa, H. & Ho, I.C. ROG, repressor of GATA, regulates the expression of cytokine genes. Immunity 12, 323–333 (2000).
Hirahara, K. et al. Repressor of GATA regulates TH2-driven allergic airway inflammation and airway hyperresponsiveness. J. Allergy Clin. Immunol. 122, 512–520 (2008).
Hirasaki, Y. et al. Repressor of GATA negatively regulates murine contact hypersensitivity through the inhibition of type-2 allergic responses. Clin. Immunol. 139, 267–276 (2011).
Miaw, S.C., Kang, B.Y., White, I.A. & Ho, I.C. A repressor of GATA-mediated negative feedback mechanism of T cell activation. J. Immunol. 172, 170–177 (2004).
Marrack, P. & Kappler, J. Control of T cell viability. Annu. Rev. Immunol. 22, 765–787 (2004).
Dunkle, A., Dzhagalov, I., Gordy, C. & He, Y.W. Transfer of CD8+ T cell memory using Bcl-2 as a marker. J. Immunol. 190, 940–947 (2013).
Stergachis, A.B. et al. Developmental fate and cellular maturity encoded in human regulatory DNA landscapes. Cell 154, 888–903 (2013).
Sun, J.C. et al. Proinflammatory cytokine signaling required for the generation of natural killer cell memory. J. Exp. Med. 209, 947–954 (2012).
Miyagi, T. et al. High basal STAT4 balanced by STAT1 induction to control type 1 interferon effects in natural killer cells. J. Exp. Med. 204, 2383–2396 (2007).
Hesslein, D.G. & Lanier, L.L. Transcriptional control of natural killer cell development and function. Adv. Immunol. 109, 45–85 (2011).
Crotty, S., Johnston, R.J. & Schoenberger, S.P. Effectors and memories: Bcl-6 and Blimp-1 in T and B lymphocyte differentiation. Nat. Immunol. 11, 114–120 (2010).
Kallies, A. et al. A role for Blimp1 in the transcriptional network controlling natural killer cell maturation. Blood 117, 1869–1879 (2011).
Yoon, H.S. et al. ZBTB32 Is an Early Repressor of the CIITA and MHC Class II Gene Expression during B Cell Differentiation to Plasma Cells. J. Immunol. 189, 2393–2403 (2012).
Shin, H.M. et al. Epigenetic modifications induced by Blimp-1 Regulate CD8(+) T cell memory progression during acute virus infection. Immunity 39, 661–675 (2013).
Bikoff, E.K., Morgan, M.A. & Robertson, E.J. An expanding job description for Blimp-1/PRDM1. Curr. Opin. Genet. Dev. 19, 379–385 (2009).
Sakurai, N. et al. The LRF transcription factor regulates mature B cell development and the germinal center response in mice. J. Clin. Invest. 121, 2583–2598 (2011).
Phan, R.T. & Dalla-Favera, R. The BCL6 proto-oncogene suppresses p53 expression in germinal-centre B cells. Nature 432, 635–639 (2004).
Phan, R.T., Saito, M., Basso, K., Niu, H. & Dalla-Favera, R. BCL6 interacts with the transcription factor Miz-1 to suppress the cyclin-dependent kinase inhibitor p21 and cell cycle arrest in germinal center B cells. Nat. Immunol. 6, 1054–1060 (2005).
Tunyaplin, C. et al. Direct repression of prdm1 by Bcl-6 inhibits plasmacytic differentiation. J. Immunol. 173, 1158–1165 (2004).
Vasanwala, F.H., Kusam, S., Toney, L.M. & Dent, A.L. Repression of AP-1 function: a mechanism for the regulation of Blimp-1 expression and B lymphocyte differentiation by the B cell lymphoma-6 protooncogene. J. Immunol. 169, 1922–1929 (2002).
Lopez-Verges, S. et al. Expansion of a unique CD57(+)NKG2Chi natural killer cell subset during acute human cytomegalovirus infection. Proc. Natl. Acad. Sci. USA 108, 14725–14732 (2011).
Foley, B. et al. Cytomegalovirus reactivation after allogeneic transplantation promotes a lasting increase in educated NKG2C+ natural killer cells with potent function. Blood 119, 2665–2674 (2012).
Foley, B. et al. Human cytomegalovirus (CMV)-induced memory-like NKG2C(+) NK cells are transplantable and expand in vivo in response to recipient CMV antigen. J. Immunol. 189, 5082–5088 (2012).
Romee, R. et al. Cytokine activation induces human memory-like NK cells. Blood 120, 4751–4760 (2012).
Hoshino, K. et al. Cutting edge: generation of IL-18 receptor-deficient mice: evidence for IL-1 receptor-related protein as an essential IL-18 binding receptor. J. Immunol. 162, 5041–5044 (1999).
Wu, C. et al. IL-12 receptor beta 2 (IL-12R beta 2)-deficient mice are defective in IL-12-mediated signaling despite the presence of high affinity IL-12 binding sites. J. Immunol. 165, 6221–6228 (2000).
Kaplan, M.H., Sun, Y.L., Hoey, T. & Grusby, M.J. Impaired IL-12 responses and enhanced development of Th2 cells in Stat4-deficient mice. Nature 382, 174–177 (1996).
Muller, U. et al. Functional role of type I and type II interferons in antiviral defense. Science 264, 1918–1921 (1994).
Fodil-Cornu, N. et al. Ly49h-deficient C57BL/6 mice: a new mouse cytomegalovirus-susceptible model remains resistant to unrelated pathogens controlled by the NK gene complex. J. Immunol. 181, 6394–6405 (2008).
Tripathy, S.K. et al. Continuous engagement of a self-specific activation receptor induces NK cell tolerance. J. Exp. Med. 205, 1829–1841 (2008).
Narni-Mancinelli, E. et al. Fate mapping analysis of lymphoid cells expressing the NKp46 cell surface receptor. Proc. Natl. Acad. Sci. USA 108, 18324–18329 (2011).
Shapiro-Shelef, M. et al. Blimp-1 is required for the formation of immunoglobulin secreting plasma cells and pre-plasma memory B cells. Immunity 19, 607–620 (2003).
Kim, S.K. et al. Generation of mucosal cytotoxic T cells against soluble protein by tissue-specific environmental and costimulatory signals. Proc. Natl. Acad. Sci. USA 95, 10814–10819 (1998).
Zheng, Y. et al. Genome-wide analysis of Foxp3 target genes in developing and mature regulatory T cells. Nature 445, 936–940 (2007).
Acknowledgements
We thank members of the Sun lab for technical support and experimental assistance, members of the Memorial Sloan-Kettering NK club for insightful comments and helpful discussions, A. Rudensky, M. van den Brink, M. Li, L. Lanier (University of California San Francisco), D. Sant'Angelo (Rutgers University) and L. Denzin (Rutgers University) for sharing antibodies and flow cytometry resources and for providing expertise critical to this study and manuscript, G. Gasteiger, K. Schluns (M.D. Anderson) and U. Koszinowski (Max von Pettenkofer-Institute) for providing many of the parent and recombinant viruses used in our study, S. Way (University of Minnesota) for providing femurs from Il12rb2−/− × Ifnar1−/− mice for use in making bone marrow chimeras, the Immunological Genome Consortium for providing the microarray data used in this study26, and J.P. Houchins and his team at R&D Systems for providing the experimental anti-Zbtb32 flow cytometry antibody used in this study. A.M.B. was supported by US National Institutes of Health T32 award (CA009149). J.C.S. was supported by the Searle Scholars Program, the Cancer Research Institute, and grants from the National Institutes of Health (AI085034 and AI100874).
Author information
Authors and Affiliations
Contributions
A.M.B. and C.L.Z. performed the experiments; T.N. provided the Zbtb32−/− mice and feedback on the manuscript; A.M.B. and J.C.S. designed the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
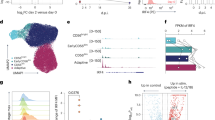
Supplementary Figure 1 Zbtb32 protein is upregulated in NK cells during MCMV infection.
Intranuclear Zbtb32 expression in splenic Ly49H+ NK cells from WT or Zbtb32–/– animals following MCMV infection (n = 3 animals per time point). Representative of 2 independent experiments.
Supplementary Figure 2 Zbtb32 is dispensable for IFN-γ production in vitro and cytotoxicity in vivo.
(a) WT or Zbtb32–/– splenocytes were stimulated for 18 h, then IFN-γ+ NK cells were identified by flow cytometry (n = 3 biological replicates per group). Data are representative of 2 – 3 experiments. (b) Equal numbers of CFSE-labeled splenocytes from WT (CFSElo), m157-transgenic (CFSEmid), or B2m–/– (CFSEhi) animals were co-transferred into WT (left) or Zbtb32–/– (right) recipients. After 24 h, the percentage of remaining cells from each population was determined in the spleen. Data are representative of 4 – 6 animals per genotype from 2 independent experiments.
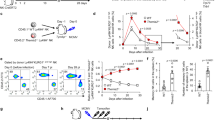
Supplementary Figure 3 Zbtb32-deficiency impairs viral antigen-independent NK cell proliferation during infection.
(a) Schematic of adoptive transfer experiments. Equal numbers of WT (CD45.1+) and Zbtb32–/–(CD45.2+) Ly49H+ NK cells were co-transferred into Ly49H-deficient hosts one day prior to infection with MCMV. (b) Expression of Zbtb32 mRNA in Ly49H+ and Ly49H– NK cells sorted from the spleens of WT animals on day 0 or 2 p.i. with MCMV (n = 7 animals per time point from 2 pooled experiments). Data are shown as fold expression relative to Ly49H+ NK cells at day 0 p.i. (c) Equal numbers of CFSE-labeled WT and Zbtb32–/– NK cells were transferred into congenic Ly49H-deficient hosts. CFSE dilution (left panel: WT, black; Zbtb32–/–, gray) and percentage of cells that divided > 5 rounds (right panel; n = 3 mice) in the Ly49H+ and Ly49H- transferred NK cells from the spleens of MCMV-infected animals on day 5 p.i. Representative of 2 independent experiments. (d) As in (c), except animals were infected with MCMVΔm157 and data are shown for Ly49H+ NK cells only (n = 3 mice). Representative of 2 independent experiments.
Supplementary Figure 4 NK cells develop normally in Zbtb32–/– animals.
(a) Percentage of NK cells in various organs in naïve WT or Zbtb32–/– mice (n = 3 mice per genotype). (b) Percentage (left panel) and absolute number (right panel) of CD122+Lin– cells that are pre-NK cells (pNK; DX5–NK1.1- ) or mature NK cells (mNK; DX5+NK1.1+) in the bone marrow of naïve WT or Zbtb32–/– mice (n = 3 mice per genotype). (c) Percentage of NK cells expressing indicated Ly49 NK receptors in naïve WT or Zbtb32–/– mice (n = 3 mice per genotype). (d) Expression of various activating receptors on NK cells from naïve WT or Zbtb32–/– mice (n = 3 mice per genotype). All panels are representative of 2 independent experiments.
Supplementary Figure 5 Zbtb32 is required in a cell-intrinsic manner for NK cell proliferation during viral infection.
Equal numbers of WT and Zbtb32–/– Ly49H+ NK cells from WT:Zbtb32–/– mixed bone marrow chimeras were co-transferred into Ly49H-deficient hosts. The percentage of transferred WT (CD45.1+) and Zbtb32–/– (CD45.2+) Ly49H+ NK cells in the spleen on days 8 and 24 p.i. with MCMV. Day 0 panel shows the percentage of each population in the spleens of the chimeric donors prior to transfer. Representative data from n = 3 mice per time point.
Supplementary Figure 6 Zbtb32–/–NK cells upregulate pSTATs, T-bet and GATA-3 but down-regulate Bcl-6, after MCMV infection.
(a) Splenic WT and Zbtb32–/– NK cells were co-transferred into Ly49H-deficient mice. Expression of intracellular phosphorylated STAT1, STAT3, STAT4, T-bet, and GATA-3 on transferred Ly49H+ NK cells in the spleen on day 3 p.i. with MCMV (n = 3 animals). Shaded histograms show expression in uninfected controls (n = 1 animal). Representative of 2 – 3 independent experiments. (b) Bcl6 mRNA abundance in splenic WT Ly49H+ NK cells sorted on indicated days p.i. with MCMV (n = 3 per time point), shown as fold expression relative to day 0.
Supplementary Figure 7 Zbtb32 regulates the proliferative burst of Ly49H-activated NK cells during MCMV infection.
Zbtb32 acts downstream of the IL-12 receptor and STAT4 (in addition to other proinflammatory cytokine signals) to antagonize Blimp-1 and instruct a “proliferation permissive” state in NK cells.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, and Supplementary Tables 1 and 2 (PDF 1899 kb)
Rights and permissions
About this article
Cite this article
Beaulieu, A., Zawislak, C., Nakayama, T. et al. The transcription factor Zbtb32 controls the proliferative burst of virus-specific natural killer cells responding to infection. Nat Immunol 15, 546–553 (2014). https://doi.org/10.1038/ni.2876
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2876
This article is cited by
-
IKAROS and AIOLOS directly regulate AP-1 transcriptional complexes and are essential for NK cell development
Nature Immunology (2024)
-
Themis2 regulates natural killer cell memory function and formation
Nature Communications (2023)
-
Cardinal features of immune memory in innate lymphocytes
Nature Immunology (2023)
-
Clonal expansion of innate and adaptive lymphocytes
Nature Reviews Immunology (2020)
-
Innate lymphoid cell memory
Cellular & Molecular Immunology (2019)