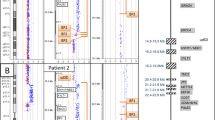
Abstract
Despite the clinical significance of balanced chromosomal abnormalities (BCAs), their characterization has largely been restricted to cytogenetic resolution. We explored the landscape of BCAs at nucleotide resolution in 273 subjects with a spectrum of congenital anomalies. Whole-genome sequencing revised 93% of karyotypes and demonstrated complexity that was cryptic to karyotyping in 21% of BCAs, highlighting the limitations of conventional cytogenetic approaches. At least 33.9% of BCAs resulted in gene disruption that likely contributed to the developmental phenotype, 5.2% were associated with pathogenic genomic imbalances, and 7.3% disrupted topologically associated domains (TADs) encompassing known syndromic loci. Remarkably, BCA breakpoints in eight subjects altered a single TAD encompassing MEF2C, a known driver of 5q14.3 microdeletion syndrome, resulting in decreased MEF2C expression. We propose that sequence-level resolution dramatically improves prediction of clinical outcomes for balanced rearrangements and provides insight into new pathogenic mechanisms, such as altered regulation due to changes in chromosome topology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jacobs, P.A., Melville, M., Ratcliffe, S., Keay, A.J. & Syme, J. A cytogenetic survey of 11,680 newborn infants. Ann. Hum. Genet. 37, 359–376 (1974).
Nielsen, J. & Wohlert, M. Chromosome abnormalities found among 34,910 newborn children: results from a 13-year incidence study in Arhus, Denmark. Hum. Genet. 87, 81–83 (1991).
Ravel, C., Berthaut, I., Bresson, J.L. & Siffroi, J.P. Prevalence of chromosomal abnormalities in phenotypically normal and fertile adult males: large-scale survey of over 10,000 sperm donor karyotypes. Hum. Reprod. 21, 1484–1489 (2006).
Funderburk, S.J., Spence, M.A. & Sparkes, R.S. Mental retardation associated with “balanced” chromosome rearrangements. Am. J. Hum. Genet. 29, 136–141 (1977).
Marshall, C.R. et al. Structural variation of chromosomes in autism spectrum disorder. Am. J. Hum. Genet. 82, 477–488 (2008).
McKusick, V.A. & Amberger, J.S. The morbid anatomy of the human genome: chromosomal location of mutations causing disease. J. Med. Genet. 30, 1–26 (1993).
Talkowski, M.E. et al. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell 149, 525–537 (2012).
Weischenfeldt, J., Symmons, O., Spitz, F. & Korbel, J.O. Phenotypic impact of genomic structural variation: insights from and for human disease. Nat. Rev. Genet. 14, 125–138 (2013).
Warburton, D. Current techniques in chromosome analysis. Pediatr. Clin. North Am. 27, 753–769 (1980).
Talkowski, M.E. et al. Next-generation sequencing strategies enable routine detection of balanced chromosome rearrangements for clinical diagnostics and genetic research. Am. J. Hum. Genet. 88, 469–481 (2011).
Talkowski, M.E. et al. Clinical diagnosis by whole-genome sequencing of a prenatal sample. N. Engl. J. Med. 367, 2226–2232 (2012).
Schluth-Bolard, C. et al. Breakpoint mapping by next generation sequencing reveals causative gene disruption in patients carrying apparently balanced chromosome rearrangements with intellectual deficiency and/or congenital malformations. J. Med. Genet. 50, 144–150 (2013).
Utami, K.H. et al. Detection of chromosomal breakpoints in patients with developmental delay and speech disorders. PLoS One 9, e90852 (2014).
Vergult, S. et al. Mate pair sequencing for the detection of chromosomal aberrations in patients with intellectual disability and congenital malformations. Eur. J. Hum. Genet. 22, 652–659 (2014).
Tabet, A.C. et al. Complex nature of apparently balanced chromosomal rearrangements in patients with autism spectrum disorder. Mol. Autism 6, 19 (2015).
Jin, F. et al. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 503, 290–294 (2013).
Rao, S.S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Köhler, S. et al. The Human Phenotype Ontology project: linking molecular biology and disease through phenotype data. Nucleic Acids Res. 42, D966–D974 (2014).
Meyerson, M. & Pellman, D. Cancer genomes evolve by pulverizing single chromosomes. Cell 144, 9–10 (2011).
Stephens, P.J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Kloosterman, W.P. et al. Chromothripsis as a mechanism driving complex de novo structural rearrangements in the germline. Hum. Mol. Genet. 20, 1916–1924 (2011).
Chiang, C. et al. Complex reorganization and predominant non-homologous repair following chromosomal breakage in karyotypically balanced germline rearrangements and transgenic integration. Nat. Genet. 44, 390–397 (2012).
Baca, S.C. et al. Punctuated evolution of prostate cancer genomes. Cell 153, 666–677 (2013).
De Gregori, M. et al. Cryptic deletions are a common finding in “balanced” reciprocal and complex chromosome rearrangements: a study of 59 patients. J. Med. Genet. 44, 750–762 (2007).
Zhang, F. et al. The DNA replication FoSTeS/MMBIR mechanism can generate genomic, genic and exonic complex rearrangements in humans. Nat. Genet. 41, 849–853 (2009).
Abyzov, A. et al. Analysis of deletion breakpoints from 1,092 humans reveals details of mutation mechanisms. Nat. Commun. 6, 7256 (2015).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Petrovski, S., Wang, Q., Heinzen, E.L., Allen, A.S. & Goldstein, D.B. Genic intolerance to functional variation and the interpretation of personal genomes. PLoS Genet. 9, e1003709 (2013).
Samocha, K.E. et al. A framework for the interpretation of de novo mutation in human disease. Nat. Genet. 46, 944–950 (2014).
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Berg, J.S. et al. An informatics approach to analyzing the incidentalome. Genet. Med. 15, 36–44 (2013).
Darnell, J.C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011).
Ascano, M. Jr. et al. FMRP targets distinct mRNA sequence elements to regulate protein expression. Nature 492, 382–386 (2012).
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012).
O'Roak, B.J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012).
Sanders, S.J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012).
De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014).
Cotney, J. et al. The autism-associated chromatin modifier CHD8 regulates other autism risk genes during human neurodevelopment. Nat. Commun. 6, 6404 (2015).
Sugathan, A. et al. CHD8 regulates neurodevelopmental pathways associated with autism spectrum disorder in neural progenitors. Proc. Natl. Acad. Sci. USA 111, E4468–E4477 (2014).
Hawrylycz, M.J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012).
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014).
Purcell, S.M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Landrum, M.J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D1, D862–D868 (2016).
Kleefstra, T. et al. Loss-of-function mutations in euchromatin histone methyl transferase 1 (EHMT1) cause the 9q34 subtelomeric deletion syndrome. Am. J. Hum. Genet. 79, 370–377 (2006).
Lu, W. et al. NFIA haploinsufficiency is associated with a CNS malformation syndrome and urinary tract defects. PLoS Genet. 3, e80 (2007).
Rosenfeld, J.A. et al. Small deletions of SATB2 cause some of the clinical features of the 2q33.1 microdeletion syndrome. PLoS One 4, e6568 (2009).
Talkowski, M.E. et al. Assessment of 2q23.1 microdeletion syndrome implicates MBD5 as a single causal locus of intellectual disability, epilepsy, and autism spectrum disorder. Am. J. Hum. Genet. 89, 551–563 (2011).
Rasmussen, M.B. et al. Neurodevelopmental disorders associated with dosage imbalance of ZBTB20 correlate with the morbidity spectrum of ZBTB20 candidate target genes. J. Med. Genet. 51, 605–613 (2014).
DeSanto, C. et al. WAC loss-of-function mutations cause a recognisable syndrome characterised by dysmorphic features, developmental delay and hypotonia and recapitulate 10p11.23 microdeletion syndrome. J. Med. Genet. 52, 754–761 (2015).
Turner, T.N. et al. Loss of δ-catenin function in severe autism. Nature 520, 51–56 (2015).
Xia, F. et al. De novo truncating mutations in AHDC1 in individuals with syndromic expressive language delay, hypotonia, and sleep apnea. Am. J. Hum. Genet. 94, 784–789 (2014).
Splawski, I. et al. Severe arrhythmia disorder caused by cardiac L-type calcium channel mutations. Proc. Natl. Acad. Sci. USA 102, 8089–8096, discussion 8086–8088 (2005).
Petrovski, S. et al. Germline de novo mutations in GNB1 cause severe neurodevelopmental disability, hypotonia, and seizures. Am. J. Hum. Genet. 98, 1001–1010 (2016).
Floris, C. et al. Two patients with balanced translocations and autistic disorder: CSMD3 as a candidate gene for autism found in their common 8q23 breakpoint area. Eur. J. Hum. Genet. 16, 696–704 (2008).
Cardoso, C. et al. Periventricular heterotopia, mental retardation, and epilepsy associated with 5q14.3-q15 deletion. Neurology 72, 784–792 (2009).
Engels, H. et al. A novel microdeletion syndrome involving 5q14.3-q15: clinical and molecular cytogenetic characterization of three patients. Eur. J. Hum. Genet. 17, 1592–1599 (2009).
Le Meur, N. et al. MEF2C haploinsufficiency caused by either microdeletion of the 5q14.3 region or mutation is responsible for severe mental retardation with stereotypic movements, epilepsy and/or cerebral malformations. J. Med. Genet. 47, 22–29 (2010).
Zweier, M. et al. Mutations in MEF2C from the 5q14.3q15 microdeletion syndrome region are a frequent cause of severe mental retardation and diminish MECP2 and CDKL5 expression. Hum. Mutat. 31, 722–733 (2010).
Saitsu, H. et al. De novo 5q14.3 translocation 121.5-kb upstream of MEF2C in a patient with severe intellectual disability and early-onset epileptic encephalopathy. Am. J. Med. Genet. A. 155A, 2879–2884 (2011).
Zweier, M. & Rauch, A. The MEF2C-related and 5q14.3q15 microdeletion syndrome. Mol. Syndromol. 2, 164–170 (2012).
Dixon, J.R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Lupiáñez, D.G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene–enhancer interactions. Cell 161, 1012–1025 (2015).
Lupiáñez, D.G., Spielmann, M. & Mundlos, S. Breaking TADs: how alterations of chromatin domains result in disease. Trends Genet. 32, 225–237 (2016).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Mencarelli, M.A. et al. 14q12 microdeletion syndrome and congenital variant of Rett syndrome. Eur. J. Med. Genet. 52, 148–152 (2009).
Ellaway, C.J. et al. 14q12 microdeletions excluding FOXG1 give rise to a congenital variant Rett syndrome–like phenotype. Eur. J. Hum. Genet. 21, 522–527 (2013).
Takagi, M. et al. A 2.0 Mb microdeletion in proximal chromosome 14q12, involving regulatory elements of FOXG1, with the coding region of FOXG1 being unaffected, results in severe developmental delay, microcephaly, and hypoplasia of the corpus callosum. Eur. J. Med. Genet. 56, 526–528 (2013).
Perche, O. et al. Dysregulation of FOXG1 pathway in a 14q12 microdeletion case. Am. J. Med. Genet. A. 161A, 3072–3077 (2013).
Ibn-Salem, J. et al. Deletions of chromosomal regulatory boundaries are associated with congenital disease. Genome Biol. 15, 423 (2014).
Deng, Y., Gao, L., Wang, B. & Guo, X. HPOSim: an R package for phenotypic similarity measure and enrichment analysis based on the human phenotype ontology. PLoS One 10, e0115692 (2015).
Brunetti-Pierri, N. et al. Duplications of FOXG1 in 14q12 are associated with developmental epilepsy, mental retardation, and severe speech impairment. Eur. J. Hum. Genet. 19, 102–107 (2011).
McDermott, S.M. et al. Drosophila Syncrip modulates the expression of mRNAs encoding key synaptic proteins required for morphology at the neuromuscular junction. RNA 20, 1593–1606 (2014).
Warburton, D. De novo balanced chromosome rearrangements and extra marker chromosomes identified at prenatal diagnosis: clinical significance and distribution of breakpoints. Am. J. Hum. Genet. 49, 995–1013 (1991).
Brand, H. et al. Cryptic and complex chromosomal aberrations in early-onset neuropsychiatric disorders. Am. J. Hum. Genet. 95, 454–461 (2014).
Chaisson, M.J. et al. Resolving the complexity of the human genome using single-molecule sequencing. Nature 517, 608–611 (2015).
Huddleston, J. et al. Reconstructing complex regions of genomes using long-read sequencing technology. Genome Res. 24, 688–696 (2014).
Lettice, L.A. et al. Enhancer-adoption as a mechanism of human developmental disease. Hum. Mutat. 32, 1492–1499 (2011).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Hanscom, C. & Talkowski, M. Design of large-insert jumping libraries for structural variant detection using Illumina sequencing. Curr. Protoc. Hum. Genet. 80, 7.22.21–7.22.29 (2014).
Higgins, A.W. et al. Characterization of apparently balanced chromosomal rearrangements from the developmental genome anatomy project. Am. J. Hum. Genet. 82, 712–722 (2008).
Köhler, S. et al. Clinical diagnostics in human genetics with semantic similarity searches in ontologies. Am. J. Hum. Genet. 85, 457–464 (2009).
Brand, H. et al. Paired-duplication signatures mark cryptic inversions and other complex structural variation. Am. J. Hum. Genet. 97, 170–176 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Tarasov, A., Vilella, A.J., Cuppen, E., Nijman, I.J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
North, B.V., Curtis, D. & Sham, P.C. A note on the calculation of empirical P values from Monte Carlo procedures. Am. J. Hum. Genet. 71, 439–441 (2002).
Durand, N.C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Acknowledgements
We are deeply grateful for the seminal work led by our co-author, Dorothy Warburton, who passed away during review of this manuscript. Dr. Warburton was a pioneer in cytogenetic research and a close colleague, mentor, and friend to so many in the cytogenetics community.
We wish to thank all subjects and families who have been enrolled in this study, as well as the countless genetic counselors and clinical geneticists who contributed to the ascertainment of subjects. This study was supported by the National Institutes of Health (grant GM061354 to M.E.T., J.F.G., C.C.M., and E.L.; grants MH095867 and HD081256 to M.E.T.), the March of Dimes (6-FY15-255 to M.E.T.), the European Molecular Biology Organization and the Marie Curie Actions of the European Commission (fellowship EMBO ALTF-183-2015 to C.R.), the Bettencourt-Schueller Foundation (young investigator award to C.R.), the Philippe Foundation (award to C.R.), the Harvard Medical School–Portugal Program in Translational and Clinical Research and Health Information (Fundação para a Ciência e a Tecnologia, HMSP-ICT/0016/2013 to C.C.M. and D.D.), the National Science Foundation (NSF Graduate Research Fellowship DGE1144152 to S.L.P.S.), the Fund for Scientific Research–Flanders (B.C. and S.V. are an FWO senior clinical investigator and an FWO postdoctoral researcher, respectively), Clinical Medicine Science and Technology Projects of Jiangsu Province (grant BL2013019 to Haibo Li and Hong Li), the Suzhou Key Medical Center (grant Szzx201505 to Haibo Li and Hong Li), and the Royal Society of New Zealand (Rutherford Discovery Fellowship to J.C.J.). This study was also supported by the Desmond and Ann Heathwood MGH Research Scholars award to M.E.T.
Author information
Authors and Affiliations
Contributions
M.E.T., J.F.G., C.C.M., E.C.T., J.C.H., W.P.K., N.d.L., and H.G.B. designed the study. C.R., H.B., R.L.C., V.P., I.B., C.C., J.T.G., M.R.S., M.J.v.R., and W.P.K. performed computational analyses. C.H., C.M.S., R.A., M.-A. Anderson, C.A., E.C., B.B.C., J.K., W.L., P.M., L.M., T. Mason, D.P., J.R., M.J.W., and A.W. performed cellular, molecular, or genomic experiments. T.K., E. Mitchell, J.C.H., M.-A. Abbott, O.A.A.-R., E.A., S.L.A.-E., F.S.A., Y.A., K.A.-Y., J.F.A., T.B., J.A.B., E.B., E.M.H.F.B., E.H.B., C.W.B., H.T.B., B.C., K.C., H.C., T.C., D.D., M.A.D., A.D., M.D'H., B.B.A.d.V., D.L.E., H.L.F., H.F., D.R.F., P.G., D.G., T.G., M.G., B.H.G., C.G., K.W.G., A.L.G., A.H.-K., D.J.H., M.A.H., R. Hill, R. Hochstenbach, J.D.H., R.J.H., M.W.H., A.M.I., M. Irons, M. Irving, J.C.J., S.J., T.J., J.P.J., M.C.J., S.G.K., D.A.K., P.M.K., Y.L., E.L., K.L., A.V.L., Haibo Li, Hong Li, E.C.L., C.L., E.J.L., D.L., M.J.M., G. Mandrile, C.L.M., D.M.-F., M.W.M., C.J.Z.M., B.M., S. Middelkamp, L.R.M., E. Moe, S. Mohammed, T. Mononen, M.E.M., G. Moya, A.W.N., Z.O., S. Parkash, S.P.P., S. Pereira, K.P., R.E.P.A., P.J.P., G.P., S.R., L.R., W.R., D.R., I.R., F.R., P.R., S.L.P.S., R. Shaheen, R. Sparkes, E.S., B.S., J.T., J.V.T., B.W.v.B., J.v.d.K., I.v.D.B., T.v.E., C.M.v.R.-A., S.V., C.M.L.V.-T., D.P.W., S.W., M.d.l.C.A.Y.-d.V., R.T.Z., B.L., H.G.B., N.d.L., W.P.K., E.C.T., C.C.M., and J.F.G. ascertained and enrolled subjects and provided phenotypic information. C.R. and M.E.T. wrote the manuscript, which was approved by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–77 and Supplementary Note. (PDF 39849 kb)
Supplementary Tables 1–12
Supplementary Tables 1–12. (XLSX 769 kb)
Rights and permissions
About this article
Cite this article
Redin, C., Brand, H., Collins, R. et al. The genomic landscape of balanced cytogenetic abnormalities associated with human congenital anomalies. Nat Genet 49, 36–45 (2017). https://doi.org/10.1038/ng.3720
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3720
This article is cited by
-
Structural rearrangements as a recurrent pathogenic mechanism for SETBP1 haploinsufficiency
Human Genomics (2024)
-
Long-read sequencing and optical mapping generates near T2T assemblies that resolves a centromeric translocation
Scientific Reports (2024)
-
Scrambling the genome in cancer: causes and consequences of complex chromosome rearrangements
Nature Reviews Genetics (2024)
-
Evaluation of genetic risk of apparently balanced chromosomal rearrangement carriers by breakpoint characterization
Journal of Assisted Reproduction and Genetics (2024)
-
Genome sequencing as a generic diagnostic strategy for rare disease
Genome Medicine (2024)