Abstract
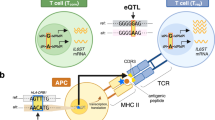
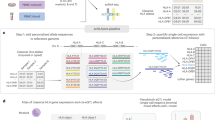
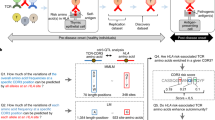
In each individual, a highly diverse T cell receptor (TCR) repertoire interacts with peptides presented by major histocompatibility complex (MHC) molecules. Despite extensive research, it remains controversial whether germline-encoded TCR–MHC contacts promote TCR–MHC specificity and, if so, whether differences exist in TCR V gene compatibilities with different MHC alleles. We applied expression quantitative trait locus (eQTL) mapping to test for associations between genetic variation and TCR V gene usage in a large human cohort. We report strong trans associations between variation in the MHC locus and TCR V gene usage. Fine-mapping of the association signals identifies specific amino acids from MHC genes that bias V gene usage, many of which contact or are spatially proximal to the TCR or peptide in the TCR–peptide–MHC complex. Hence, these MHC variants, several of which are linked to autoimmune diseases, can directly affect TCR–MHC interaction. These results provide the first examples of trans-QTL effects mediated by protein–protein interactions and are consistent with intrinsic TCR–MHC specificity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Neefjes, J., Jongsma, M.L.M., Paul, P. & Bakke, O. Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nat. Rev. Immunol. 11, 823–836 (2011).
McDevitt, H.O. & Bodmer, W.F. HL-A, immune-response genes, and disease. Lancet 1, 1269–1275 (1974).
Gutierrez-Arcelus, M., Rich, S.S. & Raychaudhuri, S. Autoimmune diseases—connecting risk alleles with molecular traits of the immune system. Nat. Rev. Genet. 17, 160–174 (2016).
Miyadera, H. & Tokunaga, K. Associations of human leukocyte antigens with autoimmune diseases: challenges in identifying the mechanism. J. Hum. Genet. 60, 697–702 (2015).
Vantourout, P. & Hayday, A. Six-of-the-best: unique contributions of γδ T cells to immunology. Nat. Rev. Immunol. 13, 88–100 (2013).
Rossjohn, J. et al. T cell antigen receptor recognition of antigen-presenting molecules. Annu. Rev. Immunol. 33, 169–200 (2015).
Rudolph, M.G., Stanfield, R.L. & Wilson, I.A. How TCRs bind MHCs, peptides, and coreceptors. Annu. Rev. Immunol. 24, 419–466 (2006).
Turner, S.J., Doherty, P.C., McCluskey, J. & Rossjohn, J. Structural determinants of T-cell receptor bias in immunity. Nat. Rev. Immunol. 6, 883–894 (2006).
Housset, D. & Malissen, B. What do TCR–pMHC crystal structures teach us about MHC restriction and alloreactivity? Trends Immunol. 24, 429–437 (2003).
Garcia, K.C. et al. A closer look at TCR germline recognition. Immunity 36, 887–888 (2012).
Klein, L., Kyewski, B., Allen, P.M. & Hogquist, K.A. Positive and negative selection of the T cell repertoire: what thymocytes see (and don't see). Nat. Rev. Immunol. 14, 377–391 (2014).
Roudier, J. Association of MHC and rheumatoid arthritis. Association of RA with HLA-DR4: the role of repertoire selection. Arthritis Res. 2, 217–220 (2000).
Robins, H.S. et al. Overlap and effective size of the human CD8+ T cell receptor repertoire. Sci. Transl. Med. 2, 47ra64 (2010).
Zvyagin, I.V. et al. Distinctive properties of identical twins' TCR repertoires revealed by high-throughput sequencing. Proc. Natl. Acad. Sci. USA 111, 5980–5985 (2014).
Gulwani-Akolkar, B. et al. Do HLA genes play a prominent role in determining T cell receptor Vα segment usage in humans? J. Immunol. 154, 3843–3851 (1995).
Miles, J.J. et al. TCRα genes direct MHC restriction in the potent human T cell response to a class I–bound viral epitope. J. Immunol. 177, 6804–6814 (2006).
Garcia, K.C. Reconciling views on T cell receptor germline bias for MHC. Trends Immunol. 33, 429–436 (2012).
Garcia, K.C., Adams, J.J., Feng, D. & Ely, L.K. The molecular basis of TCR germline bias for MHC is surprisingly simple. Nat. Immunol. 10, 143–147 (2009).
Castro, C.D., Luoma, A.M. & Adams, E.J. Coevolution of T-cell receptors with MHC and non-MHC ligands. Immunol. Rev. 267, 30–55 (2015).
Marrack, P., Scott-Browne, J.P., Dai, S., Gapin, L. & Kappler, J.W. Evolutionarily conserved amino acids that control TCR–MHC interaction. Annu. Rev. Immunol. 26, 171–203 (2008).
Scott-Browne, J.P., White, J., Kappler, J.W., Gapin, L. & Marrack, P. Germline-encoded amino acids in the αβ T-cell receptor control thymic selection. Nature 458, 1043–1046 (2009).
Van Laethem, F. et al. Lck availability during thymic selection determines the recognition specificity of the T cell repertoire. Cell 154, 1326–1341 (2013).
Van Laethem, F. et al. Deletion of CD4 and CD8 coreceptors permits generation of αβ T cells that recognize antigens independently of the MHC. Immunity 27, 735–750 (2007).
Holland, S.J. et al. The T-cell receptor is not hardwired to engage MHC ligands. Proc. Natl. Acad. Sci. USA 109, E3111–E3118 (2012).
Burrows, S.R. et al. Hard wiring of T cell receptor specificity for the major histocompatibility complex is underpinned by TCR adaptability. Proc. Natl. Acad. Sci. USA 107, 10608–10613 (2010).
Beringer, D.X. et al. T cell receptor reversed polarity recognition of a self-antigen major histocompatibility complex. Nat. Immunol. 16, 1153–1161 (2015).
Parrish, H.L., Deshpande, N.R., Vasic, J. & Kuhns, M.S. Functional evidence for TCR-intrinsic specificity for MHCII. Proc. Natl. Acad. Sci. USA 113, 3000–3005 (2016).
Rockman, M.V. & Kruglyak, L. Genetics of global gene expression. Nat. Rev. Genet. 7, 862–872 (2006).
Battle, A. et al. Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals. Genome Res. 24, 14–24 (2014).
Sottini, A., Imberti, L., Fiordalisi, G. & Primi, D. Use of variable human Vδ genes to create functional T cell receptor α chain transcripts. Eur. J. Immunol. 21, 2455–2459 (1991).
Jia, X. et al. Imputing amino acid polymorphisms in human leukocyte antigens. PLoS One 8, e64683 (2013).
Sinclair, C., Bains, I., Yates, A.J. & Seddon, B. Asymmetric thymocyte death underlies the CD4:CD8 T-cell ratio in the adaptive immune system. Proc. Natl. Acad. Sci. USA 110, E2905–E2914 (2013).
Yang, J., Lee, S.H., Goddard, M.E. & Visscher, P.M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Kichaev, G. et al. Integrating functional data to prioritize causal variants in statistical fine-mapping studies. PLoS Genet. 10, e1004722 (2014).
Wallace, C. et al. Dissection of a complex disease susceptibility region using a Bayesian stochastic search approach to fine mapping. PLoS Genet. 11, e1005272 (2015).
Wellcome Trust Case Control Consortium. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat. Genet. 44, 1294–1301 (2012).
Servin, B. & Stephens, M. Imputation-based analysis of association studies: candidate regions and quantitative traits. PLoS Genet. 3, e114 (2007).
Raychaudhuri, S. et al. Five amino acids in three HLA proteins explain most of the association between MHC and seropositive rheumatoid arthritis. Nat. Genet. 44, 291–296 (2012).
Hu, X. et al. Additive and interaction effects at three amino acid positions in HLA-DQ and HLA-DR molecules drive type 1 diabetes risk. Nat. Genet. 47, 898–905 (2015).
Patsopoulos, N.A. et al. Fine-mapping the genetic association of the major histocompatibility complex in multiple sclerosis: HLA and non-HLA effects. PLoS Genet. 9, e1003926 (2013).
Messaoudi, I., Guevara Patiño, J.A., Dyall, R., LeMaoult, J. & Nikolich-Zugich, J. Direct link between MHC polymorphism, T cell avidity, and diversity in immune defense. Science 298, 1797–1800 (2002).
Price, D.A. et al. Public clonotype usage identifies protective Gag-specific CD8+ T cell responses in SIV infection. J. Exp. Med. 206, 923–936 (2009).
Luz, J.G. et al. Structural comparison of allogeneic and syngeneic T cell receptor–peptide–major histocompatibility complex complexes: a buried alloreactive mutation subtly alters peptide presentation substantially increasing Vβ interactions. J. Exp. Med. 195, 1175–1186 (2002).
Murray, J.S. An old Twist in HLA-A: CDR3α hook up at an R65-joint. Front. Immunol. 6, 268 (2015).
Levin, A.M. et al. Exploiting a natural conformational switch to engineer an interleukin-2 'superkine'. Nature 484, 529–533 (2012).
Jerne, N.K. The somatic generation of immune recognition. Eur. J. Immunol. 1, 1–9 (1971).
Dai, S. et al. Crossreactive T cells spotlight the germline rules for αβ T cell-receptor interactions with MHC molecules. Immunity 28, 324–334 (2008).
Adams, J.J. et al. Structural interplay between germline interactions and adaptive recognition determines the bandwidth of TCR–peptide–MHC cross-reactivity. Nat. Immunol. 17, 87–94 (2016).
Feng, D., Bond, C.J., Ely, L.K., Maynard, J. & Garcia, K.C. Structural evidence for a germline-encoded T cell receptor–major histocompatibility complex interaction 'codon'. Nat. Immunol. 8, 975–983 (2007).
Blevins, S.J. et al. How structural adaptability exists alongside HLA-A2 bias in the human αβ TCR repertoire. Proc. Natl. Acad. Sci. USA 113, E1276–E1285 (2016).
Sim, B.C., Zerva, L., Greene, M.I. & Gascoigne, N.R. Control of MHC restriction by TCR Vα CDR1 and CDR2. Science 273, 963–966 (1996).
Ferreira, M.A.R. et al. Quantitative trait loci for CD4:CD8 lymphocyte ratio are associated with risk of type 1 diabetes and HIV-1 immune control. Am. J. Hum. Genet. 86, 88–92 (2010).
Klarenbeek, P.L. et al. Somatic variation of T-cell receptor genes strongly associate with HLA class restriction. PLoS One 10, e0140815 (2015).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359 (2012).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Anders, S., Pyl, P.T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
de Bakker, P.I.W. et al. A high-resolution HLA and SNP haplotype map for disease association studies in the extended human MHC. Nat. Genet. 38, 1166–1172 (2006).
Delaneau, O. & Marchini, J. Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel. Nat. Commun. 5, 3934 (2014).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Visscher, P.M., Yang, J. & Goddard, M.E. A commentary on 'common SNPs explain a large proportion of the heritability for human height' by Yang et al. (2010). Twin Res. Hum. Genet. 13, 517–524 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Mitchell, T.J. & Beauchamp, J.J. Bayesian variable selection in linear regression. J. Am. Stat. 83, 1023–1032 (1988).
Ishwaran, H. & Rao, J.S. Spike and slab gene selection for multigroup microarray data. J. Am. Stat. 100, 764–780 (2005).
Geman, S. & Geman, D. Stochastic relaxation, gibbs distributions, and the Bayesian restoration of images. IEEE Trans. Pattern Anal. Mach. Intell. 6, 721–741 (1984).
Lefranc, M.-P. et al. IMGT®, the international ImMunoGeneTics information system® 25 years on. Nucleic Acids Res. 43, D413–D422 (2015).
Berman, H.M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015).
Acknowledgements
We thank D. Golan, D. Knowles, A. Fu, M. Birnbaum, M. Gee, J. Mendoza, A. Bhaskar and T. Raj for helpful discussions and the anonymous referees for valuable comments. This work was supported by NIH grants HG0070736, 1R01GM097171-01A1, RO1AI03867 and U19AI057229, the Howard Hughes Medical Institute, the EMBO Long-Term Fellowship and a National Science Foundation Graduate Research Fellowship.
Author information
Authors and Affiliations
Contributions
E.S., J.K.P. and K.C.G. conceived the project. E.S. performed genetic analyses with input from A.B., H.B.F. and J.K.P. E.S. and L.V.S. performed structural analyses with input from K.C.G. E.S., L.V.S., K.C.G. and J.K.P. wrote the manuscript. The work was supervised by K.C.G. and J.K.P. All authors reviewed, revised and provided feedback on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–32 and Supplementary Table 1 (PDF 17162 kb)
Supplementary Table 2
TCR V genes expression. (XLSX 930 kb)
Supplementary Table 3
Ig V-genes expression. (XLSX 1070 kb)
Supplementary Table 4
TCR and Ig V genes association with short-range genetic variation and genetic variation in the MHC locus. (XLSX 88 kb)
Supplementary Table 5
Most significance association between expression of each TCR and Ig V gene and genotyped SNPs in the MHC locus. (XLSX 76 kb)
Supplementary Table 6
Nucleotide and amino acid variants in the MHC locus that are independently associated with single TCR Vα or Vβ gene expression. (XLSX 70 kb)
Supplementary Table 7
Expression variation of TCR Vα and Vβ genes explained by indepedent associations with SNP and amino acid variation in the MHC locus. (XLSX 54 kb)
Supplementary Table 8
Expression variation of TCR Vα and Vβ genes explained by indepedent associations with four-digit haplotypes for classical MHC genes. (XLSX 12 kb)
Supplementary Table 9
Probabilities that classical MHC gene amino acid positions influences expression of any TCR Vα gene. (XLSX 62 kb)
Supplementary Table 10
Probabilities that classical MHC gene amino acid positions influences expression of any TCR Vβ gene. (XLSX 61 kb)
Supplementary Table 11
A list of PDB accession codes, MHC alleles and TCR Vα gene used in the analysis of TCR–pMHC complexes. (XLSX 58 kb)
Rights and permissions
About this article
Cite this article
Sharon, E., Sibener, L., Battle, A. et al. Genetic variation in MHC proteins is associated with T cell receptor expression biases. Nat Genet 48, 995–1002 (2016). https://doi.org/10.1038/ng.3625
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3625
This article is cited by
-
The immunopathogenesis of narcolepsy type 1
Nature Reviews Immunology (2024)
-
MHC II immunogenicity shapes the neoepitope landscape in human tumors
Nature Genetics (2023)
-
Narcolepsy risk loci outline role of T cell autoimmunity and infectious triggers in narcolepsy
Nature Communications (2023)
-
Validation and promise of a TCR mimic antibody for cancer immunotherapy of hepatocellular carcinoma
Scientific Reports (2022)
-
Genetic and environmental determinants of human TCR repertoire diversity
Immunity & Ageing (2020)