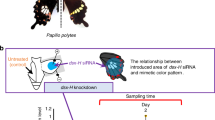
Abstract
In Batesian mimicry, animals avoid predation by resembling distasteful models. In the swallowtail butterfly Papilio polytes, only mimetic-form females resemble the unpalatable butterfly Pachliopta aristolochiae. A recent report showed that a single gene, doublesex (dsx), controls this mimicry1; however, the detailed molecular mechanisms remain unclear. Here we determined two whole-genome sequences of P. polytes and a related species, Papilio xuthus, identifying a single ∼130-kb autosomal inversion, including dsx, between mimetic (H-type) and non-mimetic (h-type) chromosomes in P. polytes. This inversion is associated with the mimicry-related locus H, as identified by linkage mapping. Knockdown experiments demonstrated that female-specific dsx isoforms expressed from the inverted H allele (dsx(H)) induce mimetic coloration patterns and simultaneously repress non-mimetic patterns. In contrast, dsx(h) does not alter mimetic patterns. We propose that dsx(H) switches the coloration of predetermined wing patterns and that female-limited polymorphism is tightly maintained by chromosomal inversion.
Similar content being viewed by others
Main
P. polytes is known to exhibit female-limited Batesian mimicry. The females have two forms: a non-mimetic form (cyrus), with wing patterns identical to those of monomorphic males, and a mimetic form (polytes) resembling the toxic model P. aristolochiae (Fig. 1a). This polymorphism is controlled by a single autosomal locus H, and the mimetic phenotype (genotype HH or Hh) is dominant2. There are two models of the underlying gene encoded in the H locus: a conceptual ‘supergene’ consisting of a series of neighboring genes tightly linked to one another3, or a regulatory gene controlling unlinked downstream genes that affect the color pattern4. It has been demonstrated that a supergene is created and fixed by chromosomal rearrangements in Heliconius numata, a species exhibiting Müllerian mimicry5. In addition, a recent study on P. polytes has shown that the mimetic phenotype is controlled by dsx, which was suggested to be fixed by chromosomal inversion1. However, the chromosomal structure and detailed molecular mechanisms involved in the female-limited polymorphic mimicry in P. polytes remain obscure.
(a) P. polytes and its model species P. aristolochiae. Scale bars, 1 cm. (b) Initial H locus screening of a small number of BC1 individuals (ten mimetic and ten non-mimetic). The mimetic phenotype is associated with prospero (0.0 cM) and survivin (4.3 cM) on chromosome LG25. No markers were characterized on LG1, LG27 and LG28. (c) Analysis of marker recombination rates led to the identification of a chromosomal interval completely associated with the mimicry wing pattern (H_800 kb). Numbers beside the markers indicate scaffold positions in the present P. polytes genome assembly, H_locus_nonmimetic_h_scaffold. Recombination rates were calculated using mimetic and non-mimetic BC1 females (n = 51). (d) An association study was performed on non-mimetic (n = 28) and mimetic (n = 26) wild-caught adult females. Three markers within the inversion breakpoints (gray box) showed strong association between SNPs and the mimicry wing pattern. The blue numbers indicate the number of SNPs corresponding to each marker. The position of dsx is indicated. No other association was found in the 1-Mb flanking regions or on a different chromosome containing ras.
Kunte et al. identified the H locus as corresponding to dsx1; in this study, we performed further crosses that confirmed this result using non-mimetic individuals of P. polytes and mimetic individuals of the subspecies P. p. alphenor (also known as Papilio alphenor). Using DNA fragment-length polymorphisms, we mapped the H locus in P. polytes to a region spanning 800 kb on chromosome 25 (Fig. 1b,c). In addition, analysis of SNPs in the 800-kb region using 54 P. polytes females captured on the Ryukyu Islands, Japan, showed that 8 SNPs in dsx were associated with the mimetic phenotype (P < 1 × 10−10; Fig. 1d).
Following this analysis, we determined the whole-genome sequences of P. polytes (Hh mimetic female) and the related species P. xuthus for comparison using the HiSeq 2000 and HiSeq 2500 systems (Supplementary Table 1). The quality of the assembled scaffolds for both species was high enough for these to be used as reference genomes (scaffold N50 values: P. polytes, 3.7 Mb; P. xuthus, 6.2 Mb; Supplementary Table 2). The total genome sizes for P. polytes and P. xuthus were 227 Mb and 244 Mb (Supplementary Table 2), encoding 12,244 and 13,102 predicted protein-coding genes, respectively (Supplementary Table 3). The number of ortholog groups among 5 Lepidoptera species is shown in Figure 2a, and a phylogenetic tree of 2,077 of these orthologous genes established the genetic relationships among the 5 species (Supplementary Fig. 1a). During de novo genome assembly using Platanus6, we found some long regions with high allelic divergence in P. polytes but not in P. xuthus; these regions were further selected on the basis of having a coverage depth ≤350, which was approximately half of the peak for homozygous loci at 600 (Fig. 2b, bottom). After clustering overlapping windows, we found 15 highly diverse (identity ≤90%) and long (≥100-kb) heterozygous regions; 14 mapped to the heterogametic sex chromosome 1 (ZW) and 1 mapped to chromosome 25 near dsx (denoted ‘hetero_130 kb’) (Fig. 2b), possibly reflecting the differences between chromosomes with the h and H alleles. In comparison with the rate of single-nucleotide variation in the whole genome (0.252%), the rate of match/mismatch (mismatch/(match + mismatch)) was 23.9% in the hetero_130 kb region, indicating that the region is extremely diverse.
(a) Venn diagram of the protein ortholog groups among five lepidopteran species. (b) Distribution of coverage depth (bottom) and the chromosomal locations of long heterozygous regions in P. polytes (n = 15; red dots and rectangles). On the basis of the coverage depths of 50-kb windows, we classified regions with coverage <350 as highly divergent. Scaffolds with lengths ≥100 kb were mapped to the chromosomes of B. mori. Fourteen regions were derived from the sex chromosome (ZW) and one mapped around dsx on chromosome 25.
By fosmid sequencing and genome walking, we obtained two large scaffolds (2.3 Mb), including different hetero_130 kb regions: 122.7 kb for the h-type chromosome and 136.7 kb for the H-type chromosome (Supplementary Fig. 2). Overlapping fosmid clones completely covered the hetero_130 kb region for the h locus and most of the H locus (including left breakpoints) (Supplementary Fig. 2a,b).In addition, we confirmed the sequences around all the breakpoints by PCR (Supplementary Fig. 2c,d) and by mate-pair libraries, sufficiently covering all regions (Supplementary Fig. 2a,b). These data verified two types of large regions with high allelic divergence around dsx in P. polytes. The hetero_130 kb region on the H-type chromosome was the most remarkable, exhibiting an inverted orientation relative to the h-type chromosome (Fig. 3a,b and Supplementary Figs. 3 and 4). Most of the hetero_130 kb region was occupied by dsx, and the intron-exon structures of the chromosomes with the h and H loci were reversed, suggesting that a simple inversion occurred near both ends of dsx. Sequence comparison of the hetero_130 kb regions corresponding to the H and h loci showed a low level of homology, which was not only direct but also in the reverse orientation (Fig. 3c), whereas some scattered regions, including exons for dsx, were highly conserved. In a comparison of the hetero_130 kb region with a corresponding region in P. xuthus, the homology between the h locus and the P. xuthus region was lower than that between the h and H loci (Fig. 3c), suggesting that the chromosomal inversion occurred after the divergence of P. polytes and P. xuthus. We found that all sequenced lepidopteran genomes, except for that of Danaus plexippus, had similarly oriented synteny with the h-type region in P. polytes (Fig. 3d). This finding suggests that the H-type chromosome originated from the h-type chromosome.
(a) Detailed gene structures for the h and H alleles. Putative breakpoints are indicated by red dotted lines. (b) Sequence alignment near both breakpoints in four individuals. Asterisks indicate the positions of nucleotides shared by the h and H alleles. (c) Graphical overview of the homology between the heterozygous regions of the h and H loci. Top, h versus H; middle, h versus the reverse complement of H; bottom, h versus a corresponding region in P. xuthus. Exons for each gene are shown in blue. Introns and intergenic regions are shown in pink. (d) Synteny around dsx on the H allele in P. polytes and other insects.
To estimate the exact breakpoints of the inversion, we compared the boundary regions of the hetero_130 kb regions for the H and h loci using DNA from four butterflies captured on Ishigaki Island and the main Okinawa Islands in Japan and determined putative breakpoints by detecting a sharp decline in sequence conservation (Fig. 3b). In comparison to dsx(h), dsx(H) was longer in introns 2, 4, 5 and 6 and in exon 6 (Fig. 3a and Supplementary Figs. 3 and 4). These structural differences imply that several insertion and deletion events might have occurred in the hetero_130 kb region on the H-type chromosome after the inversion, being maintained by the repression of recombination between the two chromosome types. In addition, we determined the sequences of dsx(h) and dsx(H) in P. p. alphenor on the basis of the Illumina genomic reads reported in the previous study (Sequence Read Archive (SRA), SRP035394)1 (Online Methods) and performed phylogenetic analysis using these sequences. Analysis of the phylogenetic trees suggested that the divergence of dsx(h) and dsx(H) occurred after the divergence of P. polytes and P. xuthus 40 million years ago7 and before the separation of subspecies for P. p. alphenor and P. p. polytes (Supplementary Fig. 1b,c). The long branches for dsx(H) in both subspecies indicate that dsx(H) has a high evolutionary rate and might have evolved a new function under positive selective pressure.
Gene annotation for both the h- and H-type chromosomes identified three independent transcripts near the left breakpoints, ubiquitously expressed transcript (UXT; transcriptional regulator)8, unknown-3-exons (U3X; long noncoding RNA emerging in the H locus) and unknown transcripts downstream of prospero (Supplementary Figs. 3 and 4), that had higher expression in the wings of mimetic females (HH or Hh) than in those of non-mimetic females (hh) (Fig. 4a,b and Supplementary Fig. 5). The structures of the 5′ UTR and transcriptional start site for UXT were altered by an inversion event between the H and h loci, whereas the ORF remained unchanged (Fig. 3b and Supplementary Fig. 3c). These results suggest that inversion on the H-type chromosome affects the regulation of neighboring genes, even for those outside the inverted region.
Expression levels of genes in female hindwings at stages P1–P2 (n = 4 for mimetic; n = 3 for non-mimetic), P4–P5 (n = 6 for mimetic; n = 3 for non-mimetic) and P10.5 (n = 3 for mimetic; n = 3 for non-mimetic), as analyzed by qRT-PCR. (a) UXT. (b) U3X. (c) F1 isoform of dsx. (d) F2 isoform of dsx. (e) F3 isoform of dsx. (f) dsx from H (dsx(H)). (g) dsx from h (dsx(h)). The expression level of RpL3 was used as the internal control. (h) Allele-specific expression of a female-limited isoform of dsx in each Hh individual at the P2 stage, as estimated by RNA-seq. The FPKM value represents fragments per kilobase of transcript per million mapped reads. Filled and open bars indicate mimetic and non-mimetic samples, respectively. Bars colored blue, red and gray represent samples from white, red and black regions (or corresponding regions) of mimetic and non-mimetic pupal hindwings, respectively (schematic at top left). Data are represented as means ± s.d.; *P < 0.05 for Student's t test, Welch's t test, two-way factorial ANOVA and Tukey's honestly significant difference (HSD) test (Online Methods). All n values indicate the number of biological replicates.
RNA sequencing (RNA-seq) assembly from mimetic (HH) and non-mimetic (hh) females showed three types of female-specific dsx isoforms (F1, F2 and F3) in the wings, confirming the results of a previous study1. Although the numbers of nonsynonymous substitutions in the three Dsx isoforms (F1, 16; F2, 14; F3, 15) between the H and h alleles (Supplementary Fig. 6) were different from that found in the previous study (21–22 substitutions)1, we obtained similar results because most substitutions occurred around the DNA-binding motif and dimerization domain1,9. The sequence differences observed when comparing the results in our study (P. p. polytes) with those obtained by Kunte et al.1 (P. p. alphenor) might be due to intraspecific variations. Comparison of the dsx sequences among Lepidoptera showed that only five encoded amino acids were specifically changed in P. polytes dsx(H) (Supplementary Fig. 7), implying that these might have central roles in mimicry.
Kunte et al.1 demonstrated that two isoforms of dsx appeared to be similarly expressed in the wings of mimetic and non-mimetic females at early (5th instar larva to first pupal (P1)) and middle (P1–P3) stages; these variants were upregulated only in mimetic females (HH) at late stages (P5–P12), suggesting an increase in dsx(H) expression during the late pupal stage in mimetic females. In contrast, our analyses suggested that the expression levels of three female-specific isoforms of dsx did not exhibit major differences between mimetic (Hh or HH) and non-mimetic (hh) wings at early to middle pupal stages (P1–P2 and P4–P5) and in each color region even at the late pupal stage (P10.5) (Fig. 4c–e). Using quantitative RT-PCR (qRT-PCR) with specific primers, we further confirmed that dsx(H) in mimetic females (Hh or HH) was highly expressed in the early pupal stages (Fig. 4f, P1–P2), whereas the expression levels of dsx(h) increased at later pupal stages (Fig. 4g, P10.5). Kunte et al. showed that the expression level of dsx(H) in mimetic wings during stages P1–P3 was rather lower than that during later pupal stages1, suggesting a difference in dsx expression profiles between the two results.In addition, RNA-seq experiments showed that dsx(H) was dominantly expressed in Hh mimetic female wings at the P2 stage (Fig. 4h and Supplementary Fig. 8), whereas dsx(H) was barely expressed in Hh males (Supplementary Fig. 8d,e). These results suggest that cis regulatory changes in dsx(H), which may have been fixed by chromosomal inversion, also contribute to the mimetic phenotype. We hypothesized that dsx(H) expression during the early pupal stage is crucial for determining the fate of the mimetic phenotype because this period is important for wing pattern formation10.
To verify the function of dsx in mimetic wing pattern formation, we performed electroporation-mediated small interfering RNA (siRNA) incorporation, enabling mosaic analysis by knocking down the expression of target genes11,12. First, we optimized our methods to minimize side effects on pigmentation patterns13 (Online Methods and Supplementary Fig. 9a–c) using siRNAs for tyrosine hydroxylase (TH) and unrelated sequences (Supplementary Fig. 9d,e, respectively). After this optimization, we incorporated siRNA designed to knock down dsx(H) but not dsx(h) into the whole hindwings of mimetic females, which resulted in non-mimetic wing patterns (Fig. 5a). In addition, electroporation of siRNA targeting dsx(H) into part of the early pupal hindwings of mimetic females resulted in severe repression of red spots and white pigmentation (Fig. 5b, magenta and green dotted arrows in the right hindwing). Ectopic white patterns for non-mimetic females emerged at the predicted position (Fig. 5b, blue arrows). We concluded that dsx(H) not only induces the mimetic wing patterns but also simultaneously represses emergence of the non-mimetic wing patterns (Fig. 5e). In contrast, incorporation of siRNA targeting dsx(h) in mimetic females did not influence the wing phenotype (Fig. 5c). When we knocked down both dsx(H) and dsx(h), we observed the same phenotype as that obtained with siRNA to dsx(H) alone (Fig. 5b,d). The experiments were performed on more than three individuals for each siRNA (Supplementary Fig. 10), and the reduced expression of each target gene was confirmed by RT-PCR (Supplementary Fig. 9f). These results imply that dsx(h) is not involved in mimetic or non-mimetic wing pattern formation.
(a–d) The phenotypes of mimetic females with knockdown of dsx(H) (a,b), dsx(h) (c), and dsx(H) and dsx(h) (d). Each siRNA was injected into either of the pupal wings immediately after pupation and electroporated into the entire dorsal (a) or ventral (b–d) side. (e) Schematic of the dsx(H) RNA interference phenotype. Green and magenta arrowheads represent the repression of mimetic white and red spots, respectively. Blue arrowheads represent the emergence of non-mimetic white regions. Whole-changing patterns (a) and mosaic patterns (b–d) are shown. Scale bars, 1 cm. Two other replicates are shown in Supplementary Figure 10.
New functional analyses undertaken here led to an understanding of the functional role of dsx(H) in changing wing coloration from a non-mimetic to a mimetic pattern. The appearance of the non-mimetic pattern on the wings of mimetic female through knockdown of dsx(H) suggests that the pigmentation pattern is preset. We speculate that both mimetic and non-mimetic patterns are predetermined by genes other than dsx during wing development14,15 and that dsx(H) merely selects the pigmentation processes for the mimetic pattern and represses the non-mimetic pattern in fate-determined wings (Supplementary Fig. 11a). Another possibility is that dsx(H) might be involved in determining mimetic pre-pattern formation in mimetic wings (Supplementary Fig. 11b). The present study shows that H-type transcripts from three genes near the left breakpoint of the inversion were more highly expressed in mimetic females (Fig. 4a,b and Supplementary Fig. 5). Some of these transcripts might be involved in dsx(H) regulation or pre-patterning, although further examination is necessary. Because the inverted region characterized here by means of genome assemblies is the longest region with high allelic divergences across all the autosomes in P. polytes, genomic mutations over 100 kb may be maintained because of repression of recombination at these sites. In addition, although it remains unclear whether the inversion event is linked to emergence of the polymorphism underlying mimicry, the inversion has contributed not only to maintaining altered regulation of dsx and/or neighboring genes but also to differentiation of the function of the encoded Dsx protein, which explains the polymorphic mimicry in this insect.
Female-limited Batesian mimicry is widely observed among several butterfly species16 and may be controlled by similar systems as those in P. polytes. To make the present two Papilio genome data sets available to other researchers, we have built and released PapilioBase, which allows users to perform BLAST searches and download the sequences of all genes and genome scaffolds.
Methods
Insect rearing.
Adult P. polytes females were purchased from Chokan-kabira. P. xuthus were collected in Kashiwa, Japan, or provided by S. Shimizu (Center for Environmental Studies, Tachikawa, Japan). Larvae were reared on Citrus unshiu (Rutaceae) leaves or on an artificial diet (5.6 ml of water, 144 μl of 10% formalin solution, 20 μg of chloramphenicol, 0.8 g for 1st to 2nd instar larvae or 1.2 g for 3rd to 5th instar larvae of Insecta F-II (Nihonnosankogyo) and 0.8 g for 1st to 2nd instar larvae or 0.4 g for 3rd to 5th instar larvae of Citrus natsudaidai leaf powder) under long-day conditions (light:dark = 16 h:8 h) at 25 °C. Pupal samples were staged by the length of time after pupal ecdysis.
De novo assembly and annotation of two Papilio genomes.
Genomic DNA was extracted from one wild-caught larva of P. xuthus (Tokyo, Japan) and one inbred larva of P. polytes. The draft genomes for P. polytes and P. xuthus were assembled with Illumina reads using Platanus (version 1.2.1)6, and validation and annotation of protein-coding genes were performed. Details have been provided in the Supplementary Note.
The rate of single-nucleotide variation in the P. polytes genome.
We estimated the rate of single-nucleotide variation across the whole genome on the basis of the mapping of paired-end reads. The procedures were as follows:
-
1
All paired-end reads were mapped by Bowtie2 (ref. 17), discarding alignments that had edit distances >5.
-
2
A paired-end read was excluded if its insert size did not range from 0.5 to 1.5 times the mean insert size.
-
3
The remaining mapped reads were piled up using SAMtools18, removing bases whose quality was <30 (SAMtools mpileup -Q 30).
-
4
A site with a coverage depth not in the range of 2 to 20 times the mean depth was excluded from SNV determination.
-
5
An SNV was confirmed if the rate of the mismatches was ≥0.25 for both the forward and reverse strands.
As a result, the rate of single-nucleotide variation was estimated as 0.252% (536,696/213,005,066). In addition to estimating the average rate of single-nucleotide variation, we calculated the heterozygosity distribution for every 10-kb non-overlapping window to investigate bias.
We also estimated the substitution rate between the haplotypes in the long inverted region of the H locus. First, the mimetic-type sequence was reversed to ensure the direction was even. Next, all repetitive elements, detected by RepeatMasker, were removed. Finally, the remaining two sequences (74,080 bp and 74,021 bp) were aligned globally using Mafft19. The rate (mismatch/(match + mismatch)) was 23.9% (13,991/58,639), which is markedly higher than that of the whole genome.
Detection of highly diverse and long heterozygous regions of P. polytes.
We searched the regions that were highly diverse between homologous chromosomes (identity ≤90%) using long lengths of sequence (≥100 kb). First, all paired-end reads were mapped to scaffolds using Bowtie2, and coverage depths were derived on the basis of mapped reads with identity ≥90%. Here the H locus was represented as a non-mimetic-type locus. Next, a mean coverage depth was calculated for each window with a size of 100 kb. To exclude influences from gaps and repetitive sequences, we did not use values that were >900 or <150 for the calculation. The coverage depths of targeted heterozygous regions were expected to be half those of homozygous regions, and we identified windows in which the coverage depth was <350. Finally, after clustering overlapping windows, 15 regions, including the H locus, were detected. These regions were mapped to the chromosomes of Bombyx mori, according to the sum of bit scores reported from BLASTN alignments between repeat-masked sequences (tool for masking: RepeatMasker (version 4.0.5)).
Orthologs among lepidopteran species.
We classified all the proteins from five lepidopteran species (P. polytes, P. xuthus, B. mori, Danaus plexippus and Heliconius melpomene) into orthologous groups using Proteinortho20. This tool constructs groups on the basis of all-against-all alignment with BLASTP. The definition of an orthologous relationship between proteins was as follows: e value ≤1 × 10−5, identity ≥25% and alignment coverage ≥50% for both sequences. Protein sequences were obtained from KAIKObase for B. mori, from MonarchBase for D. plexippus and from the Butterfly Genome Database for H. melpomene.
Construction of a phylogenetic tree of concatenated proteins.
We determined orthologous groups of proteins for 5 lepidopteran species and Drosophila melanogaster (see the preceding section) and extracted 2,077 groups that had a one-to-one relationship across all species. For each group, multiple alignment was performed by Mafft19, and the sites containing gaps (‘-’) or ambiguous characters (‘X’) were excluded. All alignments were concatenated, and 748,665 sites were used for the following phylogenetic analysis. The phylogenetic tree was constructed with RAxML21 (Supplementary Fig. 1a), which makes use of the maximum-likelihood method. Here we applied the JTT substitution matrix with a gamma model of rate heterogeneity (-m PROTGAMMAJTT), and the number of replicates for bootstrap analysis was 100.
Construction of a phylogenetic tree for dsx.
A phylogenetic tree for dsx was constructed using five lepidopteran species (Supplementary Fig. 1b,c). Nucleotide sequences for dsx (isoform F3) from non-mimetic (n = 10; 22.7 Gb) and mimetic (n = 10; 26.8 Gb) P. p. alphenor were obtained from SRA (SRP035394) (Supplementary Tables 4 and 5). After mapping these reads to the genome scaffolds assembled in this study using Bowtie2 software17, we constructed each coding sequence (CDS) using SAMtools and the Genome Analysis Toolkit (Supplementary Note). The tree constructed using the maximum-likelihood method was generated following the same procedure as described for concatenated proteins (see the preceding section) (Supplementary Fig. 1b). In addition to using the maximum-likelihood method, we constructed the phylogenetic tree using the Bayesian algorithm method in MrBayes22 software (3.2.2) (Supplementary Fig. 1c).
Linkage mapping.
Two closely related Papilio species, P. polytes (Minamidaitōjima, Japan; no mimetic forms have been isolated thus far from this locality) and P. p. alphenor (Marinduque, Philippines), were used to construct interspecific hybrids to yield considerable variation at sequence-based markers, which facilitated linkage mapping. In brief, P. polytes males were crossed with mimetic P. p. alphenor females by hand-pairing; subsequently, male hybrids that were inevitably phenotypically ‘non-mimetic’ but contained at least half genetically mimetic (Hh) forms were backcrossed with non-mimetic P. polytes females to generate female backcrossed F1 (BC1) individuals. In the present study, we generated 102 BC1 specimens, with 51 mimetic (genotype Hh) and 51 non-mimetic (genotype hh) phenotypes, as expected. We prepared genomic DNA from the thorax muscle of each individual, amplified it using GenomiPhi reagent (GE Healthcare) and used it for the mapping DNA panel (n = 102). We first constructed an EST database for P. polytes. ESTs for candidate markers such as ribosomal proteins, which are mostly single copy and widely distributed across the genome, were selected, and we searched for orthologs in B. mori in KAIKObase. Among these, several ESTs predicted to be representative loci for each of the B. mori linkage groups (from LG1 to LG28) were used for chromosome mapping in P. polytes. Using these primers (Supplementary Table 6), PCR was performed on the genomic DNA of the mapping panel, and products were visualized on 1% agarose gels. Amplicon length variations and/or RFLPs were used to genotype segregating alleles in mapping families. Recombination distances were calculated using MAPMAKER version 3.0 software23. Linkage groups were determined with the GROUP command in the software, performed at a logarithm of odds (LOD) score of >3.0. P. p. alphenor specimens were imported by permission of the Ministry of Agriculture, Forestry and Fisheries, Japan (permission numbers 15-Meisyoku-1130, 17-Meisyoku-919, 19-Meisyoku-1112 and 23-Meisyoku-462 to H. Hori).
Association study.
Twenty PCR-based markers were used to estimate the association between genotype and phenotype for 28 non-mimetic and 26 mimetic wild-caught female adults from Okinawa and Ishigaki in Japan, which were genotyped by sequence variations (SNPs) and by visualization of differences in amplicon size (Supplementary Table 7). We estimated the association between genotype and phenotype by testing the hypothesis that polymorphic differences at each site (SNPs or differences in size of the PCR fragment) in each individual occurred regardless of a mimetic or non-mimetic phonotype. For each polymorphic site, we counted the population of individuals with each polymorphism and then calculated P value using the χ2 test to test the hypothesis. Thus, the region including dsx with a lower P value was identified as the genomic region where the genotype and phenotype were strongly related.
Assembly of RNA sequencing data and dsx structures.
RNA-seq reads were sequenced with the HiSeq 2000 platform (Illumina). To construct transcripts from homozygotes for the H locus (HH), we sequenced one paired-end library derived from the mimetic wings of one HH female pupa at the P2 stage to obtain 76 mega reads and performed de novo assembly using Trinity 2013-2-16 (ref. 24). For non-mimetic females (hh), reads from 5 paired-end libraries at the wandering and P10.5 stages were mixed (256 mega reads in total), and assembly was performed as described above. Isoforms of dsx were searched using BLAST against the assembled contigs, and contig hits were aligned to the genomic scaffolds to identify exon-intron structures using Exonerate 2.2.0 (ref. 25). After we removed redundant contigs, five and six variations were identified for the hh and HH genotypes, respectively. The structures of the male-specific dsx isoform on the h and H alleles were identified using RNA-seq data derived from heterozygous male (Hh).
Homology between heterozygous regions of h, H and P. xuthus.
Schematic views of homology between heterozygous regions of h, H and P. xuthus were constructed using the LAGAN26 program and visualized by VISTA27. VISTA was used to generate pairwise alignments between two species. Each graph is a plot of nucleotide identity, with a 70% identity threshold and a 100-bp sliding window centered on a given position. Annotated genes are shown at the top of the alignment.
Allele-specific dsx expression.
In the wings of individuals heterozygous at the H locus (Hh), dsx was transcribed from both the h (dsx(h)) and H (dsx(H)) alleles. The ratio of dsx(H)/dsx(h) expression levels was estimated using RNA-seq data: two female wing libraries and one male wing library for the wandering stage, four female wing libraries and one male wing library for the P2 stage, and three female wing libraries for the P10.5 stage. RNA-seq reads were mapped to all female-specific isoforms of the dsx ORF for dsx(h) and dsx(H) using Bowtie 1.0.0 (ref. 28). Normalized counted data were obtained by dividing the ORF length by the number of total reads (FPKM). For male individuals heterozygous at the H locus (Hh), only exon 1 of the dsx ORF was used as a template for mapping in Bowtie; this exon is common to known female- and male-specific isoforms.
RT-PCR.
Total RNA was extracted from the hindwings of each individual using TRI reagent (Sigma). After treatment of RNA with approximately 2.5 U of DNase I (Takara) per 1 μg of RNA for 15 min at 37 °C and phenol-chloroform extraction, reverse transcription was performed using a First-Strand cDNA Synthesis kit (GE Healthcare) with random hexamer primers. A StepOne Real-Time PCR system (Applied Biosystems) with Power SYBR Green PCR Master Mix (Applied Biosystems) was used for quantification of gene expression levels. Standard curves were generated for all primer pairs (Supplementary Fig. 12 and Supplementary Table 6), and normalized relative gene expression values were estimated using ribosomal protein L3 (RpL3) expression levels as an internal control. The expression levels shown are relative to maximal expression (set to 1). Statistical analysis was performed on data from individuals in each of the three pupal stages. Statistical significance at early and mid-pupal stages was determined using a Student's t test for UXT, U3X, dsx F2 and dsx F3 at both stages and for dsx F1 and dsx from the h allele at the early pupal stage (in the case of homoscedasticity) or a Welch's t test for dsx from the H allele at both stages and for dsx F1 and dsx from the h allele at the mid-pupal stage (in the case of heteroscedasticity). For the late pupal stage, two-way factorial ANOVA was performed. If significant differences were detected, we used Tukey's HSD as a post-hoc test.
In vivo electroporation.
In vivo electroporation was modified from the method described by Ando and Fujiwara11. A P. polytes individual collected just after pupation was anesthetized at 4 °C before the experiment. Experimental procedures are described in Supplementary Figure 9a–c. Pupal wings were opened and fixed on parafilm laid under the pupal hindwing (Supplementary Fig. 9a,b). siRNA solution (250 μM) dissolved in injection buffer (Sigma-Aldrich) was then loaded into a glass capillary processed into a needle-like shape using a single-stage glass microelectrode puller (PP-830, Narishige). Using the glass capillary, a solution containing siRNA (1 μl) (Supplementary Fig. 12 and Supplementary Table 6) was injected into the pupal hindwing via a FemtoJet Microinjector (Eppendorf). After injection, solidified 1% agarose in PBS was laid between the pupal wing and the parafilm, and PBS was placed around the M1 and M3 regions on the ventral side of the pupal hindwing. Positive and negative electrodes were attached to the PBS and the 1% agarose gel, respectively (when the dorsal pattern was analyzed, the positive and negative electrodes were attached to the 1% agarose gel and the PBS, respectively). Thereafter, 7.5 V was applied to the pupal hindwing (in five 280-ms pulses/5 s) (Supplementary Fig. 9a,c). It is known that electroporation causes serious side effects on pigmentation patterns of the wings13. To avoid these effects, we ran trials at 25, 20, 15 and 7.5 V and found that there were no side effects on wings at 7.5 V. In addition, to avoid direct contact with electrodes, we applied them to the pupal wings indirectly via PBS and 1% agarose. After eclosion, phenotypes were observed under a stereomicroscope (M165 FC, Leica).
URLs.
The genome database for P. polytes and P. xuthus is available at PapilioBase (http://papilio.bio.titech.ac.jp/papilio.html or http://papilio.nig.ac.jp/). The protein sequences used to construct the protein ortholog groups were obtained from KAIKObase (http://sgp.dna.affrc.go.jp/KAIKObase/) for Bombyx mori, MonarchBase (http://monarchbase.umassmed.edu/) for Danaus plexippus and the Butterfly Genome Project database (http://www.butterflygenome.org/) for Heliconius melpomene. To construct the dsx nucleotide sequences, we used SAMtools (http://samtools.sourceforge.net/) and the Genome Analysis Toolkit (https://www.broadinstitute.org/gatk/).
Accession codes.
Raw sequencing information has been deposited in the DNA Data Bank of Japan. Accession information is as follows: short reads for the P. polytes and P. xuthus genomes, BioProjects PRJDB2954 and PRJDB2956, respectively; transcriptome sequences, BioProjects PRJDB2955 and PRJDB2957; assembled genome scaffolds of P. polytes and P. xuthus, GenBank DF820621–DF824493 and DF824494–DF830065, respectively.
References
Kunte, K. et al. Doublesex is a mimicry supergene. Nature 507, 229–232 (2014).
Clarke, C.A. & Sheppard, P.M. The genetics of the mimetic butterfly Papilio polytes L. Phil. Trans. R. Soc. Lond. B 263, 431–458 (1972).
Clarke, C.A. & Sheppard, P.M. Super-genes and mimicry. Heredity 14, 175–185 (1960).
Loehlin, D.W. & Carroll, S.B. Evolutionary biology: sex, lies and butterflies. Nature 507, 172–173 (2014).
Joron, M. et al. Chromosomal rearrangements maintain a polymorphic supergene controlling butterfly mimicry. Nature 477, 203–206 (2011).
Kajitani, R. et al. Efficient de novo assembly of highly heterozygous genomes from whole-genome shotgun short reads. Genome Res. 24, 1384–1395 (2014).
Zakharov, E.V., Caterino, M.S. & Sperling, F.A. Molecular phylogeny, historical biogeography, and divergence time estimates for swallowtail butterflies of the genus Papilio (Lepidoptera: Papilionidae). Syst. Biol. 53, 193–215 (2004).
Schroer, A., Schneider, S., Ropers, H. & Nothwang, H. Cloning and characterization of UXT, a novel gene in human Xp11, which is widely and abundantly expressed in tumor tissue. Genomics 56, 340–343 (1999).
An, W., Cho, S., Ishii, H. & Wensink, P.C. Sex-specific and non-sex-specific oligomerization domains in both of the doublesex transcription factors from Drosophila melanogaster. Mol. Cell. Biol. 16, 3106–3111 (1996).
Nijhout, H.F. Pattern formation on lepidopteran wings: determination of an eyespot. Dev. Biol. 80, 267–274 (1980).
Ando, T. & Fujiwara, H. Electroporation-mediated somatic transgenesis for rapid functional analysis in insects. Development 140, 454–458 (2013).
Yamaguchi, J., Mizoguchi, T. & Fujiwara, H. siRNAs induce efficient RNAi response in Bombyx mori embryos. PLoS ONE 6, e25469 (2011).
Golden, K., Sagi, V., Markwarth, N., Chen, B. & Monteiro, A. In vivo electroporation of DNA into the wing epidermis of the butterfly, Bicyclus anynana. J. Insect Sci. 7, 1–8 (2007).
Monteiro, A., Glaser, G., Stockslager, S., Glansdorp, N. & Ramos, D. Comparative insights into questions of lepidopteran wing pattern homology. BMC Dev. Biol. 6, 52 (2006).
Brakefield, P.M. et al. Development, plasticity and evolution of butterfly eyespot patterns. Nature 384, 236–242 (1996).
Kunte, K. The diversity and evolution of Batesian mimicry in Papilio swallowtail butterflies. Evolution 63, 2707–2716 (2009).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Katoh, K. & Standley, D.M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Lechner, M. et al. Proteinortho: detection of (co-)orthologs in large-scale analysis. BMC Bioinformatics 12, 124 (2011).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Ronquist, F. & Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Lander, E.S. et al. MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1, 174–181 (1987).
Grabherr, M.G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Slater, G.S. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31 (2005).
Brudno, M. et al. LAGAN and Multi-LAGAN: efficient tools for large-scale multiple alignment of genomic DNA. Genome Res. 13, 721–731 (2003).
Mayor, C. et al. VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16, 1046–1047 (2000).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Acknowledgements
We thank A. Toyoda, Y. Kohara, R. Futahashi, T. Kojima, K.K. Kojima, H. Yoshikawa, M. Hasebe, Y. Kondo, S. Kawamura, H. Maekawa and J.M. Otaki for helpful suggestions on our research. We also thank R. Kurokawa for technical assistance with data analysis in the association study. We also thank T. Kojima for helpful comments on the manuscript. This work was supported by Grants-in-Aid for Scientific Research on Priority Areas ‘Comparative Genomics’ from the Ministry of Education, Culture, Sports, Science and Technology of Japan 18017007 and 20017007 (to H.F.), Grant-in-Aid for Scientific Research 22128005 (to H.F.) and KAKENHI for Innovative Areas 221S0002.
Author information
Authors and Affiliations
Contributions
H.N., T. Iijima and J.Y. provided organisms and isolated the RNA and DNA. R.K., Y.S., S.S., A.F., S.K., H. Hirakawa, S.T., K.O. and T. Itoh designed and performed genome sequencing, assembly and annotation. M.O., H. Hori, T. Iijima and J.Y. performed linkage analyses. H.M., R.K. and T. Itoh built the genome browser. H. Hori, K.I., R.K. and T. Itoh sequenced fosmids. H.N., T. Iijima, J.Y., T.A. and H.F. designed and performed the experiments. H.N., T. Iijima, R.K., J.Y., T. Itoh and H.F. wrote the manuscript, and J.Y., T. Itoh and H.F. edited the manuscript. H. Hori, T. Itoh and H.F. supervised this project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Phylogenetic trees of concatenated protein and Dsx proteins.
(a) Phylogenetic tree of concatenated proteins. Multiple alignments of 2,077 ortholog groups were concatenated, and a phylogenetic tree was constructed using 748,665 sites. (b) Phylogenetic tree of Dsx proteins. A total of 264 Dsx sites were used. (c) Phylogenetic tree of Dsx proteins (Bayesian method using MrBayes). A total of 264 Dsx sites were used.
Supplementary Figure 2 Details of the assembly for the hetero_130kb regions on h and H.
Schematics of fosmid clones, the mate-pair libraries of 5.5 kb and the scaffolds of the previous version of the assembly (Version 121225), which were used for the characterization of hetero_130kb on h (a) and H (b). Dashed red lines indicate the breakpoints. All fosmid clones were sequenced and aligned to the assembled scaffolds. Hetero_130kb was covered completely on h with six fosmid clones and largely on H with nine fosmid clones, verifying two different scaffolds, scaff651 and scaff938, in both of which the dsx gene is encoded in the inverted direction. All breakpoints were validated by PCR and sequencing. The regions sequenced using PCR are shown by rectangles with the PCR primers (black dots connected by lines). n denotes the numbers of individuals used in the PCR analysis. The mate-pair libraries of 5.5 kb are partially illustrated on H around the breakpoints (shown as many short black lines) to demonstrate the reliability of the orientation of the scaffolds near breakpoints. Their coverage depths are shown in the blue chart. (c) Detailed information on the PCR validation around left breakpoint on H. The gap between scaff34139 and scaff938 in the previous assembly was filled by PCR (the reverse primer is specific to the H chromosome) and sequencing. The 1,961-bp nucleotide sequence was included in fosmids 006O10 and 002D08 as expected. (d) Detail of the PCR validation around the right breakpoint on the H chromosome.
Supplementary Figure 3 Annotation overview of h and H around the left breakpoint.
(a) Gene structures predicted in the 200-kb genomic region including the inversion. (b) Enlarged view of 20 kb around the left breakpoint. U3X is transcribed only from H. The two dotted boxes indicate the regions where some RNA-seq reads were mapped on H but not on h; this is described in detail in Supplementary Figure 5. (c) An enlarged view of 200 bp near the left breakpoint. UXT8,33 crosses the breakpoint at its 5′ UTR (yellow boxes); therefore, the sequences between the transcriptional start point and 95 bp upstream of a translational start are very different. Asterisks indicate the same nucleotides in h and H.
Supplementary Figure 4 Annotation overview of h and H around the right breakpoint.
(a) Gene structures predicted in the 200-kb genomic region including the inversion. (b) Enlarged view of 20 kb around the right breakpoint. (c) Nucleotide sequences around the breakpoint are shown. Asterisks indicate the same nucleotides in h and H.
Supplementary Figure 5 RNA-seq analysis of the unknown transcripts downstream of prospero.
Mapping with RNA-seq reads from homozygous H (HH, top) and homozygous h (hh, bottom). The positions of the new transcripts detected in mimetic individuals are shown by dotted boxes. Transcriptional regions were detected with the mapped reads using TopHat and Cufflinks. Results were visualized by Integrative Genomics Viewer (http://www.broadinstitute.org/igv/home).
Supplementary Figure 6 Amino acid substitutions in three female-specific isoforms of Dsx.
The common regions among the three isoforms (F1–F3) are shown in the second column; the other regions are in the third column. Red amino acids indicate substitutions between h and H for each allele. Underlined sequences represent the Dsx DNA-binding motif (DM, smart00301) and the Dsx dimerization domain (Dsx dimer, pfam08828) identified in a conserved domain9,34 search in BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi).
Supplementary Figure 7 Amino acid sequence substitutions in Dsx.
Amino acid sequences of Dsx from the h and H alleles in P. polytes (Pp), P. xuthus (Px), D. plexippus (Dp), B. mori (Bm) and Manduca sexta (Ms) were aligned. Asterisks indicate stop codons. Amino acids conserved among all species are indicated in blue. Variable amino acids are shown in different colors. The positions of substitutions between h and H in P. polytes are marked by filled circles, and amino acid sequences are shown in red and orange, respectively. Red-filled circles indicate the positions of the substitutions that occurred only on H in P. polytes. Alignments based on the female-specific F3 isoform of Dsx in P. polytes (a), the C-terminal portion of the female-specific F1 isoform (b), the C-terminal portion of the female-specific F2 isoform (c) and male-specific isoforms (d) are shown. Functional domains were predicted on the basis of a previous report in B. mori35. The sequences of the female-specific Dsx isoforms in B. mori (Bm) were obtained from GenBank (KF255811, AGS48298 and NP_001036871 for isoforms 1, 2 and 3, respectively). Others were obtained from the Genome Project, from Monarchbase (http://monarchbase.umassmed.edu) for D. plexippus (Dp) and from Manducabase (http://agripestbase.org/manduca/) for M. sexta (Ms). A superscript number beside the species name indicates that the sequences were obtained from each transcriptome database (OGS2_20140407_proteins for Ms1 and Monarch genes OGS2.0 for Dp1) and were predicted from each genomic sequence (Manduca_Assembly 1.0 ScaffoldDB for Ms2 and Monarch genome assembly v3 for Dp2).
Supplementary Figure 8 Allele-specific expression of dsx.
Expression levels of dsx from the h and H alleles were analyzed using RNA-seq data from heterozygotes for H (Hh) in the wandering (a), P2 (b) and P10.5 (c) stages. RNA-seq reads were mapped to ORF sequences of the dsx transcript from the h allele (dsx(h)) and H allele (dsx(H)), followed by expression level (FPKM) estimation. (d) In an Hh male, dsx expression from the H allele is much lower than that from the h allele. (e) The ratio of dsx(H)/dsx(h) expression levels at each stage. In P2, dsx(H) levels were more than two-fold higher than those of dsx(h) in an Hh female.
Supplementary Figure 9 Functional analyses using in vivo electroporation.
(a) Schematic of the functional analysis procedure. Pupal wings were opened and fixed onto parafilm to easily perform the analysis. Next, 1 µl of injection buffer containing siRNA (250 µM) was injected into the pupal hindwing with a glass capillary via a FemtoJet Microinjector (Eppendorf) (b). After injection, solidified 1% agarose in PBS was laid between the pupal wing and parafilm. Then, PBS solution was put onto the ventral side of the hindwing. Positive and negative electrodes were attached to the PBS solution and 1% agarose gel, respectively (in cases where the dorsal pattern was analyzed, the positive and negative electrodes were reversed), followed by the application of voltage pulses (c). After in vivo electroporation, we evaluated the phenotype of each adult-stage individual. (d) The effects of melanin synthesis gene tyrosine hydroxylase (si-TH)-targeting siRNA. The right and left sides show the wings treated and untreated with siRNA, respectively. siRNA was injected into the pupal wings just after pupation and electroporated around the M1 and M3 regions on the ventral side of the hindwings. The dotted blue line indicates the area that was expected to introduce si-TH. A reduction in black pigment was observed, which corresponded to the expected phenotype36,37. Three biological replicates were performed. (e) A negative control using Universal Negative Control siRNA (Nippongene) under the same conditions. No phenotypic effects were detected. Three biological replicates were performed. Scale bars, 1 cm. (f) Verification of reduction in target gene expression in the RNAi experiments. We estimated the ratio of reduction in expression of the target gene between untreated and treated wings for each siRNA in the same individual by RT-PCR using RpL3 as an internal control. Each mRNA expression level was measured at stages P4–P5 for dsx and at stage P12 for si-TH. An asterisk indicates statistical significance (P < 0.05) in a one-sample t test. Note that we dissected both sides of the cell layers (dorsal and ventral) together for estimation, whereas siRNA was incorporated by electroporation only into the ventral side and phenotypic effects were always observed on this side. Lower efficiency of reduction in target gene expression may be explained by this.
Supplementary Figure 10 Phenotype of dsx RNAi for each individual in the hindwings of P. polytes.
The phenotype of each individual after knockdown of dsx(H) (a), dsx(H) and dsx(h) (b), or dsx(h) (c) in hindwings is shown. siRNA was injected into the pupal hindwings just after pupation and electroporated into the ventral side of the hindwings. Scale bars, 1 cm.
Supplementary Figure 11 Model for the function of dsx(H) in wing pattern formation.
(a) The hypothetical function of dsx(H) as a pigmentation selector. (b) Model for pre-pattern determination of mimetic wing patterns by dsx(H).
Supplementary Figure 12 Schematics of dsx structures.
Genes were obtained from the de novo–assembled transcriptome and aligned with genomic fragments. (a) Variations of dsx obtained from the assembled RNA-seq data of five h homozygous (hh) females. Note that alternative splicing is abundantly apparent between the third and fourth exons (green box). At least five female-specific isoforms were found to encode three types of ORFs (blue box in F1, F2 and F3; red vertical lines indicate the positions of each stop codon). The male-specific isoform shown at the bottom was obtained from additional RNA-seq data for a male individual. (b) Variations of dsx in H homozygotes (HH) that were obtained from RNA-seq data for one HH female. The F2′ and F3′ translational start positions differed from those of F1 and F2 (MN is substituted for MVSVGAWRRRSPDEC). Scale bars, 500 bp. The sequences for the 3′ UTR were not exactly determined. Isoform-specific primers A to C were designed against common sequences in hh and HH. Primer D in a and primer E in b were designed to detect only transcripts from hh and HH, respectively. Green arrows indicate the targeted portions in the qRT-PCR analysis. The red arrowheads labeled “h” and “H” indicate the targets selected for the dsx(h) and dsx(H) siRNAs, respectively. The red arrowheads labeled “H/h” indicate targets common to both h- and H-type alleles. (c) Differences in the target sequences for the siRNAs for dsx(H) and for dsx(h).
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–12 and Supplementary Tables 1–6. (PDF 2296 kb)
Supplementary Table 7
Detailed information for the association study. (XLS 34 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Nishikawa, H., Iijima, T., Kajitani, R. et al. A genetic mechanism for female-limited Batesian mimicry in Papilio butterfly. Nat Genet 47, 405–409 (2015). https://doi.org/10.1038/ng.3241
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3241
This article is cited by
-
Evolutionary genomics of camouflage innovation in the orchid mantis
Nature Communications (2023)
-
Sex-specific morphs: the genetics and evolution of intra-sexual variation
Nature Reviews Genetics (2023)
-
Whole genome sequencing of spotted stem borer, Chilo partellus, reveals multiple genes encoding enzymes for detoxification of insecticides
Functional & Integrative Genomics (2022)
-
Genome-wide identification and gene-editing of pigment transporter genes in the swallowtail butterfly Papilio xuthus
BMC Genomics (2021)
-
A combined RAD-Seq and WGS approach reveals the genomic basis of yellow color variation in bumble bee Bombus terrestris
Scientific Reports (2021)