Abstract
Elucidating genetic causes of cholestasis has proved to be important in understanding the physiology and pathophysiology of the liver. Here we show that protein-truncating mutations in the tight junction protein 2 gene (TJP2) cause failure of protein localization and disruption of tight-junction structure, leading to severe cholestatic liver disease. These findings contrast with those in the embryonic-lethal knockout mouse, highlighting differences in redundancy in junctional complexes between organs and species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Accession codes
References
Clayton, R.J. et al. Am. J. Dis. Child. 117, 112–124 (1969).
Bull, L.N. et al. Nat. Genet. 18, 219–224 (1998).
de Vree, J.M. et al. Proc. Natl. Acad. Sci. USA 95, 282–287 (1998).
Strautnieks, S.S. et al. Nat. Genet. 20, 233–238 (1998).
Davit-Spraul, A. et al. Hepatology 51, 1645–1655 (2010).
Mitic, L.L. et al. Am. J. Physiol. Gastrointest. Liver Physiol. 279, G250–G254 (2000).
Itoh, M. et al. J. Cell Biol. 147, 1351–1363 (1999).
Holczbauer, Á. et al. J. Histochem. Cytochem. 61, 294–305 (2013).
Xu, J. et al. Mol. Cell. Biol. 28, 1669–1678 (2008).
Fanning, A.S. et al. Mol. Biol. Cell 23, 577–590 (2012).
Chlenski, A. et al. Biochim. Biophys. Acta 1493, 319–324 (2000).
Carlton, V.E. et al. Nat. Genet. 34, 91–96 (2003).
Grosse, B. et al. Hepatology 55, 1249–1259 (2012).
Li, H. et al. Bioinformatics 25, 2078–2079 (2009).
DePristo, M.A. et al. Nat. Genet. 43, 491–498 (2011).
McKenna, A. et al. Genome Res. 20, 1297–1303 (2010).
Plagnol, V. et al. Bioinformatics 28, 2747–2754 (2012).
McLaren, W. et al. Bioinformatics 26, 2069–2070 (2010).
Conrad, D.F. et al. Nature 464, 704–712 (2010).
Acknowledgements
M.S. was funded by an Alex Mowat PhD Studentship. R.J.T. and B.E.C. received funding for this work from a consumables grant awarded by the King's College Hospital Department of Research and Development. Targeted resequencing and whole-exome sequencing were supported by the National Institute for Health Research (NIHR) Biomedical Research Centre based at Guy's and St Thomas' NHS Foundation Trust and King's College London. The views expressed are those of the authors and not necessarily those of the NHS, the NIHR or the UK Department of Health. C.A.J. was supported by a Sir Jules Thorn Award for Biomedical Research (JTA/09). Additional funding for this project included US National Institutes of Health (NIH) grant R56 DK094828 to L.N.B. and R.J.T., the University of California, San Francisco (UCSF)–King's College Health Partners Faculty Fellowship Travel Grant (UCSF Academic Senate) to L.N.B., US NIH grant U01 DK062500 to P. Rosenthal, US NIH grants U01 DK062453 and UL1 TR000154 to R.J.S. and US NIH grant U01 DK062456 to J.C.M. Further whole-exome sequencing was undertaken by the University of Washington Center for Mendelian Genomics (UW CMG) and was funded by the National Human Genome Research Institute and by NHLBI grant 1U54HG006493 to D. Nickerson, J. Shendure and M. Bamshad.
Author information
Authors and Affiliations
Consortia
Contributions
M.S. performed the majority of the experimental work, analyzed and interpreted data, and wrote the manuscript. S.S., E.P. and C.V.L. performed experiments. P.R. and B.E.C. helped design the targeted capture. D.A.P., J.D.S. and M.A.S. analyzed whole-exome sequencing data. L.J.N. helped with protein blotting. B.E.W. performed electron microscopy. B.M.K., S.L. and P.M. provided cases and sequence data. G.M.-V. and T.G. provided cases. J.C.M. and R.J.S. run ChiLDREN and provided samples and clinical data. The University of Washington Center for Mendelian Genomics performed whole-exome sequencing for ChiLDREN cases. A.S.K. undertook histopathological analysis. C.A.J. and L.N.B. directed the whole-exome sequencing projects. R.J.T. initiated the project, analyzed the data and wrote the manuscript. All authors commented on and edited the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Further information appears in the Supplementary Note
Integrated supplementary information
Supplementary Figure 1 Flow chart of work undertaken.
Targeted resequencing (TRS), whole-exome sequencing (WES) and Sanger sequencing (SS) were conducted in one center. Whole-exome sequencing was conducted in three other laboratories, as shown. Sample numbers for each are indicated. TJP2 was identified as the disease-causing gene by targeted resequencing and whole-exome sequencing independently. All subsequent analyses were conducted in light of this knowledge.
Supplementary Figure 2 Pedigrees of the eight families found to be mutated in TJP2.
Mutated individuals are indicated with filled shapes. Unfilled shapes are used for untested relatives. DNA was not available from unaffected individuals, with the exception of family 8, where three unaffected individuals were tested. All three were heterozygous for the disease-causing mutation and are marked with dots. The deceased child in family 2 died of cardiac disease. The cause of death in family 5 is not known. The three deceased children in family 6 all had chronic obstructive pulmonary disease and cholestatic liver disease.
Supplementary Figure 3 Quantitative analysis of TJP2 mRNA levels in five patients and six controls.
TJP2 expression levels were measured in liver tissue by TaqMan-based quantitative RT-PCR. The expression in all is expressed relative to the mean of the control samples. Each data point represents a single liver sample, although all were tested in triplicate.
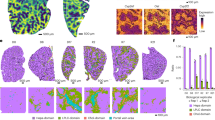
Supplementary Figure 4 Transmission electron microscopy of tight-junction structure in liver biopsies.
All four panels show tight-junction complexes between adjacent hepatocytes and biliary canaliculi. In each panel, a red asterisk indicates the canalicular space. Panel a is from a liver biopsy specimen obtained at presentation in patient 1. Panel b is taken from the explanted liver of patient 5b. Panel c is from a patient with BSEP deficiency and panel d from a patient with FIC1 deficiency. Tight junctions appear to extend deeper into the paracellular, or lateral, space in the TJP2-mutated patients, with diminution of the most electron-dense part of the zona occludens (shown with arrows in c and d). In all cases, scale bar, 500 nm; OsO4/uranyl acetate/lead citrate.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–4 and Supplementary Tables 1–5 (PDF 1509 kb)
Rights and permissions
About this article
Cite this article
Sambrotta, M., Strautnieks, S., Papouli, E. et al. Mutations in TJP2 cause progressive cholestatic liver disease. Nat Genet 46, 326–328 (2014). https://doi.org/10.1038/ng.2918
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2918
This article is cited by
-
Angeborene Cholestasesyndrome im Erwachsenenalter: Relevantes für Diagnostik und Therapie
Die Gastroenterologie (2023)
-
Rare variant contribution to cholestatic liver disease in a South Asian population in the United Kingdom
Scientific Reports (2023)
-
Clinical outcomes of surgical management for rare types of progressive familial intrahepatic cholestasis: a case series
Surgical Case Reports (2022)
-
ZO-2/Tjp2 suppresses Yap and Wwtr1/Taz-mediated hepatocyte to cholangiocyte transdifferentiation in the mouse liver
npj Regenerative Medicine (2022)