Abstract
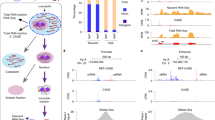
Despite the conventional distinction between them, promoters and enhancers share many features in mammals, including divergent transcription and similar modes of transcription factor binding. Here we examine the architecture of transcription initiation through comprehensive mapping of transcription start sites (TSSs) in human lymphoblastoid B cell (GM12878) and chronic myelogenous leukemic (K562) ENCODE Tier 1 cell lines. Using a nuclear run-on protocol called GRO-cap, which captures TSSs for both stable and unstable transcripts, we conduct detailed comparisons of thousands of promoters and enhancers in human cells. These analyses identify a common architecture of initiation, including tightly spaced (110 bp apart) divergent initiation, similar frequencies of core promoter sequence elements, highly positioned flanking nucleosomes and two modes of transcription factor binding. Post-initiation transcript stability provides a more fundamental distinction between promoters and enhancers than patterns of histone modification and association of transcription factors or co-activators. These results support a unified model of transcription initiation at promoters and enhancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007).
Core, L.J., Waterfall, J. & Lis, J. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Seila, A.C. et al. Divergent transcription from active promoters. Science 322, 1849–1851 (2008).
Almada, A.E., Wu, X., Kriz, A.J., Burge, C.B. & Sharp, P.A. Promoter directionality is controlled by U1 snRNP and polyadenylation signals. Nature 499, 360–363 (2013).
Ntini, E. et al. Polyadenylation site–induced decay of upstream transcripts enforces promoter directionality. Nat. Struct. Mol. Biol. 20, 923–928 (2013).
Trinklein, N.D. et al. An abundance of bidirectional promoters in the human genome. Genome Res. 14, 62–66 (2004).
Oler, A.J. et al. Human RNA polymerase III transcriptomes and relationships to Pol II promoter chromatin and enhancer-binding factors. Nat. Struct. Mol. Biol. 17, 620–628 (2010).
Kim, T.K. et al. Widespread transcription at neuronal activity–regulated enhancers. Nature 465, 182–187 (2010).
Hah, N., Murakami, S., Nagari, A., Danko, C.G. & Kraus, W.L. Enhancer transcripts mark active estrogen receptor binding sites. Genome Res. 23, 1210–1223 (2013).
Wang, D. et al. Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature 474, 390–394 (2011).
Koch, F. et al. Transcription initiation platforms and GTF recruitment at tissue-specific enhancers and promoters. Nat. Struct. Mol. Biol. 18, 956–963 (2011).
Heintzman, N.D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Heintzman, N.D. & Ren, B. Finding distal regulatory elements in the human genome. Curr. Opin. Genet. Dev. 19, 541–549 (2009).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: what, how, and why? Mol. Cell 49, 825–837 (2013).
Struhl, K. Transcriptional noise and the fidelity of initiation by RNA polymerase II. Nat. Struct. Mol. Biol. 14, 103–105 (2007).
Ørom, U.A. & Shiekhattar, R. Long non-coding RNAs and enhancers. Curr. Opin. Genet. Dev. 21, 194–198 (2011).
Lam, M.T., Li, W., Rosenfeld, M.G. & Glass, C.K. Enhancer RNAs and regulated transcriptional programs. Trends Biochem. Sci. 39, 170–182 (2014).
Seila, A.C., Core, L.J., Lis, J.T. & Sharp, P.A. Divergent transcription: a new feature of active promoters. Cell Cycle 8, 2557–2564 (2009).
Melgar, M.F., Collins, F.S. & Sethupathy, P. Discovery of active enhancers through bidirectional expression of short transcripts. Genome Biol. 12, R113 (2011).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Kruesi, W.S., Core, L.J., Waters, C.T., Lis, J.T. & Meyer, B.J. Condensin controls recruitment of RNA polymerase II to achieve nematode X-chromosome dosage compensation. Elife 2, e00808 (2013).
Kwak, H., Fuda, N.J., Core, L.J. & Lis, J.T. Precise maps of RNA polymerase reveal how promoters direct initiation and pausing. Science 339, 950–953 (2013).
Andersson, R. et al. Nuclear stability and transcriptional directionality separate functionally distinct RNA species. bioRxiv 10.1101/005447 (29 August 2014).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Orkin, S.H. Regulation of globin gene expression in erythroid cells. Eur. J. Biochem. 231, 271–281 (1995).
Venters, B.J. & Pugh, B.F. Genomic organization of human transcription initiation complexes. Nature 502, 53–58 (2013); retraction 513, 444 (2014).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Shiraki, T. et al. Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage. Proc. Natl. Acad. Sci. USA 100, 15776–15781 (2003).
Maruyama, K. & Sugano, S. Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides. Gene 138, 171–174 (1994).
Affymetrix ENCODE Transcriptome Project & Cold Spring Harbor Laboratory ENCODE Transcriptome Project. Post-transcriptional processing generates a diversity of 5′-modified long and short RNAs. Nature 457, 1028–1032 (2009).
Piper, J. et al. Wellington: a novel method for the accurate identification of digital genomic footprints from DNase-seq data. Nucleic Acids Res. 41, e201 (2013).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008).
Clouaire, T. et al. Cfp1 integrates both CpG content and gene activity for accurate H3K4me3 deposition in embryonic stem cells. Genes Dev. 26, 1714–1728 (2012).
Schones, D.E. et al. Dynamic regulation of nucleosome positioning in the human genome. Cell 132, 887–898 (2008).
Gaffney, D.J. et al. Controls of nucleosome positioning in the human genome. PLoS Genet. 8, e1003036 (2012).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Coulombe, B. & Burton, Z.F. DNA bending and wrapping around RNA polymerase: a “revolutionary” model describing transcriptional mechanisms. Microbiol. Mol. Biol. Rev. 63, 457–478 (1999).
Foley, J.W. & Sidow, A. Transcription-factor occupancy at HOT regions quantitatively predicts RNA polymerase recruitment in five human cell lines. BMC Genomics 14, 720 (2013).
Wamstad, J.A. et al. Dynamic and coordinated epigenetic regulation of developmental transitions in the cardiac lineage. Cell 151, 206–220 (2012).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Zentner, G.E., Tesar, P.J. & Scacheri, P.C. Epigenetic signatures distinguish multiple classes of enhancers with distinct cellular functions. Genome Res. 21, 1273–1283 (2011).
Creyghton, M.P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl. Acad. Sci. USA 107, 21931–21936 (2010).
Stergachis, A.B. et al. Developmental fate and cellular maturity encoded in human regulatory DNA landscapes. Cell 154, 888–903 (2013).
Pekowska, A. et al. H3K4 tri-methylation provides an epigenetic signature of active enhancers. EMBO J. 30, 4198–4210 (2011).
Shilatifard, A. The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu. Rev. Biochem. 81, 65–95 (2012).
Damgaard, C.K. et al. A 5′ splice site enhances the recruitment of basal transcription initiation factors in vivo. Mol. Cell 29, 271–278 (2008).
Banerji, J., Rusconi, S. & Schaffner, W. Expression of a β-globin gene is enhanced by remote SV40 DNA sequences. Cell 27, 299–308 (1981).
Kaida, D. et al. U1 snRNP protects pre-mRNAs from premature cleavage and polyadenylation. Nature 468, 664–668 (2010).
Berg, M.G. et al. U1 snRNP determines mRNA length and regulates isoform expression. Cell 150, 53–64 (2012).
Wu, X. & Sharp, P.A. Divergent transcription: a driving force for new gene origination? Cell 155, 990–996 (2013).
Green, P. et al. Transcription-associated mutational asymmetry in mammalian evolution. Nat. Genet. 33, 514–517 (2003).
Kent, W.J. et al. in The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Ashe, H.L., Monks, J., Wijgerde, M., Fraser, P. & Proudfoot, N.J. Intergenic transcription and transinduction of the human β-globin locus. Genes Dev. 11, 2494–2509 (1997).
Wang, I.X. et al. RNA-DNA differences are generated in human cells within seconds after RNA exits polymerase II. Cell. Rep. 6, 906–915 (2014).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet 17, 10–12 (2011).
Carninci, P. et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nat. Genet. 38, 626–635 (2006).
Sandelin, A. et al. Mammalian RNA polymerase II core promoters: insights from genome-wide studies. Nat. Rev. Genet. 8, 424–436 (2007).
Viterbi, A.J. Error bounds for convolutional codes and an asymptotically optimum decoding algorithm. IEEE Trans. Inf. Theory 13, 260–269 (1967).
Rabiner, L.R. A tutorial on hidden Markov models and selected applications in speech recognition. Proc. IEEE 77, 257–286 (1989).
Cabili, M.N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 25, 1915–1927 (2011).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Beaudoing, E., Freier, S., Wyatt, J.R., Claverie, J.M. & Gautheret, D. Patterns of variant polyadenylation signal usage in human genes. Genome Res. 10, 1001–1010 (2000).
Durbin, R., Eddy, S.R., Krogh, A. & Mitchison, G.J. Biological Sequence Analysis: Probabilistic Models of Proteins and Nucleic Acids (Cambridge University Press, Cambridge, UK, 1998).
Johnson, J.W. A heuristic method for estimating the relative weight of predictor variables in multiple regression. Multivariate Behav. Res. 35, 1–19 (2000).
Acknowledgements
Research reported in this publication was supported by NIGMS (National Institute of General Medical Sciences) and NHGRI (National Human Genome Research Institute) grants from the US National Institutes of Health under award numbers GM25232 to J.T.L. and HG0070707 to A.S. and J.T.L., respectively. The content is solely the responsibility of the authors and does not necessarily represent the official views of the US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
L.J.C. and J.T.L. designed the experiments. L.J.C. and C.T.W. produced the data sets. A.L.M. designed and implemented software for data analysis. A.L.M., L.J.C., A.S., J.T.L. and C.G.D. analyzed the data and interpreted the results. L.J.C., A.L.M., A.S., J.T.L. and C.G.D. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Tables 1 and 2. (PDF 8931 kb)
Supplementary Data Set
A unified architecture of initiation regions at mammalian promoters and enhancers. (ZIP 2685 kb)
Rights and permissions
About this article
Cite this article
Core, L., Martins, A., Danko, C. et al. Analysis of nascent RNA identifies a unified architecture of initiation regions at mammalian promoters and enhancers. Nat Genet 46, 1311–1320 (2014). https://doi.org/10.1038/ng.3142
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3142
This article is cited by
-
Transcription directionality is licensed by Integrator at active human promoters
Nature Structural & Molecular Biology (2024)
-
Prioritizing genes associated with brain disorders by leveraging enhancer-promoter interactions in diverse neural cells and tissues
Genome Medicine (2023)
-
Decoding enhancer complexity with machine learning and high-throughput discovery
Genome Biology (2023)
-
eRNA co-expression network uncovers TF dependency and convergent cooperativity
Scientific Reports (2023)
-
RNA polymerase II dynamics shape enhancer–promoter interactions
Nature Genetics (2023)