Abstract
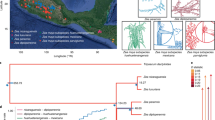
Domestication and plant breeding are ongoing 10,000-year-old evolutionary experiments that have radically altered wild species to meet human needs. Maize has undergone a particularly striking transformation. Researchers have sought for decades to identify the genes underlying maize evolution1,2, but these efforts have been limited in scope. Here, we report a comprehensive assessment of the evolution of modern maize based on the genome-wide resequencing of 75 wild, landrace and improved maize lines3. We find evidence of recovery of diversity after domestication, likely introgression from wild relatives, and evidence for stronger selection during domestication than improvement. We identify a number of genes with stronger signals of selection than those previously shown to underlie major morphological changes4,5. Finally, through transcriptome-wide analysis of gene expression, we find evidence both consistent with removal of cis-acting variation during maize domestication and improvement and suggestive of modern breeding having increased dominance in expression while targeting highly expressed genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Briggs, W.H., McMullen, M.D., Gaut, B.S. & Doebley, J. Linkage mapping of domestication loci in a large maize-teosinte backcross resource. Genetics 177, 1915–1928 (2007).
Wright, S.I. et al. The effects of artificial selection of the maize genome. Science 308, 1310–1314 (2005).
Chia, J.-M. et al. Maize HapMap2 identifies extant variation from a genome in flux. Nat. Genet. published online, doi:10.1038/ng.2313 (3 June 2012).
Doebley, J., Stec, A. & Hubbard, L. The evolution of apical dominance in maize. Nature 386, 485–488 (1997).
Wang, H. et al. The origin of the naked grains of maize. Nature 436, 714–719 (2005).
Piperno, D.R., Ranere, A.J., Holst, I., Iriarte, J. & Dickau, R. Starch grain and phytolith evidence for early ninth millennium BP maize from the Central Balsas River Valley, Mexico. Proc. Natl. Acad. Sci. USA 106, 5019–5024 (2009).
Matsuoka, Y. et al. A single domestication for maize shown by multilocus microsatellite genotyping. Proc. Natl. Acad. Sci. USA 99, 6080–6084 (2002).
van Heerwaarden, J. et al. Genetic signals of origin, spread, and introgression in a large sample of maize landraces. Proc. Natl. Acad. Sci. USA 108, 1088–1092 (2011).
Caicedo, A.L. et al. Genome-wide patterns of nucleotide polymorphism in domesticated rice. PLoS Genet. 3, 1745–1756 (2007).
Lam, H.M. et al. Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat. Genet. 42, 1053–1059 (2010).
Wilkes, H.G. Teosinte: The Closest Relative of Maize (The Bussey Institute of Harvard University, Cambridge, Massachusetts, 1967).
Chen, H., Patterson, N. & Reich, D. Population differentiation as a test for selective sweeps. Genome Res. 20, 393–402 (2010).
Fang, Z. et al. Megabase-scale inversion polymorphism in the wild ancestor of maize. Genetics published online, doi:10.1534/genetics.112.138578 (27 April 2012).
Purugganan, M.D. & Fuller, D.Q. Archaeological data reveal slow rates of evolution during plant domestication. Evolution 65, 171–183 (2011).
Olsen, K.M. et al. Selection under domestication: evidence for a sweep in the rice waxy genomic region. Genetics 173, 975–983 (2006).
Palaisa, K., Morgante, M., Tingey, S. & Rafalski, A. Long-range patterns of diversity and linkage disequilibrium surrounding the maize Y1 gene are indicative of an asymmetric selective sweep. Proc. Natl. Acad. Sci. USA 101, 9885–9890 (2004).
Camus-Kulandaivelu, L. et al. Maize adaptation to temperate climate: relationship between population structure and polymorphism in the Dwarf8 gene. Genetics 172, 2449–2463 (2006).
Brown, P.J. et al. Distinct genetic architectures for male and female inflorescence traits of maize. PLoS Genet. 7, e1002383 (2011).
Buckler, E.S. et al. The genetic architecture of maize flowering time. Science 325, 714–718 (2009).
Gallais, A. & Hirel, B. An approach to the genetics of nitrogen use efficiency in maize. J. Exp. Bot. 55, 295–306 (2004).
Jackson, D. & Hake, S. Control of phyllotaxy in maize by the abphyl1 gene. Development 126, 315–323 (1999).
Gualberti, G. et al. Mutations in the Dof zinc finger genes DAG2 and DAG1 influence with opposite effects the germination of Arabidopsis seeds. Plant Cell 14, 1253–1263 (2002).
Sasaki, A. et al. Green revolution: a mutant gibberellin-synthesis gene in rice—new insight into the rice variant that helped to avert famine over thirty years ago. Nature 416, 701–702 (2002).
Wang, Y. et al. Molecular tailoring of farnesylation for plant drought tolerance and yield protection. Plant J. 43, 413–424 (2005).
Eastmond, P.J. SUGAR-DEPENDENT1 encodes a patatin domain triacylglycerol lipase that initiates storage oil breakdown in germinating Arabidopsis seeds. Plant Cell 18, 665–675 (2006).
Sekhon, R.S. et al. Genome-wide atlas of transcription during maize development. Plant J. 66, 553–563 (2011).
Hufford, K.M., Canaran, P., Ware, D.H., McMullen, M.D. & Gaut, B.S. Patterns of selection and tissue-specific expression among maize domestication and crop improvement loci. Plant Physiol. 144, 1642–1653 (2007).
Stupar, R.M. et al. Gene expression analyses in maize inbreds and hybrids with varying levels of heterosis. BMC Plant Biol. 8, 33 (2008).
Beadle, G.W. Teosinte and the origin of maize. J. Hered. 30, 245–247 (1939).
Ryu, C.H. et al. OsMADS50 and OsMADS56 function antagonistically in regulating long day (LD)-dependent flowering in rice. Plant Cell Environ. 32, 1412–1427 (2009).
Lo, S.F. et al. A novel class of gibberellin 2–oxidases control semidwarfism, tillering, and root development in rice. Plant Cell 20, 2603–2618 (2008).
Schnable, P.S. et al. The B73 maize genome: complexity, diversity, and dynamics. Science 326, 1112–1115 (2009).
McMullen, M.D. et al. Genetic properties of the maize nested association mapping population. Science 325, 737–740 (2009).
Wolfgruber, T.K. et al. Maize centromere structure and evolution: sequence analysis of centromeres 2 and 5 reveals dynamic loci shaped primarily by retrotransposons. PLoS Genet. 5, e1000743 (2009).
Thornton, K. libsequence: a C++ class library for evolutionary genetic analysis. Bioinformatics 19, 2325–2327 (2003).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Hudson, R.R., Boos, D.D. & Kaplan, N.L. A statistical test for detecting geographic subdivision. Mol. Biol. Evol. 9, 138–151 (1992).
Hudson, R.R. Two-locus sampling distributions and their application. Genetics 159, 1805–1817 (2001).
Swanson-Wagner, R.A. et al. Pervasive gene content variation and copy number variation in maize and its undomesticated progenitor. Genome Res. 20, 1689–1699 (2010).
Irizarry, R.A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003).
Springer, N.M. et al. Maize inbreds exhibit high levels of copy number variation (CNV) and presence/absence variation (PAV) in genome content. PLoS Genet. 5, e1000734 (2010).
Lawrence, C.J., Dong, O.F., Polacco, M.L., Seigfried, T.E. & Brendel, V. MaizeGDB, the community database for maize genetics and genomics. Nucleic Acids Res. 32, D393–D397 (2004).
Davison, A.C. & Hinkley, D.V. Bootstrap Methods and Their Application (Cambridge University Press, New York, 1997).
Acknowledgements
The authors would like to thank T. Kono, S. Watson and M. Watson for photographs of inflorescences, P. Brown for help with QTL delineation, B.S. Gaut, A.M. Gonzales and two anonymous reviewers for comments on an earlier version of the manuscript and M. Grote for statistical advice. This work was supported by funding to the maize diversity project from the US National Science Foundation (NSF; IOS-0820619 to E.S.B., J.D. and M.D.M.) and USDA-ARS (to E.S.B., M.D.M. and D.W.), as well as from USDA Hatch Funds (to P.T. and N.M.S.), the Chinese 973 program (2007CB815701 to J.W.), the Chinese Ministry of Agriculture 984 program (2010-Z13 to G.Z.), the Shenzhen Municipal Government Basic Research Program (to J.W.), the US DOE Great Lakes Bioenergy Research Center (DOE Office of Science; BER DE-FC02-07ER64494), the Office of Science of the US DOE (contract DE-AC02-05CH11231 to the US DOE Joint Genome Institute) and by grants from the US NSF (IOS-0922703 to J.R.-I.) and the USDA–National Institute of Food and Agriculture (2009-01864 to J.R.-I.).
Author information
Authors and Affiliations
Contributions
J.D., M.D.M., E.S.B., D.W. and J.R.-I. designed the project. M.B.H., J.v.H., T.P. and J.R.-I. performed most data analyses. J.D. developed wild and landrace inbred lines. E.S.B., S.M.K., J.L., M.D.M. and D.W. contributed sequence data for inbred maize and parviglumis. K.E.G. and R.J.E. developed libraries and managed sequencing for inbred maize and parviglumis. X.X., S.Y., J.W. and G.Z. directed sequencing for landrace maize, mexicana and Tripsacum. E.S.B., J.R.-I., D.W. and X.X. directed bioinformatics analyses. J.-M.C. and C.S. performed read mapping, SNP calling and annotation, and analysis of coding sequence. E.S.B., J.-M.C. and J.C.G. performed quality control filtering of SNPs. N.M.S., R.A.S.-W. and P.T. generated Nimblegen expression data for maize and parviglumis. S.M.K. provided early access expression data. L.M.S. reanalyzed QTL data for domestication traits. R.A.C. analyzed site frequency spectra. M.B.H., J.v.H., T.P., P.L.M. and J.R.-I. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–5, 8 and 9 and Supplementary Figures 1–15 (PDF 10057 kb)
Supplementary Table 6
Domestication candidates (XLSX 189 kb)
Supplementary Table 7
Improvement candidates (XLSX 163 kb)
Rights and permissions
About this article
Cite this article
Hufford, M., Xu, X., van Heerwaarden, J. et al. Comparative population genomics of maize domestication and improvement. Nat Genet 44, 808–811 (2012). https://doi.org/10.1038/ng.2309
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2309
This article is cited by
-
Intergrative metabolomic and transcriptomic analyses reveal the potential regulatory mechanism of unique dihydroxy fatty acid biosynthesis in the seeds of an industrial oilseed crop Orychophragmus violaceus
BMC Genomics (2024)
-
Genomic insight into the origin, domestication, dispersal, diversification and human selection of Tartary buckwheat
Genome Biology (2024)
-
Cytoplasmic genome contributions to domestication and improvement of modern maize
BMC Biology (2024)
-
Identifying yield-related genes in maize based on ear trait plasticity
Genome Biology (2023)
-
Fine-Scale analysis of both wild and cultivated horned galls provides insight into their quality differentiation
BMC Plant Biology (2023)