Abstract
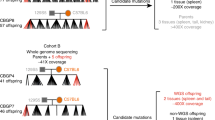
J.B.S. Haldane proposed in 1947 that the male germline may be more mutagenic than the female germline1. Diverse studies have supported Haldane's contention of a higher average mutation rate in the male germline in a variety of mammals, including humans2,3. Here we present, to our knowledge, the first direct comparative analysis of male and female germline mutation rates from the complete genome sequences of two parent-offspring trios. Through extensive validation, we identified 49 and 35 germline de novo mutations (DNMs) in two trio offspring, as well as 1,586 non-germline DNMs arising either somatically or in the cell lines from which the DNA was derived. Most strikingly, in one family, we observed that 92% of germline DNMs were from the paternal germline, whereas, in contrast, in the other family, 64% of DNMs were from the maternal germline. These observations suggest considerable variation in mutation rates within and between families.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Haldane, J.B.S. The mutation rate of the gene for haemophilia, and its segregation ratios in males and females. Ann. Eugen. 13, 262–271 (1947).
Bohossian, H.B., Skaletsky, H. & Page, D.C. Unexpectedly similar rates of nucleotide substitution found in male and female hominids. Nature 406, 622–625 (2000).
Taylor, J., Tyekucheva, S., Zody, M., Chiaromonte, F. & Makova, K.D. Strong and weak male mutation bias at different sites in the primate genomes: insights from the human-chimpanzee comparison. Mol. Biol. Evol. 23, 565–573 (2006).
Awadalla, P. et al. Direct measure of the de novo mutation rate in autism and schizophrenia cohorts. Am. J. Hum. Genet. 87, 316–324 (2011).
Gauthier, J. et al. De novo mutations in the gene encoding the synaptic scaffolding protein SHANK3 in patients ascertained for schizophrenia. Proc. Natl. Acad. Sci. USA 107, 7863–7868 (2010).
Lee, C., Iafrate, A.J. & Brothman, A.R. Copy number variations and clinical cytogenetic diagnosis of constitutional disorders. Nat. Genet. 39, S48–S54 (2007).
Roach, J.C. et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328, 636–639 (2010).
Xue, Y. et al. Human Y chromosome base-substitution mutation rate measured by direct sequencing in a deep-rooting pedigree. Curr. Biol. 19, 1453–1457 (2009).
Crow, J.F. How much do we know about spontaneous human mutation rates? Environ. Mol. Mutagen. 21, 122–129 (1993).
Haldane, J.B.S. The rate of spontaneous mutation of a human gene. J. Genet. 31, 317–326 (1935).
Kondrashov, A.S. Direct estimates of human per nucleotide mutation rates at 20 loci causing Mendelian diseases. Hum. Mutat. 21, 12–27 (2003).
Kondrashov, A.S. & Crow, J.F. A molecular approach to estimating the human deleterious mutation rate. Hum. Mutat. 2, 229–234 (1993).
Lynch, M. Rate, molecular spectrum, and consequences of human mutation. Proc. Natl. Acad. Sci. USA 107, 961–968 (2010).
Nachman, M.W. & Crowell, S.L. Estimate of the mutation rate per nucleotide in humans. Genetics 156, 297–304 (2000).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Chen, F.C. & Li, W.H. Genomic divergences between humans and other hominoids and the effective population size of the common ancestor of humans and chimpanzees. Am. J. Hum. Genet. 68, 444–456 (2001).
Lynch, M. Evolution of the mutation rate. Trends Genet. 26, 345–352 (2010).
Cooper, G.M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005).
Acknowledgements
We would like to thank A. Kernytsky, G. McVean, T. Massingham, J. Thorne, J. Hussin, A. Motsinger, Coriell Cell Repositories and members of the 1000 Genomes analysis group for their help and support. D.F.C., S.J.L., Y.Z., C.T. and M.E.H. were funded by the Wellcome Trust (grant number 077014/Z/05/Z). J.E.M.K., F.C., Y.I., M.Z., G.A.R. and P.A. were funded by the Ministry of Development, Exploration and Innovation (grant number PSR-SIIRI-195) in Quebec and a Genome Quebec Award for Population and Medical Genomics to P.A.
Author information
Authors and Affiliations
Consortia
Contributions
M.E.H. and P.A. conceived the study. D.F.C., J.E.M.K., M.A.D., M.D., R.C., E.A.S. and P.A. developed statistical methodologies. D.F.C., J.E.M.K., M.A.D., C.L.H., K.V.G., E.A.S., M.E.H. and P.A. analyzed the data. F.C., Y.I., G.A.R., C.T., M.Z., S.J.L. and Y.Z. generated validation data. D.F.C., P.A. and M.E.H. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The author declare no competing financial interests.
Additional information
A full list of members is provided in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 2–3 and Supplementary Note. (PDF 1320 kb)
Supplementary Table 1
Validation and haplotyping results (XLS 3486 kb)
Rights and permissions
About this article
Cite this article
the 1000 Genomes Project. Variation in genome-wide mutation rates within and between human families. Nat Genet 43, 712–714 (2011). https://doi.org/10.1038/ng.862
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.862
This article is cited by
-
The unusual gene architecture of polyubiquitin is created by dual-specific splice sites
Genome Biology (2024)
-
Statistical methods for assessing the effects of de novo variants on birth defects
Human Genomics (2024)
-
Meta-analysis of 46,000 germline de novo mutations linked to human inherited disease
Human Genomics (2024)
-
MECP2 germline mosaicism plays an important part in the inheritance of Rett syndrome: a study of MECP2 germline mosaicism in males
BMC Medicine (2023)
-
Evolution of the germline mutation rate across vertebrates
Nature (2023)