Abstract
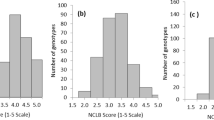
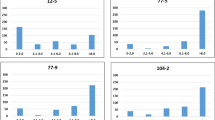
Nested association mapping (NAM) offers power to resolve complex, quantitative traits to their causal loci. The maize NAM population, consisting of 5,000 recombinant inbred lines (RILs) from 25 families representing the global diversity of maize, was evaluated for resistance to southern leaf blight (SLB) disease. Joint-linkage analysis identified 32 quantitative trait loci (QTLs) with predominantly small, additive effects on SLB resistance. Genome-wide association tests of maize HapMap SNPs were conducted by imputing founder SNP genotypes onto the NAM RILs. SNPs both within and outside of QTL intervals were associated with variation for SLB resistance. Many of these SNPs were within or near sequences homologous to genes previously shown to be involved in plant disease resistance. Limited linkage disequilibrium was observed around some SNPs associated with SLB resistance, indicating that the maize NAM population enables high-resolution mapping of some genome regions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Holley, R.N. & Goodman, M.M. New sources of resistance to southern corn leaf blight from tropical hybrid maize derivatives. Plant Dis. 73, 562–564 (1989).
Bent, A.F. & Mackey, D. Elicitors, effectors, and R genes: The new paradigm and a lifetime supply of questions. Annu. Rev. Phytopathol. 45, 399–436 (2007).
Broglie, K. et al. Polynucleotides and methods for making plants resistant to fungal pathogens. United States Patent number 7,619,133 (2009).
Fukuoka, S. et al. Loss of function of a proline-containing protein confers durable disease resistance in rice. Science 325, 998–1001 (2009).
Fu, D. et al. A kinase-start gene confers temperature-dependent resistance to wheat stripe rust. Science 323, 1357–1360 (2009).
Manosalva, P.M. et al. A germin-like protein gene family functions as a complex quantitative trait locus conferring broad-spectrum disease resistance in rice. Plant Physiol. 149, 286–296 (2009).
Poland, J.A., Balint-Kurti, P.J., Wisser, R.J., Pratt, R.C. & Nelson, R.J. Shades of gray: The world of quantitative disease resistance. Trends Plant Sci. 14, 21–29 (2009).
Krattinger, S.G. et al. A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323, 1360–1363 (2009).
McMullen, M.D. et al. Genetic properties of the maize nested association mapping population. Science 325, 737–740 (2009).
Buckler, E.S. et al. The genetic architecture of maize flowering time. Science 325, 714–718 (2009).
Yu, J., Holland, J.B., McMullen, M. & Buckler, E.S. Genetic design and statistical power of nested association mapping in maize. Genetics 178, 539–551 (2008).
Gore, M.A. et al. A first-generation haplotype map of maize. Science 326, 1115–1117 (2009).
Tian, F. et al. Genome wide association identifies genes that have contributed to maize improvement. Nat. Genet. advance online publication, doi:10.1038/ng.746 (9 January 2010).
Balint-Kurti, P.J. et al. Precise mapping of quantitative trait loci for resistance to southern leaf blight, caused by Cochliobolus heterostrophus race O, and flowering time using advanced intercross maize lines. Genetics 176, 645–657 (2007).
Wisser, R. The genetic architecture of disease resistance in maize: a synthesis of published studies. Phytopathology 96, 120–129 (2006).
Balint-Kurti, P.J. & Carson, M.L. Analysis of quantitative trait loci for resistance to southern leaf blight in juvenile maize. Phytopathology 96, 221–225 (2006).
Balint-Kurti, P.J. et al. Identification of quantitative trait loci for resistance to southern leaf blight and days to anthesis in a maize recombinant inbred line population. Phytopathology 96, 1067–1071 (2006).
Balint-Kurti, P.J. et al. Identification of quantitative trait loci for resistance to southern leaf blight and days to anthesis in two maize recombinant inbred line populations. Phytopathology 98, 315–320 (2008).
Zwonitzer, J. et al. Use of selection with recurrent backcrossing and QTL mapping to identify loci contributing to southern leaf blight resistance in a highly resistant maize line. Theor. Appl. Genet. 118, 911–925 (2009).
Zwonitzer, J.C. et al. Mapping resistance quantitative trait loci for three foliar diseases in a maize recombinant inbred line population-evidence for multiple disease resistance? Phytopathology 100, 72–79 (2010).
Valdar, W., Holmes, C.C., Mott, R. & Flint, J. Mapping in structured populations by resample model averaging. Genetics 182, 1263–1277 (2009).
Valdar, W. et al. Genome-wide genetic association of complex traits in heterogeneous stock mice. Nat. Genet. 38, 879–887 (2006).
Atwell, S. et al. Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465, 627–631 (2010).
Brachi, B. et al. Linkage and association mapping of Arabidopsis thaliana flowering time in nature. PLoS Genet. 6, e1000940 (2010).
Morris, E.R. & Walker, J.C. Receptor-like protein kinases: The keys to response. Curr. Opin. Plant Biol. 6, 339–342 (2003).
Nicaise, V., Roux, M. & Zipfel, C. Recent advances in PAMP-triggered immunity against bacteria: Pattern recognition receptors watch over and raise the alarm. Plant Physiol. 150, 1638–1647 (2009).
Dong, X. NPR1, all things considered. Curr. Opin. Plant Biol. 7, 547–552 (2004).
Cao, H., Glazebrook, J., Clarke, J.D., Volko, S. & Dong, X.N. The Arabidopsis NPR1 gene that controls systemic acquired resistance encodes a novel protein containing ankyrin repeats. Cell 88, 57–63 (1997).
Despres, C., DeLong, C., Glaze, S., Liu, E. & Fobert, P.-R. The Arabidopsis NPR1/NIM1 protein enhances the DNA binding activity of a subgroup of the TGA family of bZIP transcription factors. Plant Cell 12, 279–290 (2000).
Chern, M., Fitzgerald, H.A., Canlas, P.E., Navarre, D.A. & Ronald, P.C. Overexpression of a rice NPR1 homolog leads to constitutive activation of defense response and hypersensitivity to light. Mol. Plant Microbe Interact. 18, 511–520 (2005).
Chern, M., Canlas, P.E., Fitzgerald, H.A. & Ronald, P.C. Rice NRR, a negative regulator of disease resistance, interacts with Arabidopsis NPR1 and rice NH1. Plant J. 43, 623–635 (2005).
Mignotte, B. & Vayssiere, J.-L. Mitochondria and apoptosis. Eur. J. Biochem. 252, 1–15 (1998).
Sacco, M.A. et al. The cyst nematode SPRYSEC protein RBP-1 elicits Gpa2- and rangap2-dependent plant cell death. PLoS Pathog. 5, e1000564 (2009).
Wisser, R.J., Sun, Q., Hulbert, S.H., Kresovich, S. & Nelson, R.J. Identification and characterization of regions of the rice genome associated with broad-spectrum, quantitative disease resistance. Genetics 169, 2277–2293 (2005).
Maldonado, A.M., Doerner, P., Dixon, R.A., Lamb, C.J. & Cameron, R.K. A putative lipid transfer protein involved in systemic resistance signalling in Arabidopsis. Nature 419, 399–403 (2002).
Boyle, P. et al. The BTB/POZ domain of the Arabidopsis disease resistance protein NPR1 interacts with the repression domain of TGA2 to negate its function. Plant Cell 21, 3700–3713 (2009).
Kou, Y., Qiu, D., Wang, L., Li, X. & Wang, S. Molecular analyses of the rice tubby-like protein gene family and their response to bacterial infection. Plant Cell Rep. 28, 113–121 (2009).
Gutterson, N. & Reuber, T.L. Regulation of disease resistance pathways by AP2/ERF transcription factors. Curr. Opin. Plant Biol. 7, 465–471 (2004).
Taj, G., Kumar, A., Bansal, K.C. & Garg, G.K. Signaling components in programmed cell death: Molecular and functional similarities between animals and plants. in Recent Advances in Plant Biotechnology and Its Applications (eds. Kumar, A. & Sopory, S.) 391–409 (I.K. International Publishing, New Dehli, India, 2008).
Nawrath, C., Heck, S., Parinthawong, N. & Metraux, J.-P. Eds5, an essential component of salicylic acid-dependent signaling for disease resistance in Arabidopsis, is a member of the mate transporter family. Plant Cell 14, 275–286 (2002).
Zeng, L.-R. et al. Spotted leaf11, a negative regulator of plant cell death and defense, encodes a u-box/armadillo repeat protein endowed with E3 ubiquitin ligase activity. Plant Cell 16, 2795–2808 (2004).
Chern, M., Fitzgerald, H.A., Canlas, P.E., Navarre, D.A. & Ronald, P.C. Overexpression of a rice NPR1 homolog leads to constitutive activation of defense response and hypersensitivity to light. Mol. Plant Microbe Interact. 18, 511–520 (2005).
Zhou, N., Tootle, T.-L. & Glazebrook, J. Arabidopsis PAD3, a gene required for camalexin biosynthesis, encodes a putative cytochrome P450 monooxygenase. Plant Cell 11, 2419–2428 (1999).
Takemoto, D., Hayashi, M., Doke, N., Nishimura, M. & Kawakita, K. Molecular cloning of a defense-response-related cytochrome P450 gene from tobacco. Plant Cell Physiol. 40, 1232–1242 (1999).
Zhou, J., Loh, Y.-T., Bressan, R.A. & Martin, G.B. The tomato gene Pti1 encodes a serine/threonine kinase that is phosphorylated by Pto and is involved in the hypersensitive response. Cell 83, 925–935 (1995).
Loh, Y.T. & Martin, G.B. The Pto bacterial resistance gene and the Fen insecticide sensitivity gene encode functional protein kinases with serine/threonine specificity. Plant Physiol. 108, 1735–1739 (1995).
Vogel, J.P., Raab, T.K., Schiff, C. & Somerville, S.C. PMR6, a pectate lyase-like gene required for powdery mildew susceptibility in Arabidopsis. Plant Cell 14, 2095–2106 (2002).
Gu, Y.-Q. et al. Tomato transcription factors Pti4, Pti5, and Pti6 activate defense responses when expressed in Arabidopsis. Plant Cell 14, 817–831 (2002).
Shah, J. Lipids, lipases, and lipid-modifying enzymes in plant disease resistance. Annu. Rev. Phytopathol. 43, 229–260 (2005).
Zhang, Y., Fan, W., Kinkema, M., Li, X. & Dong, X. Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene. Proc. Natl. Acad. Sci. USA 96, 6523–6528 (1999).
Schnable, P.S. et al. The B73 maize genome: complexity, diversity, and dynamics. Science 326, 1112–1115 (2009).
Liu, K. et al. Genetic structure and diversity among maize inbred lines as inferred from DNA microsatellites. Genetics 165, 2117–2128 (2003).
Gilmour, A.R., Cullis, B., Gogel, B., Welham, S.J. & Thompson, R. Asreml User's Guide. Release 2.0 (VSN International, Hemel Hempstead, UK, 2005).
Hooker, A.L. Southern leaf blight of corn present status and future prospects. J. Environ. Qual. 1, 244–248 (1972).
Haldane, J.B.S. & Waddington, C.H. Inbreeding and linkage. Genetics 16, 357–374 (1931).
R Development Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, Austria, 2010).
Acknowledgements
We thank M. Eller, D. Rhyne, D. Stephens, G. Van Esbroek, C. Herring and the staffs of the North Carolina Central Crops Research Station and 27 Farms, Homestead, Florida, USA, for assistance in the field. We also thank J. Green and C.-R. Shyu for providing the photographs for Supplementary Figure 1. This research was supported by the US National Science Foundation (DBI-0321467 and IOS-0820619) and funds were provided by USDA-ARS. K.L.K. is supported by Pioneer Hi-Bred International.
Author information
Authors and Affiliations
Contributions
E.S.B., M.D.M., S.K. and J.B.H. developed the NAM population. K.L.K., R.J.W., A.R.B., M.A.O.-R., J.C.Z., P.J.B.-K. and J.B.H. conducted the field experiments. E.S.B. and D.W. created HapMap data. P.J.B. and E.S.B. developed GWAS methods and software. K.L.K. and J.B.H. analyzed the data. K.L.K., R.J.W., P.J.B.-K. and J.B.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1–9 and Supplementary Note (PDF 1624 kb)
Supplementary Table 7
Information on 245 significantly associated SNPs. (XLS 138 kb)
Rights and permissions
About this article
Cite this article
Kump, K., Bradbury, P., Wisser, R. et al. Genome-wide association study of quantitative resistance to southern leaf blight in the maize nested association mapping population. Nat Genet 43, 163–168 (2011). https://doi.org/10.1038/ng.747
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.747
This article is cited by
-
Diploid Interspecific Recombinant Inbred Lines for Genetic Mapping in Potato
American Journal of Potato Research (2024)
-
A genome-wide association study for resistance to Fusarium wilt (Fusarium oxysporum f. sp. vasinfectum) race 4 in diploid cotton (Gossypium arboreum) and resistance transfer to tetraploid Gossypium hirsutum
Molecular Genetics and Genomics (2024)
-
Hybrid allele-specific ChIP-seq analysis identifies variation in brassinosteroid-responsive transcription factor binding linked to traits in maize
Genome Biology (2023)
-
Nested association mapping population in crops: current status and future prospects
Journal of Crop Science and Biotechnology (2023)
-
Development of southern corn leaf blight (SCLB) resistant and high-popping volume composite popcorn using phenotypic and marker-assisted selection (MAS)
Cereal Research Communications (2023)