Abstract
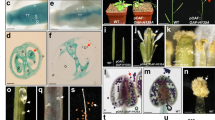
It is commonly thought that deep phylogenetic conservation of plant microRNAs (miRNAs) and their targets1,2 indicates conserved regulatory functions. We show that the blind (bl) mutant of Petunia hybrida3 and the fistulata (fis) mutant of Antirrhinum majus4,5, which have similar homeotic phenotypes, are recessive alleles of two homologous miRNA-encoding genes. The BL and FIS genes control the spatial restriction of homeotic class C genes6,7 to the inner floral whorls, but their ubiquitous early floral expression patterns are in contradiction with a potential role in patterning C gene expression. We provide genetic evidence for the unexpected function of the MIRFIS and MIRBL genes in the center of the flower and propose a dynamic mechanism underlying their regulatory role. Notably, Arabidopsis thaliana, a more distantly related species, also contains this miRNA module but does not seem to use it to confine early C gene expression to the center of the flower.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Axtell, M.J. & Bartel, D.P. Antiquity of microRNAs and their targets in land plants. Plant Cell 17, 1658–1673 (2005).
Baulcombe, D. RNA silencing in plants. Nature 431, 356–363 (2004).
Vallade, J., Maizonnier, D. & Cornu, A. La morphogenese florale chez le petunia. I. Analyze d'un mutant à corolle staminée. Can. J. Bot. 65, 761–764 (1987).
McSteen, P.C.M., Vincent, C.A., Doyle, S., Carpenter, R. & Coen, E.S. Control of floral homeotic gene-expression and organ morphogenesis in Antirrhinum. Development 125, 2359–2369 (1998).
Motte, P., Saedler, H. & Schwarz-Sommer, Z. STYLOSA and FISTULATA: regulatory components of the homeotic control of Antirrhinum floral organogenesis. Development 125, 71–84 (1998).
Davies, B., Cartolano, M. & Schwarz-Sommer, Z. Flower development: The Antirrhinum perspective. Adv. Bot. Res. Inc. Adv. Plant Pathol. 44, 278–319 (2006).
Jack, T. Molecular and genetic mechanisms of floral control. Plant Cell 16, S1–S17 (2004).
Navarro, C. et al. Molecular and genetic interactions between STYLOSA and GRAMINIFOLIA in the control of Antirrhinum vegetative and reproductive development. Development 131, 3649–3659 (2004).
Kater, M.M. et al. Multiple AGAMOUS homologs from cucumber and petunia differ in their ability to induce reproductive organ fate. Plant Cell 10, 171–182 (1998).
Davies, B. et al. PLENA and FARINELLI: redundancy and regulatory interactions between two Antirrhinum MADS-box factors controlling flower development. EMBO J. 18, 4023–4034 (1999).
Maes, T. et al. Petunia Ap2-like genes and their role in flower and seed development. Plant Cell 13, 229–244 (2001).
Keck, E., McSteen, P., Carpenter, R. & Coen, E. Separation of genetic functions controlling organ identity in flowers. EMBO J. 22, 1058–1066 (2003).
Jones-Rhoades, M.W., Bartel, D.P. & Bartel, B. MicroRNAs and their regulatory roles in plants. Annu. Rev. Plant Biol. 57, 19–53 (2006).
Xie, Z. et al. Expression of Arabidopsis MIRNA genes. Plant Physiol. 138, 2145–2154 (2005).
Gusmaroli, G., Tonellia, C. & Mantovani, R. Regulation of the CCAAT-Binding NF-Y subunits in Arabidopsis thaliana. Gene 283, 41–48 (2002).
Schwab, R. et al. Specific effects of microRNAs on the plant transcriptome. Dev. Cell 8, 517–527 (2005).
Hong, R.L., Hamaguchi, L., Busch, M.A. & Weigel, D. Regulatory elements of the floral homeotic gene AGAMOUS identified by phylogenetic footprinting and shadowing. Plant Cell 15, 1296–1309 (2003).
Nikovics, K. et al. The balance between the MIR164A and CUC2 genes controls leaf margin serration in Arabidopsis. Plant Cell 18, 2929–2945 (2006).
Hornstein, E. & Shomron, N. Canalization of development by microRNAs. Nat. Genet. 38, S20–S24 (2006).
Chen, X. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303, 2022–2025 (2004).
Gandikota, M. et al. The miRNA156/157 recognition element in the 3′ UTR of the Arabidopsis SBP box gene SPL3 prevents early flowering by translational inhibition in seedlings. Plant J. 49, 683–693 (2007).
Ingram, C.G. et al. Dual role for fimbriata in regulating floral homeotic genes and cell division in Antirrhinum. EMBO J. 16, 6521–6534 (1997).
Gomez-Mena, C., de Folter, S., Costa, M.M.R., Angenent, G.C. & Sablowski, R. Transcriptional program controlled by the floral homeotic gene AGAMOUS during early organogenesis. Development 132, 429–438 (2005).
Ingram, G.C. et al. Parallels between unusual floral organs and fimbriata, genes controlling flower development in Arabidopsis and Antirrhinum. Plant Cell 7, 1501–1510 (1995).
Shigyo, M., Hasebe, M. & Ito, M. Molecular evolution of the AP2 subfamily. Gene 366, 256–265 (2006).
Mlotshwa, S., Yang, Z., Kim, Y. & Chen, X. Floral patterning defects induced by Arabidopsis APETALA2 and microRNA172 expression in Nicotiana benthamiana. Plant Mol. Biol. 61, 781–793 (2006).
Van den Broeck, D. et al. Transposon display identifies individual transposable elements in high copy number lines. Plant J. 13, 121–129 (1998).
Valoczi, A., Varallyay, E., Kauppinen, S., Burgyan, J. & Havelda, Z. Spatio-temporal accumulation of microRNAs is highly coordinated in developing plant tissues. Plant J. 47, 140–151 (2006).
Acknowledgements
We thank J. Burgyan for information on in situ analyses with miRNA; J. Stuurman and J. Moore (University of Berne) for the bl-2 allele and B. Davies, P. Huijser, H. Saedler and P. Schulze-Lefert for discussions and comments. This work was supported in part by a grant from the Deutsche Forschungsgemeinschaft/SFB572 (Z.S.-S.).
Author information
Authors and Affiliations
Contributions
All authors contributed to the experiments, which were conceptually designed mainly by M.V. and Z.S.-S. Z.S.-S. wrote the manuscript, with support from M.V., M.C. and T.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Neighbor-joining tree of Antirrhinum, Petunia and Arabidopsis NF-YA family members. (PDF 957 kb)
Supplementary Fig. 2
Detection of the in situ expression patterns of Antirrhinum NF-YA genes by different methods. (PDF 117 kb)
Supplementary Fig. 3
RNA blot analysis of microRNA expression, and analysis of gene expression by qRT-PCR. (PDF 106 kb)
Supplementary Table 1
Position of cleavage sites in the MRE of Antirrhinum and Petunia NF-YA transcripts. (PDF 28 kb)
Supplementary Table 2
Oligonucleotide sequences. (PDF 60 kb)
Rights and permissions
About this article
Cite this article
Cartolano, M., Castillo, R., Efremova, N. et al. A conserved microRNA module exerts homeotic control over Petunia hybrida and Antirrhinum majus floral organ identity. Nat Genet 39, 901–905 (2007). https://doi.org/10.1038/ng2056
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng2056
This article is cited by
-
Ectopic expression of AtNF-YA6-VP16 in petals results in a novel petal phenotype in Torenia fournieri
Planta (2022)
-
Identification of microRNAs from transcriptome data in gurmar (Gymnema sylvestre)
Horticulture, Environment, and Biotechnology (2019)
-
Multiple microRNAs Regulate the Floral Development and Sex Differentiation in the Dioecious Cucurbit Coccinia grandis (L.) Voigt
Plant Molecular Biology Reporter (2019)
-
A miR172 target-deficient AP2-like gene correlates with the double flower phenotype in roses
Scientific Reports (2018)
-
Analysis of microRNA reveals cleistogamous and chasmogamous floret divergence in dimorphic plant
Scientific Reports (2018)