Abstract
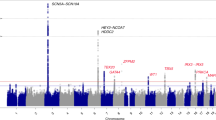
Extremes of the electrocardiographic QT interval, a measure of cardiac repolarization, are associated with increased cardiovascular mortality. We identified a common genetic variant influencing this quantitative trait through a genome-wide association study on 200 subjects at the extremes of a population-based QT interval distribution of 3,966 subjects from the KORA cohort in Germany, with follow-up screening of selected markers in the remainder of the cohort. We validated statistically significant findings in two independent samples of 2,646 subjects from Germany and 1,805 subjects from the US Framingham Heart Study. This genome-wide study identified NOS1AP (CAPON), a regulator of neuronal nitric oxide synthase, as a new target that modulates cardiac repolarization. Approximately 60% of subjects of European ancestry carry at least one minor allele of the NOS1AP genetic variant, which explains up to 1.5% of QT interval variation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Busjahn, A. et al. QT interval is linked to 2 long-QT syndrome loci in normal subjects. Circulation 99, 3161–3164 (1999).
Carter, N. et al. QT interval in twins. J. Hum. Hypertens. 14, 389–390 (2000).
Newton-Cheh, C. et al. QT interval is a heritable quantitative trait with evidence of linkage to chromosome 3 in a genome-wide linkage analysis: The Framingham Heart Study. Heart Rhythm 2, 277–284 (2005).
Schouten, E.G. et al. QT interval prolongation predicts cardiovascular mortality in an apparently healthy population. Circulation 84, 1516–1523 (1991).
Dekker, J.M., Schouten, E.G., Klootwijk, P., Pool, J. & Kromhout, D. Association between QT interval and coronary heart disease in middle-aged and elderly men. The Zutphen Study. Circulation 90, 779–785 (1994).
Elming, H. et al. The prognostic value of the QT interval and QT interval dispersion in all-cause and cardiac mortality and morbidity in a population of Danish citizens. Eur. Heart J. 19, 1391–1400 (1998).
Sharp, D.S., Masaki, K., Burchfiel, C.M., Yano, K. & Schatz, I.J. Prolonged QTc interval, impaired pulmonary function, and a very lean body mass jointly predict all-cause mortality in elderly men. Ann. Epidemiol. 8, 99–106 (1998).
de Bruyne, M.C. et al. Prolonged QT interval predicts cardiac and all-cause mortality in the elderly. The Rotterdam Study. Eur. Heart J. 20, 278–284 (1999).
Okin, P.M. et al. Assessment of QT interval and QT dispersion for prediction of all-cause and cardiovascular mortality in American Indians: the Strong heart study. Circulation 101, 61–66 (2000).
Dekker, J.M., Crow, R.S., Hannan, P.J., Schouten, E.G. & Folsom, A.R. Heart rate-corrected QT interval prolongation predicts risk of coronary heart disease in black and white middle-aged men and women: the ARIC study. J. Am. Coll. Cardiol. 43, 565–571 (2004).
Priori, S.G. & Napolitano, C. Genetics of cardiac arrhythmias and sudden cardiac death. Ann. NY Acad. Sci. 1015, 96–110 (2004).
Yang, P. et al. Allelic variants in long-QT disease genes in patients with drug-associated torsades de pointes. Circulation 105, 1943–1948 (2002).
Splawski, I. et al. Variant of SCN5A sodium channel implicated in risk of cardiac arrhythmia. Science 297, 1333–1336 (2002).
Perz, S. et al. Does computerized ECG analysis provide sufficiently consistent QT interval estimates for genetic research? in Analysis of Biomedical Signals and Images Vol. 17 (eds. Jan, J., Kozumplik, J. and Provaznik, I.) 47–49 (VUTIUM Press, Brno, Czech Republic, 2004).
Emison, E.S. et al. A common sex-dependent mutation in a RET enhancer underlies Hirschsprung disease risk. Nature 434, 857–863 (2005).
Klein, R.J. et al. Complement factor H polymorphism in age-related macular degeneration. Science 308, 385–389 (2005).
Haines, J.L. et al. Complement factor H variant increases the risk of age-related macular degeneration. Science 308, 419–421 (2005).
Edwards, A.O. et al. Complement factor H polymorphism and age-related macular degeneration. Science 308, 421–424 (2005).
Robertson, A. The nature of quantitative genetic variation. in Heritage from Mendel (ed. Brink, R.A.) 265–280 (Univ. of Wisconsin Press, Madison, Wisconsin, 1967).
Risch, N.J. & Zhang, H. Mapping quantitative trait loci with extreme discordant sib pairs: sampling considerations. Am. J. Hum. Genet. 58, 836–843 (1996).
Cohen, J.C. et al. Multiple rare alleles contribute to low plasma levels of HDL cholesterol. Science 305, 869–872 (2004).
Pfeufer, A. et al. Common variants in myocardial ion channel genes modify the QT interval in the general population: results from the KORA study. Circ. Res. 96, 693–701 (2005).
Bezzina, C.R. et al. A common polymorphism in KCNH2 (HERG) hastens cardiac repolarization. Cardiovasc. Res. 59, 27–36 (2003).
Satagopan, J.M., Venkatraman, E.S. & Begg, C.B. Two-stage designs for gene-disease association studies with sample size constraints. Biometrics 60, 589–597 (2004).
Boerwinkle, E., Chakraborty, R. & Sing, C.F. The use of measured genotype information in the analysis of quantitative phenotypes in man. I. Models and analytical methods. Ann. Hum. Genet. 50, 181–194 (1986).
Page, G.P. & Amos, C.I. Comparison of linkage-disequilibrium methods for localization of genes influencing quantitative traits in humans. Am. J. Hum. Genet. 64, 1194–1205 (1999).
Skol, A.D., Scott, L.J., Abecasis, G.R. & Boehnke, M. Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat. Genet. 38, 209–213 (2006).
Jaffrey, S.R., Snowman, A.M., Eliasson, M.J., Cohen, N.A. & Snyder, S.H. CAPON: a protein associated with neuronal nitric oxide synthase that regulates its interactions with PSD95. Neuron 20, 115–124 (1998).
De Jongh, K.S., Warner, C. & Catterall, W.A. Subunits of purified calcium channels. Alpha 2 and delta are encoded by the same gene. J. Biol. Chem. 265, 14738–14741 (1990).
Klugbauer, N., Marais, E. & Hofmann, F. Calcium channel alpha2delta subunits: differential expression, function, and drug binding. J. Bioenerg. Biomembr. 35, 639–647 (2003).
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94 (1997).
Lesage, F. et al. TWIK-1, a ubiquitous human weakly inward rectifying K+ channel with a novel structure. EMBO J. 15, 1004–1011 (1996).
Fang, M. et al. Dexras1: a G protein specifically coupled to neuronal nitric oxide synthase via CAPON. Neuron 28, 183–193 (2000).
Massion, P.B., Pelat, M., Belge, C. & Balligand, J.L. Regulation of the mammalian heart function by nitric oxide. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 142, 144–150 (2005).
Khan, S.A. et al. Neuronal nitric oxide synthase negatively regulates xanthine oxidoreductase inhibition of cardiac excitation-contraction coupling. Proc. Natl. Acad. Sci. USA 101, 15944–15948 (2004).
Murata, M. et al. SAP97 interacts with Kv1.5 in heterologous expression systems. Am. J. Physiol. Heart Circ. Physiol. 281, H2575–H2584 (2001).
Leonoudakis, D., Mailliard, W., Wingerd, K., Clegg, D. & Vandenberg, C. Inward rectifier potassium channel Kir2.2 is associated with synapse-associated protein SAP97. J. Cell Sci. 114, 987–998 (2001).
Kim, E. & Sheng, M. Differential K+ channel clustering activity of PSD-95 and SAP97, two related membrane-associated putative guanylate kinases. Neuropharmacology 35, 993–1000 (1996).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Wichmann, H.E., Gieger, C. & Illig, T. KORA-gen–resource for population genetics, controls and a broad spectrum of disease phenotypes. Gesundheitswesen 67 (Suppl.), S26–S30 (2005).
Mortara, D.W. Source consistency filtering. Application to resting ECGs. J. Electrocardiol. 25(Suppl.), 200–206 (1992).
Bailey, J.J. et al. Recommendations for standardization and specifications in automated electrocardiography: bandwidth and digital signal processing. A report for health professionals by an ad hoc writing group of the Committee on Electrocardiography and Cardiac Electrophysiology of the Council on Clinical Cardiology, American Heart Association. Circulation 81, 730–739 (1990).
Matsuzaki, H. et al. Genotyping over 100,000 SNPs on a pair of oligonucleotide arrays. Nat. Methods 1, 109–111 (2004).
Weir, B. Genetic Data Analysis II (Sinauer Associates, Cumberland, Massachusetts, 1996).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Acknowledgements
The authors wish to thank G. Tomaselli, J. Nathans, S. Lin and D.J. Cutler for numerous helpful discussions and D. Levy, E. Benjamin and R. D'Agostino at FHS for contributions to the electrocardiographic QT measurement study. This work was supported in part by the D.W. Reynolds Clinical Cardiovascular Research Center, Johns Hopkins University, the US National Institutes of Health, and the German Federal Ministry of Education and Research (BMBF) both in the context of the program Bioinformatics for the Functional Analysis of Mammalian Genomes (BFAM) and the German National Genome Research Network (NGFN). The authors want to thank H. Löwel, C. Meisinger, R. Holle and J. John from the KORA Study Group. The FHS replication study is a contribution from the Framingham Heart Study of the National Heart, Lung, and Blood Institute (NHLBI) of the National Institutes of Health and Boston University School of Medicine, supported by NHLBI's Framingham Heart Study (Contract No. N01-HC-25195) and the Cardiogenomics Program for Genomic Applications (5U01HL066582), a GlaxoSmithKline Competitive Grants Award Program for Young Investigators (CNC) and NIH (K23HL080025, CNC). Some of the electrocardiographic measurements were supported by an unrestricted grant from Pfizer, Inc.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
A.C. is a paid member of the Affymetrix Scientific Advisory Board. The terms of this arrangement are being managed by the Johns Hopkins University in accordance with its conflict of interest policies.
Supplementary information
Supplementary Table 1
Summary of stage I results stratified by short and long QT interval. (PDF 90 kb)
Supplementary Table 2
Candidate gene list and SNP coverage on Affymetrix Centurion arrays. (PDF 42 kb)
Supplementary Table 3
Genome-wide association study of QTc_RAS: screening results from stages I and II. (PDF 140 kb)
Supplementary Table 4
Association study of QTc_RAS emphasizing candidate genes assayed genome-wide: screening results from stages I and II. (PDF 140 kb)
Supplementary Table 5
Significance of additional CAPON SNPs in the entire KORA S4 sample. (PDF 137 kb)
Supplementary Table 6
Sequenom SNP primers, probes and conditions. (PDF 65 kb)
Supplementary Table 7
Primers used in sequencing conserved regions in CAPON. (PDF 10 kb)
Rights and permissions
About this article
Cite this article
Arking, D., Pfeufer, A., Post, W. et al. A common genetic variant in the NOS1 regulator NOS1AP modulates cardiac repolarization. Nat Genet 38, 644–651 (2006). https://doi.org/10.1038/ng1790
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1790
This article is cited by
-
Stem cell models of inherited arrhythmias
Nature Cardiovascular Research (2024)
-
Outlining cardiac ion channel protein interactors and their signature in the human electrocardiogram
Nature Cardiovascular Research (2023)
-
Evaluating Common NOS1AP Variants in Patients with Implantable Cardioverter Defibrillators for Secondary Prevention
Journal of Interventional Cardiac Electrophysiology (2022)
-
Association of T66A polymorphism in CASQ2 with PR interval in a Chinese population
Herz (2021)
-
Insights into the genetics of blood pressure in black South African individuals: the Birth to Twenty cohort
BMC Medical Genomics (2018)