Abstract
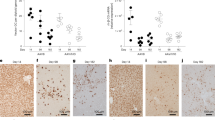
Recombinant adeno-associated virus serotype 2 (rAAV2) is a promising vector for gene therapy because it can achieve long-term stable transgene expression in animals and human subjects after direct administration of vectors into various target tissues1. In the liver, although stable transgene expression primarily results from extrachromosomal vector genomes2, a series of experiments has shown that vector genomes integrate into host chromosomes in hepatocytes3,4,5 at a low frequency2. Despite the low integration efficiency, recent reports of retroviral insertional mutagenesis in mice6 and two human subjects7,8 have raised concerns about the potential for rAAV2-mediated insertional mutagenesis. Here we characterize rAAV2-targeted chromosomal integration sites isolated from selected or non-selected hepatocytes in vector-injected mouse livers. We document frequent chromosomal deletions of up to 2 kb at integration sites (14 of 14 integrations, 100%; most of the deletions were <0.3 kb) and preferred integration into genes (21 of 29 integrations, 72%). In addition, all of the targeted genes analyzed (20 of 20 targeted genes, 100%) were expressed in the liver. This is the first report to our knowledge on host chromosomal effects of rAAV2 integration in animals, and it provides insights into the nature of rAAV2 vector integration into chromosomes in quiescent somatic cells in animals and human subjects.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kay, M.A. et al. Evidence for gene transfer and expression of factor IX in haemophilia B patients treated with an AAV vector. Nat. Genet. 24, 257–261 (2000).
Nakai, H. et al. Extrachromosomal recombinant adeno-associated virus vector genomes are primarily responsible for stable liver transduction in vivo. J. Virol. 75, 6969–6976 (2001).
Miao, C.H. et al. The kinetics of rAAV integration in the liver. Nat. Genet. 19, 13–15 (1998).
Nakai, H., Iwaki, Y., Kay, M.A. & Couto, L.B. Isolation of recombinant adeno-associated virus vector-cellular DNA junctions from mouse liver. J. Virol. 73, 5438–5447 (1999).
Chen, S.J., Tazelaar, J., Moscioni, A.D. & Wilson, J.M. In vivo selection of hepatocytes transduced with adeno-associated viral vectors. Mol. Ther. 1, 414–422 (2000).
Li, Z. et al. Murine leukemia induced by retroviral gene marking. Science 296, 497 (2002).
Marshall, E. Clinical research. Gene therapy a suspect in leukemia-like disease. Science 298, 34–35 (2002).
Marshall, E. Gene therapy. Second child in French trial is found to have leukemia. Science 299, 320 (2003).
Rutledge, E.A. & Russell, D.W. Adeno-associated virus vector integration junctions. J. Virol. 71, 8429–8436 (1997).
Yang, C.C. et al. Cellular recombination pathways and viral terminal repeat hairpin structures are sufficient for adeno-associated virus integration in vivo and in vitro. J. Virol. 71, 9231–9247 (1997).
Miller, D.G., Rutledge, E.A. & Russell, D.W. Chromosomal effects of adeno-associated virus vector integration. Nat. Genet. 30, 147–148 (2002).
Waterston, R.H. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Grompe, M. et al. Loss of fumarylacetoacetate hydrolase is responsible for the neonatal hepatic dysfunction phenotype of lethal albino mice. Genes Dev. 7, 2298–2307 (1993).
Overturf, K. et al. Hepatocytes corrected by gene therapy are selected in vivo in a murine model of hereditary tyrosinaemia type I. Nat. Genet. 12, 266–273 (1996).
Miki, R. et al. Delineating developmental and metabolic pathways in vivo by expression profiling using the RIKEN set of 18,816 full-length enriched mouse cDNA arrays. Proc. Natl. Acad. Sci. USA 98, 2199–2204 (2001).
Su, A.I. et al. Large-scale analysis of the human and mouse transcriptomes. Proc. Natl. Acad. Sci. USA 99, 4465–4470 (2002).
Vijaya, S., Steffen, D.L. & Robinson, H.L. Acceptor sites for retroviral integrations map near DNase I-hypersensitive sites in chromatin. J. Virol. 60, 683–692 (1986).
Sandmeyer, S.B., Hansen, L.J. & Chalker, D.L. Integration specificity of retrotransposons and retroviruses. Annu. Rev. Genet. 24, 491–518 (1990).
Scherdin, U., Rhodes, K. & Breindl, M. Transcriptionally active genome regions are preferred targets for retrovirus integration. J. Virol. 64, 907–912 (1990).
Chalker, D.L. & Sandmeyer, S.B. Ty3 integrates within the region of RNA polymerase III transcription initiation. Genes Dev. 6, 117–128 (1992).
Pryciak, P.M. & Varmus, H.E. Nucleosomes, DNA-binding proteins, and DNA sequence modulate retroviral integration target site selection. Cell 69, 769–780 (1992).
Muller, H.P. & Varmus, H.E. DNA bending creates favored sites for retroviral integration: an explanation for preferred insertion sites in nucleosomes. EMBO J. 13, 4704–4714 (1994).
Pruss, D., Bushman, F.D. & Wolffe, A.P. Human immunodeficiency virus integrase directs integration to sites of severe DNA distortion within the nucleosome core. Proc. Natl. Acad. Sci. USA 91, 5913–5917 (1994).
Leclercq, I. et al. Host sequences flanking the human T-cell leukemia virus type 1 provirus in vivo. J. Virol. 74, 2305–2312 (2000).
Schroder, A.R. et al. HIV-1 integration in the human genome favors active genes and local hotspots. Cell 110, 521–529 (2002).
Nakai, H. et al. Helper-independent and AAV-ITR-independent chromosomal integration of double-stranded linear DNA vectors in mice. Mol. Ther. 7, 101–111 (2003).
Diehn, M. et al. SOURCE: a unified genomic resource of functional annotations, ontologies, and gene expression data. Nucleic Acids Res. 31, 219–223 (2003).
Lander, E.S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Acknowledgements
We thank J. Park for data analysis. This work was supported by grants from the National Heart, Lung, and Blood Institute of the US National Institutes of Health to M.A.K.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
M.A.K. is on the Scientific Advisory Board of Avigen Inc., a company involved in AAV gene therapeutics.
Supplementary information
Rights and permissions
About this article
Cite this article
Nakai, H., Montini, E., Fuess, S. et al. AAV serotype 2 vectors preferentially integrate into active genes in mice. Nat Genet 34, 297–302 (2003). https://doi.org/10.1038/ng1179
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1179
This article is cited by
-
Lack of germline transmission in male mice following a single intravenous administration of AAV5-hFVIII-SQ gene therapy
Gene Therapy (2023)
-
Characterization of integration frequency and insertion sites of adenovirus DNA into mouse liver genomic DNA following intravenous injection
Gene Therapy (2022)
-
Intracellular trafficking of adeno-associated virus (AAV) vectors: challenges and future directions
Gene Therapy (2021)
-
ITR-Seq, a next-generation sequencing assay, identifies genome-wide DNA editing sites in vivo following adeno-associated viral vector-mediated genome editing
BMC Genomics (2020)
-
Safety and efficacy evaluations of an adeno-associated virus variant for preparing IL10-secreting human neural stem cell-based therapeutics
Gene Therapy (2019)