Abstract
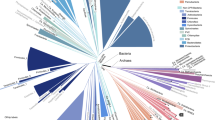
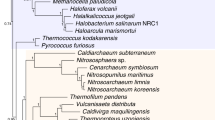
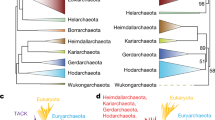
Universal trees of life based on small-subunit (SSU) ribosomal RNA (rRNA) support the separate mono/holophyly of the domains Archaea (archaebacteria), Bacteria (eubacteria) and Eucarya (eukaryotes) and the placement of extreme thermophiles at the base of the Bacteria1,2,3,4. The concept of universal tree reconstruction recently has been upset by protein trees that show intermixing of species from different domains5,6. Such tree topologies have been attributed to either extensive horizontal gene transfer7 or degradation of phylogenetic signals because of saturation for amino acid substitutions8. Here we use large combined alignments of 23 orthologous proteins conserved across 45 species from all domains to construct highly robust universal trees. Although individual protein trees are variable in their support of domain integrity, trees based on combined protein data sets strongly support separate monophyletic domains. Within the Bacteria, we placed spirochaetes as the earliest derived bacterial group. However, elimination from the combined protein alignment of nine protein data sets, which were likely candidates for horizontal gene transfer, resulted in trees showing thermophiles as the earliest evolved bacterial lineage. Thus, combined protein universal trees are highly congruent with SSU rRNA trees in their strong support for the separate monophyly of domains as well as the early evolution of thermophilic Bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Fox, G.E., Magrum, L.J., Balch, W.E., Wolfe, R.S. & Woese, C.R. Classification of methanogenic Bacteria by 16S ribosomal RNA characterization. Proc. Natl. Acad. Sci. USA 74, 4537–4541 (1977).
Woese, C.R. Bacterial evolution. Microbiol. Rev. 51, 221–271 (1987).
Olsen, G.J., Woese, C.R. & Overbeek, R. The winds of (evolutionary) change: breathing new life into microbiology. J. Bacteriol. 176, 1–6 (1994).
Pace, N.R. Origin of life—facing up to the physical setting. Cell 65, 531–533 (1991).
Golding, G.B. & Gupta, R.S. Protein-based phylogenies support a chimeric origin for the eukaryotic genome. Mol. Biol. Evol. 12, 1–6 (1995).
Brown, J.R. & Doolittle, W.F. Archaea and the prokaryote-eukaryote transition. Microbiol. Mol. Biol. Rev. 61, 456–502 (1997).
Doolittle, W.F. Phylogenetic classification and the universal tree. Science 284, 2124–2128 (1999).
Forterre, P. & Philippe, H. Where is the root of the universal tree of life? BioEssays 21, 871–879 (1999).
Xiong, J., Inoue, K. & Bauer, C.E. Tracking molecular evolution of photosynthesis by characterization of a major photosynthesis gene cluster from Heliobacillus mobilis. Proc. Natl. Acad. Sci. USA 95, 14851–14856 (1998).
Baldauf, S.L., Roger, A.J., Wenk-Siefert, I. & Doolittle, W.F. A kingdom-level phylogeny of eukaryotes based on combined protein data. Science 290, 972–977 (2000).
Gray, M.W. The endosymbiont hypothesis revisited. Int. Rev. Cytol. 141, 233–357 (1992).
Ibba, M., Morgan, S., Curnow, A.W., Pridmore, D.R., Vothknecht, U.C., Gardner, W., Lin, W., Woese, C.R. & Söll, D. A euryarchaeal lysyl-tRNA synthetase: resemblance to class I synthetases. Science 278, 1119–1122 (1997).
Teichmann, S.A. & Mitchison, G. Is there a phylogenetic signal in prokaryote proteins? J. Mol. Evol. 49, 98–107 (1999).
Bond, J.P. & Francklyn, C. Proteobacterial histidine-biosynthetic pathways are paraphyletic. J. Mol. Evol. 50, 339–347 (2000).
Brown, J.R., Zhang, J. & Hodgson, J.E. A bacterial antibiotic resistance gene with eukaryotic origins. Curr. Biol. 8, R365–R367 (1998).
Kurland, C.G. & Andersson, S.G.E. Origin and evolution of the mitochondrial proteome. Microbiol. Mol. Biol. Rev. 64, 786–820 (2000).
Chihade, J.W., Brown, J.R., Schimmel, P. & Ribas de Pouplana, L. Origin of mitochondria in relation to evolutionary history of eukaryotic alanyl-tRNA synthetase. Proc. Natl. Acad. Sci. USA 97, 12153–12157 (2000).
Miller, S.L. & Lazcano, A. The origin of life—Did it occur at high temperatures? J. Mol. Evol. 41, 689–692 (1995).
Galtier, N., Tourasse, N. & Gouy, M. A nonhyperthermophilic common ancestor to extant life forms. Science 283, 220–221 (1999).
Rivera, M.C. & Lake, J.A. Evidence that eukaryotes and eocyte prokaryotes are immediate relatives. Science 257, 74–76 (1992).
Hassanin, A. Lecointre, G. & Tillier, S. The evolutionary signal of homoplasy in protein-coding gene sequences and its consequences for a priori weighting in phylogeny. C.R. Acad. Sci. 321, 611–620 (1998).
Springer, M.S., Amrine, H.M., Burk, A. & Stanhope, M.J. Additional support for Afrotheria and Paenungulata, the performance of mitochondrial versus nuclear genes, and the impact of data partitions with heterogeneous base composition. Syst. Biol. 48, 65–75 (1999).
Shah, H.N., Gharbia, S.E. & Collins, M.D. The Gram stain: a declining synapomorphy in an emerging evolutionary tree. Rev. Med. Microbiol. 8, 103–100 (1997).
Nelson, K.E. et al. Evidence for lateral gene transfer between Archaea and Bacteria from genome sequence of Thermotoga maritima. Nature 399, 323–329 (1999).
Logsdon, J.M., Jr. & Faguy, D.M. Evolutionary genomics: Thermotoga heats up lateral gene transfer. Curr. Biol. 9, R747–R751 (1999).
Altschul, S.F., Madden, T.L., Schäffer, A.A., Zhang, J., Zhang, Z., Miller, W. & Lipman, D.J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Thompson, J.D., Higgins, D.G. & Gibson, T. J. CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalities and weight matrix choice., Nucleic Acids Res. 22, 4673–4680 (1994).
Felsenstein, J. PHYLIP (Phylongeny Inference Package), version 3.6. (Department of Genetics, University of Washington, Seattle, 2000.)
Swofford, D.L. PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. (Sinauer Associates, Sunderland, MA, 1999).
Strimmer, K. & von Haeseler, A. Quartet puzzling: a quartet maximum likelihood method for reconstructing tree topologies. Mol. Biol. Evol. 13, 964–969 (1996).
Acknowledgements
We thank the GlaxoSmithKline Bioformatics Division for their support of this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brown, J., Douady, C., Italia, M. et al. Universal trees based on large combined protein sequence data sets. Nat Genet 28, 281–285 (2001). https://doi.org/10.1038/90129
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/90129
This article is cited by
-
Estimating transcriptome complexities across eukaryotes
BMC Genomics (2023)
-
Bioinformatics approaches for classification and investigation of the evolution of the Na/K-ATPase alpha-subunit
BMC Ecology and Evolution (2022)
-
ezTree: an automated pipeline for identifying phylogenetic marker genes and inferring evolutionary relationships among uncultivated prokaryotic draft genomes
BMC Genomics (2018)
-
Comparative analyses of whole-genome protein sequences from multiple organisms
Scientific Reports (2018)
-
A fresh look at the evolution and diversification of photochemical reaction centers
Photosynthesis Research (2015)