Abstract
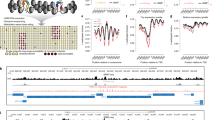
Chromatin is the highly complex structure consisting of DNA and hundreds of associated proteins. Most chromatin proteins exert their regulatory and structural functions by binding to specific chromosomal loci. Knowledge of the identity of these in vivo target loci is essential for the understanding of the functions and mechanisms of action of chromatin proteins. We report here large-scale mapping of in vivo binding sites of chromatin proteins, using a novel approach based on a combination of targeted DNA methylation and microarray technology. We show that three distinct chromatin proteins in Drosophila melanogaster cells each associate with specific sets of genes. HP1 binds predominantly to pericentric genes and transposable elements. GAGA factor associates with euchromatic genes that are enriched in (GA)n motifs. A Drosophila homolog of Saccharomyces cerevisiae Sir2p is associated with several active genes and is excluded from heterochromatin. High-resolution, genome-wide maps of target loci of chromatin proteins ('chromatin profiles') provide new insights into chromatin structure and gene regulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
van Steensel, B. & Henikoff, S. Identification of in vivo DNA targets of chromatin proteins using tethered dam methyltransferase. Nature Biotechnol. 18, 424–428 (2000).
James, T.C. et al. Distribution patterns of HP1, a heterochromatin-associated nonhistone chromosomal protein of Drosophila. Eur. J. Cell Biol. 50, 170–180 (1989).
Pollack, J.R. et al. Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nature Genet. 23, 41–46 (1999).
Charlesworth, B., Jarne, P. & Assimacopoulos, S. The distribution of transposable elements within and between chromosomes in a population of Drosophila melanogaster. III Element abundances in heterochromatin. Genet. Res. 64, 183–197 (1994).
Pimpinelli, S. et al. Transposable elements are stable structural components of Drosophila melanogaster heterochromatin. Proc. Natl. Acad. Sci. USA 92, 3804–3808 (1995).
Carmena, M. & Gonzalez, C. Transposable elements map in a conserved pattern of distribution extending from β-heterochromatin to centromeres in Drosophila melanogaster. Chromosoma 103, 676–684 (1995).
Henikoff, S. Heterochromatin function in complex genomes. Biochim. Biophys. Acta 1470, O1–O8 (1999).
Tsukiyama, T., Becker, P.B. & Wu, C. ATP-dependent nucleosome disruption at a heat-shock promoter mediated by binding of GAGA transcription factor. Nature 367, 525–532 (1994).
Benyajati, C. et al. Multiple isoforms of GAGA factor, a critical component of chromatin structure. Nucleic Acids Res. 25, 3345–3353 (1997).
Biggin, M.D. & Tjian, R. Transcription factors that activate the Ultrabithorax promoter in developmentally staged extracts. Cell 53, 699–711 (1988).
Soeller, W.C., Oh, C.E. & Kornberg, T.B. Isolation of cDNAs encoding the Drosophila GAGA transcription factor. Mol. Cell. Biol. 13, 7961–7970 (1993).
Strutt, H., Cavalli, G. & Paro, R. Co-localization of Polycomb protein and GAGA factor on regulatory elements responsible for the maintenance of homeotic gene expression. EMBO J. 16, 3621–3632 (1997).
O'Brien, T., Wilkins, R.C., Giardina, C. & Lis, J.T. Distribution of GAGA protein on Drosophila genes in vivo. Genes Dev. 9, 1098–1110 (1995).
Guarente, L. Sir2 links chromatin silencing, metabolism, and aging. Genes Dev. 14, 1021–1026 (2000).
Gartenberg, M.R. The Sir proteins of Saccharomyces cerevisiae: mediators of transcriptional silencing and much more. Curr. Opin. Microbiol. 3, 132–137 (2000).
Gotta, M. et al. Localization of Sir2p: the nucleolus as a compartment for silent information regulators. EMBO J. 16, 3243–3255 (1997).
Cuperus, G., Shafaatian, R. & Shore, D. Locus specificity determinants in the multifunctional yeast silencing protein Sir2. EMBO J. 19, 2641–2651 (2000).
Cockell, M.M., Perrod, S. & Gasser, S.M. Analysis of Sir2p domains required for rDNA and telomeric silencing in Saccharomyces cerevisiae. Genetics 154, 1069–1083 (2000).
Frye, R.A. Phylogenetic classification of prokaryotic and eukaryotic Sir2-like proteins. Biochem. Biophys. Res. Commun. 273, 793–798 (2000).
Blat, Y. & Kleckner, N. Cohesins bind to preferential sites along yeast chromosome III, with differential regulation along arms versus the centric region. Cell 98, 249–259 (1999).
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2001).
Iyer, V.R. et al. Genomic binding sites of the yeast cell-cycle transcription factors SBF and MBF. Nature 409, 533–538 (2001).
Wines, D.R., Talbert, P.B., Clark, D.V. & Henikoff, S. Introduction of a DNA methyltransferase into Drosophila to probe chromatin structure in vivo. Chromosoma 104, 332–340 (1996).
Henikoff, S., Ahmad, K., Platero, J.S. & van Steensel, B. Heterochromatic deposition of centromeric histone H3-like proteins. Proc. Natl. Acad. Sci. USA 97, 716–721 (2000).
de Lange, T. et al. Structure and variability of human chromosome ends. Mol. Cell. Biol. 10, 518–827 (1990).
Kopczynski, C.C. et al. A high throughput screen to identify secreted and transmembrane proteins involved in Drosophila embryogenesis. Proc. Natl. Acad. Sci. USA 95, 9973–9978 (1998).
Acknowledgements
We thank E. Giniger for coordinating the assembly of the Northwest Fly Consortium microarray; C. Neal for help with microarray construction and analysis; P. Ng for help with the EST annotation; J. Smothers for the anti-HP1 antibody; J. O'Brien for technical assistance; and members of the Henikoff lab for suggestions.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
van Steensel, B., Delrow, J. & Henikoff, S. Chromatin profiling using targeted DNA adenine methyltransferase. Nat Genet 27, 304–308 (2001). https://doi.org/10.1038/85871
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/85871
This article is cited by
-
Genome-wide maps of nucleolus interactions reveal distinct layers of repressive chromatin domains
Nature Communications (2022)
-
Mapping beads on strings
Nature Methods (2022)
-
Rice protein-binding microarrays: a tool to detect cis-acting elements near promoter regions in rice
Planta (2021)
-
Efficient chromatin profiling of H3K4me3 modification in cotton using CUT&Tag
Plant Methods (2020)
-
Recent advances in the spatial organization of the mammalian genome
Journal of Biosciences (2020)