Abstract
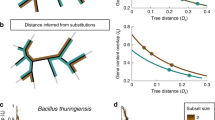
Gene order in bacteria is poorly conserved during evolution1,2,3. For example, although many homologous genes are shared by the proteobacteria Escherichia coli, Haemophilus influenzae and Helicobacter pylori, their relative positions are very different in each genome, except local functional clusters such as operons3,4,5,6. The complete sequences of the more closely related bacterial genomes, such as pairs of Chlamydia7,8,9, H. pylori10,11 and Mycobacterium species12, now allow identification of the processes and mechanisms involved in genome evolution. Here we provide evidence that a substantial proportion of rearrangements in gene order results from recombination sites that are determined by the positions of the replication forks. Our observations suggest that replication has a major role in directing genome evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mushegian, A.R. & Koonin, E.V. Gene order is not conserved in bacterial evolution. Trends Genet. 12, 289–290 (1996).
Huynen, M.A. & Bork, P. Measuring genome evolution. Proc. Natl Acad. Sci. USA 95, 5849–5856 (1998).
Casjens, S. The diverse and dynamic structures of bacterial genomes. Annu. Rev. Genet. 32, 339–377 (1998).
Tatusov, R.L. et al. Metabolism and evolution of Hemophilus influenzae deduced from a whole genome comparison with Escherichia coli. Current Biol. 3, 279–291 (1996).
Kolsko, A.B. Dynamic bacterial genome organization. Mol. Microbiol. 24, 241–248 (1997).
Tamames, J., Casari, G., Ouzounis, C. & Valencia, A. Conserved clusters of functionally related genes in two bacterial genomes. J. Mol. Evol. 44, 66–73 (1997).
Kalman, S. et al. Comparative genomes of Chlamydia pneumoniae and C. trachomatis. Nature Genet. 21, 385–389 (1999).
Stephens, R.S. et al. Genome sequence of an obligate intracellular pathogen of humans, Chlamydia trachomatis. Science 282, 754–759 (1998).
Read, T.D. et al. Genome sequences of Chlamydia trachomatis MoPn and Chlamydia pneumoniae AR39. Nucleic Acids Res. 28, 1397–1406 (2000).
Tomb, J.F. et al. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 388, 539–547 (1997).
Alm, R.A. et al. Genomic-sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature 397, 176–180 (1999).
Cole, S.T. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Parkhill, J. et al. The genome sequence of the food-borne pathogen Campylobacter jejuna reveals hypervariable sequences. Nature 403, 665–668 (2000).
Shapiro, J.A. Molecular model for the transposition and replication of bacteriophage Mu and other transposable elements. Proc. Natl Acad. Sci. USA 76, 1933–1937 (1979).
Liu, S.-H. & Sanderson, K.E. Rearrangements in the genome of the bacterium Salmonella typhi. Proc. Natl Acad. Sci. USA 92, 1018–1022 (1995).
Shmid, M.B. & Roth, J.R. Selection and endpoint distribution of bacterial inversion mutations. Genetics 105, 539–557 (1983).
Rebollo, J.-E., François, V. & Louar, J.-M. Detection and possible role of two large nondivisible zones on the Escherichia coli chromosome. Proc. Natl Acad. Sci. USA 85, 9391–9395 (1988).
Itaya, M. Physical map of the Bacillus subtilis 166 genome: evidence for the inversion of an approximately 1900 kb continuous DNA segment, the translocation of an approximately 100kb segment and the duplication of a 5kb segment. Microbiology 143, 3723–3732 (1997).
Caro, L.G. & Berg, C.M. Chromosome replication in some strains of Escherichia coli K12. Cold Spring Harb. Symp. Quant. Biol. 33, 559–573 (1968).
Schmid, M.B & Roth, J.R. Gene location affects expression level in Salmonella typhimurium. J. Bacteriol. 169, 2872–2875 (1987).
Brewer, B.J. When polymerases collide: replication and the transcriptional organization of the E. coli chromosome. Cell 53, 679–686 (1988).
Ikeda H., Moriya, K. & Matsumoto, T. In vitro study of illegitimate recombination: involvement of DNA gyrase. Cold Spring Harb. Symp. Quant. Biol. 45, 399–408 (1980).
Michel, B., Ehrlich, S.D. & Uzest, M. DNA double-strand breaks caused by replication arrest. EMBO J. 16, 430–438 (1997).
Bierne H., Ehrlich, S.D. & Michel, B. Deletions at stalled replication forks occur by two different pathways. EMBO J. 16, 3332–3340 (1997).
Kuzminov, A. & Stahl, F.W. Double-strand end repair via the RecBC pathway in Escherichia coli primes DNA replication. Genes Dev. 13, 345–356 (1999).
Newport, J. & Yan, H. Organization of DNA into foci during replication. Curr. Opin. Cell Biol. 8, 365–368 (1996).
Lemon, K.P. & Grossman, A.D. Localization of bacterial DNA polymerase: evidence for a factory model of replication. Science 28, 1516–1519 (1998).
Pearson, W.R. Rapid and sensitive sequence comparison with FASTP and FASTA. Methods Enzymol. 183, 63 (1990).
Lobry, J.R. Origin of replication of Mycoplasma genitalium. Science 272, 745–746 (1996).
Tillier, E.R.M. & Collins, R.A. The contributions of replication orientation, gene direction and signal sequences to base-composition asymmetries in bacterial genomes. J. Mol. Evol. 50, 249–257 (2000).
Acknowledgements
We thank W.F. Doolittle for discussion and B. Funnell for critical reading of the manuscript. R.A.C. is a fellow of the Canadian Institute for Advanced Research (CIAR). This work was funded by the National Sciences and Engineering Research Council (NSERC) grant to R.A.C.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tillier, E., Collins, R. Genome rearrangement by replication-directed translocation. Nat Genet 26, 195–197 (2000). https://doi.org/10.1038/79918
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/79918
This article is cited by
-
Comparative in silico genome analysis of Clostridium perfringens unravels stable phylogroups with different genome characteristics and pathogenic potential
Scientific Reports (2021)
-
Organelle inheritance and genome architecture variation in isogamous brown algae
Scientific Reports (2020)
-
Codon usage trend in genes associated with obesity
Biotechnology Letters (2020)
-
Micro-evolution of three Streptococcus species: selection, antigenic variation, and horizontal gene inflow
BMC Evolutionary Biology (2019)
-
Genome-wide detection of genetic loci associated with soybean aphid resistance in soybean germplasm PI 603712
Euphytica (2017)