Abstract
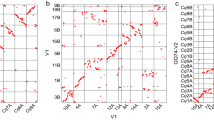
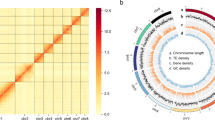
Arabidopsis thaliana has emerged as a model system for studies of plant genetics and development, and its genome has been targeted for sequencing1 by an international consortium (the Arabidopsis Genome Initiative; http://genome-www.stanford.edu/Arabidopsis/agi.html). To support the genome-sequencing effort, we fingerprinted more than 20,000 BACs (ref. 2) from two high-quality publicly available libraries3,4,5, generating an estimated 17-fold redundant coverage of the genome, and used the fingerprints to nucleate assembly of the data by computer. Subsequent manual revision of the assemblies resulted in the incorporation of 19,661 fingerprinted BACs into 169 ordered sets of overlapping clones ('contigs'), each containing at least 3 clones. These contigs are ideal for parallel selection of BACs for large-scale sequencing and have supported the generation of more than 5.8 Mb of finished genome sequence submitted to GenBank; analysis of the sequence has confirmed the integrity of contigs constructed using this fingerprint data. Placement of contigs onto chromosomes can now be performed, and is being pursued by groups involved in both sequencing and positional cloning studies. To our knowledge, these data provide the first example of whole-genome random BAC fingerprint analysis of a eucaryote, and have provided a model essential to efforts aimed at generating similar databases of fingerprint contigs to support sequencing of other complex genomes, including that of human.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bevan, M. et al. Objective: the complete sequence of a plant genome. Plant Cell 9, 476–478 ( 1997).
Shizuya, H. et al. Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor-based vector. Proc. Natl Acad. Sci. USA 89, 8794– 8797 (1992).
Mozo, T., Fischer, S., Shizuya, H. & Altmann, T. Construction and characterization of the IGF Arabidopsis BAC library. Mol. Gen. Genet. 258, 562–570 (1998).
Mozo, T., Fischer, S., Meier-Ewert, S., Lehrach, H. & Altmann, T. Use of the IGF BAC library for physical mapping of the Arabidopsis thaliana genome. Plant J. 16, 377–384 (1998).
Choi, S.D., Creelman, R., Mullet, J. & Wing, R.A. Construction and characterization of a bacterial artificial chromosome library from Arabidopsis thaliana. Weeds World 2, 17– 20 (1995).
Marra, M.A. et al. High throughput fingerprint analysis of large-insert clones. Genome Res. 7, 1072–1084 (1997).
Coulson, A., Sulston, J., Brenner, S. & Karn, J. Toward a physical map of the genome of the nematode Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 83, 7821– 7825 (1986).
Gregory, S.G., Howell, G.R. & Bentley, D.R. Genome mapping by fluorescent fingerprinting. Genome Res. 7, 1162–1168 (1997).
Wilson, R.K. & Mardis, E.R. in Genome Analysis: A Laboratory Manual (eds Birren, B., Green, E.D., Klapholz, S., Myers, R.M. & Roskams, J.) 397–454 (Cold Spring Harbor Laboratory Press, Plainview, 1997).
Olson, M.V. et al. Random-clone strategy for genomic restriction mapping in yeast. Proc. Natl Acad. Sci. USA 83, 7826– 7830 (1986).
Goffeau, A. et al. Life with 6000 genes. Science 274, 563–567 (1996).
Mewes, H.W. et al. Overview of the yeast genome. Nature 387 (suppl.), 7–65 (1997 ).
The C. elegans Genome Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012– 2018 (1998).
Riles, L. et al. Physical maps of the six smallest chromosomes of Saccharomyces cerevisiae at a resolution of 2.6 kilobase pairs. Genetics 134, 81–150 ( 1993).
Sulston, J. et al. Software for genome mapping by fingerprinting techniques. Comput. Appl. Biosci. 4, 125– 132 (1988).
Sulston, J., Mallett, F., Durbin, R. & Horsnell, T. Image analysis of restriction enzyme fingerprint autoradiograms. Comput. Appl. Biosci. 5, 101–106 ( 1989).
Soderlund, C., Longden I. & Mott, R. FPC: a system for building contigs from restriction fingerprinted clones. Comput. Appl. Biosci. 13, 523–535 (1997).
Parsons, J.D. Miropeats: graphical DNA sequence comparisons. Comput. Appl. Biosci. 11, 615–619 ( 1995).
Staden, R. The Staden sequence analysis package. Mol. Biotechnol. 5, 233–241 (1996).
Wong, G.K., Yu, J., Thayer, E.C. & Olson, M.V. Multiple-complete-digest restriction fragment mapping: generating sequence-ready maps for large-scale DNA sequencing. Proc. Natl Acad. Sci. USA 13, 5225–5230 (1997).
Acknowledgements
We thank T. Altmann and R. Wing for providing the scientific community access to their high-quality A. thaliana BAC libraries; E. Mardis, S. Chissoe, W. Barbazuk and S. Gorski for comments and discussion; D. Panussis for design and engineering expertise; M. Holman for maintaining data in web-accessible formats; C. McCabe, N. Florence, D. Scheer, S. Sasso, L. Belaygorod and C. Franklin for expert assistance in fingerprinting; A. Favello for laboratory management; staff at Washington University Genome Sequencing Center for technical support; and D. Preuss, G. Copenhaver, T. Kuromori and many others for providing contig anchoring information. Funding for this work was provided by Monsanto Company.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Marra, M., Kucaba, T., Sekhon, M. et al. A map for sequence analysis of the Arabidopsis thaliana genome . Nat Genet 22, 265–270 (1999). https://doi.org/10.1038/10327
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/10327
This article is cited by
-
Whole Genome Profiling provides a robust framework for physical mapping and sequencing in the highly complex and repetitive wheat genome
BMC Genomics (2012)
-
The peach genome
Tree Genetics & Genomes (2012)
-
Physical mapping and BAC-end sequence analysis provide initial insights into the flax (Linum usitatissimum L.) genome
BMC Genomics (2011)
-
A hybrid BAC physical map of potato: a framework for sequencing a heterozygous genome
BMC Genomics (2011)
-
Physical mapping in highly heterozygous genomes: a physical contig map of the Pinot Noir grapevine cultivar
BMC Genomics (2010)