Abstract
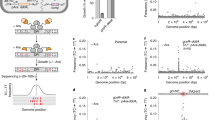
Genome sequencing projects are predicting large numbers of novel proteins, whose interactions with other proteins must mediate the function of cellular processes. To analyse these networks, we used the yeast two-hybrid system on a genome-wide scale to identify 25 interactions among the proteins of Escherichia coli bacteriophage T7. Among these is a set of six interactions connecting proteins that function in DMA replication and DMA packaging. Remarkably, two genes, arranged such that one entirely overlaps the other and uses a different reading frame, encode interacting proteins. Several of the interactions reflect intramolecular associations of different domains of the same polypeptide, suggesting that the two-hybrid assay may be useful in the analysis of protein folding. This global approach to protein–protein interactions may be applicable to the analysis of more complex genomes whose sequences are becoming available.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Fields, S. & Song, O.-K. A novel genetic system to detect protein-protein interactions. Nature 340, 245–246 (1989).
Chien, C.-T., Bartel, P.L., Sternglanz, R. & Fields, S. The two-hybrid system: A method to identify and clone genes for proteins that interact with a protein of interest. Proc. Natl. Acad. Sci. USA 88, 9578–9582 (1991).
Dunn, J.J. & Studier, F.W. Complete nucleotide sequence of bacteriophage T7 DNA and the locations of T7 genetic elements. J. molec. Biol. 166, 477–535 (1983).
Bendixen, C., Gangloff, S. & Rothstein, R. A yeast mating-selection scheme for detection of protein-protein interactions. Nucl. Acids Res. 22, 1778–1779 (1994).
Ma, J. & Ptashne, M. A new class of yeast transcriptional activators. Cell 51, 113–119 (1987).
Breeden, L. & Nasmyth, K. Regulation of the yeast HO gene. Cold Spring Harb. Symp. quant. Biol. 50, 643–650 (1985).
Bartel, P., Chien, C.-T., Sternglanz, R. & Fields, S. Elimination of false positives that arise in using the two-hybrid system. BioTechniques 14, 920–924 (1993).
Kirn, Y.T., Tabor, S., Bortner, C., Griffith, J.D. & Richardson, C.C. J. Biol. Chem. 267, 15022–15031 (1992).
Patel, S.S. & Hingorani, M.M. Oligomeric structure of bacteriophage T7 DNA primase/helicase proteins. J. Biol. Chem. 268, 10668–10675 (1993).
Nakai, H. & Richardson, C.C. Leading and lagging strand synthesis at the replication fork of bacteriophage T7: distinct properties of T7 gene 4 protein as a helicase and primase. J. Biol. Chem. 263, 9818–9830 (1988).
Patel, S.S., Rosenberg, A.H., Studier, F.W. & Johnson, K.A. Large scale purification and biochemical characterization of T7 primase/helicase proteins: evidence for homodimer and heterodimer formation. J. Biol. Chem. 267, 15013–15021 (1992).
Nakai, H. & Richardson, C.C. Interactions of the DNA polymerase and gene 4 protein of bacteriophage T7: protein-protein and protein-DNA interactions involved in RNA-primed DNA synthesis. J. Biol. Chem. 261, 15208–15216 (1986).
Steven, A. & Tins, B. The structure of bacteriophage T7. In Electron Microscopy of Proteins (eds. Harris, J. & Home, R.) 1–36 (Academic Press, London, 1986).
Roeder, G. & RD.Bacteriophage T7 morphogenesis: Phage-related particles in cells infected with wild-type and mutant T7 phage. Virology. 76, 263–285 (1977).
Matsuo-Kato, H., Fujisawa, H. & Minagawa, T. Structure and assembly of bacteriophage T3 tails. Virology. 109, 157–164 (1981).
Mooney, P.O., North, R. & Molineux, I.J. The role of T7 gene 2 protein in DNA replication. Nucl. Acids Res. 8, 3043–3053 (1980).
Studier, F.W. Identification and mapping of five new genes in bacteriophage T7. J. Molec. Biol. 153, 493–502 (1981).
Spoerel, N., Herrlich, P. & Bickle, T.A. A novel bacteriophage defence mechanism: the anti-restriction protein. Nature 278, 30–34 (1979).
Rahmsdorf, H.J. et al. Protein kinase induction in E. coli by bacteriophage T7. Proc. Natl. Acad. Sci. USA 71, 586–589 (1974).
Casjens, S., Eppler, K., Parr, R. & Poteete, A.R. Nucleotide sequence of the bacteriophage P22 gene 79 to 3 region: identification of a new gene required for lysis. Virology 171, 588–698 (1989).
Sousa, R., Chung, Y.J., Rose, J.R. & Wang, B.-C. Crystal structure of bacteriophage T7 RNA polymerase at 3.3 A resolution. Nature 364, 593–599 (1993).
Fields, S. & Sternglanz, R. The two-hybrid system: An assay for protein–protein interactions. Trends Genet. 10, 286–292 (1994).
Kirn, Y.T., Tabor, S., Churchich, J.E. & Richardson, C.C. Interactions of gene 2.5 protein and DNA polymerase of bacteriophage T7. J. Biol. Chem. 267, 15032–15040 (1992).
Moffatt, B.A. & Studier, F.W. T7 lysozyme inhibits transcription by T7 RNA polymerase. Cell 49, 221–227 (1987).
Huang, Z.J., Curtin, K.D. & Rosbash, M. PER protein interactions and temperature compensation of a circadian clock in Drosophila. Science 267, 1169–1172 (1995).
Printen, J.A. & Sprague, G.F., Protein-protein interactions in the yeast pheromone response pathway: Ste5p interacts with all members of the MAP kinase cascade. Genetics 138, 609–619 (1994).
Finley, R.L.J. & Brent, R. Interaction mating reveals binary and ternary connections between Drosophila cell cycle regulators. Proc. Natl. Acad. Sci. USA 91, 12980–12984 (1994).
Gietz, R.D. & Sugino, A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 74, 527–534 (1988).
Bartel, P.L., Chien, C.-T,, Sternglanz, R. & Fields, S. Using the two-hybrid system to detect protein-protein interactions. In Cellular Interactions in Development: A Practical Approach (ed. Hartley, DA) 153–179 (Oxford University Press, Oxford,1993).
Schiestl, R.H., Manivasakam, P., Woods, R.A. & Gietz, R.D. Introducing DNA into yeast by transformation. Methods 5, 79–85 (1993).
Sherman, F., Fink, G.R. & Hicks, J.B. Methods in yeast genetics (Cold Spring Harbor Laboratory, Cold Spring Harbor, 1986).
Hoffman, C.S. & Winston, F. A ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli . Gene 57, 267–272 (1987).
Hall, M.N., Hereford, L. & Herskowitz, I. Targeting of E coli β-galactosidase to the nucleus in yeast. Cell 36, 1057–1065 (1984).
Altschul, S.F., Gisch, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–10 (1990).
Lancy, E.D., Lifsics, M.R., Munson, P. & Maurer, R. Nucleotide sequences of dnaE, the gene for the polymerase subuntt of DNA polymerase III in Salmonella typhimurium, and a variant that facilitates growth in the absence of another polymerase subunrt. J. Bacteriol. 171, 5581–5586 (1989).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Bartel, P., Roecklein, J., SenGupta, D. et al. A protein linkage map of Escherichia coli bacteriophage T7. Nat Genet 12, 72–77 (1996). https://doi.org/10.1038/ng0196-72
Issue Date:
DOI: https://doi.org/10.1038/ng0196-72
This article is cited by
-
Gene Ontology GAN (GOGAN): a novel architecture for protein function prediction
Soft Computing (2022)
-
Programmable design of orthogonal protein heterodimers
Nature (2019)
-
The yeast two-hybrid assay: still finding connections after 25 years
Nature Methods (2014)
-
Phage lysis: Three steps, three choices, one outcome
Journal of Microbiology (2014)