Abstract
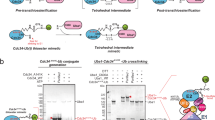
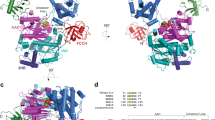
It is widely accepted that ubiquitin-conjugating enzymes contain an active site asparagine that serves as an oxyanion hole, thereby stabilizing a negatively charged transition state intermediate and promoting ubiquitin transfer. Using structural and biochemical approaches to study the role of the conserved asparagine to ubiquitin conjugation by Ubc13–Mms2, we conclude that the importance of this residue stems primarily from its structural role in stabilizing an active site loop.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Pickart, C.M. & Eddins, M.J. Biochim. Biophys. Acta 1695, 55–72 (2004).
Kerscher, O., Felberbaum, R. & Hochstrasser, M. Annu. Rev. Cell Dev. Biol. 22, 159–180 (2006).
Wu, P.Y. et al. EMBO J. 22, 5241–5250 (2003); erratum 26, 4051 (2007).
Yunus, A.A. & Lima, C.D. Nat. Struct. Mol. Biol. 13, 491–499 (2006).
Wenzel, D.M., Stoll, K.E. & Klevit, R.E. Biochem. J. 433, 31–42 (2010).
Sakata, E. et al. Structure 18, 138–147 (2010).
Kamadurai, H.B. et al. Mol. Cell 36, 1095–1102 (2009).
Plechanovová, A., Jaffray, E.G., Tatham, M.H., Naismith, J.H. & Hay, R.T. Nature 489, 115–120 (2012).
Pruneda, J.N. et al. Mol. Cell 47, 933–942 (2012).
Dou, H., Buetow, L., Sibbet, G.J., Cameron, K. & Huang, D.T. Nat. Struct. Mol. Biol. 19, 876–883 (2012).
Bernier-Villamor, V., Sampson, D.A., Matunis, M.J. & Lima, C.D. Cell 108, 345–356 (2002).
Eddins, M.J., Carlile, C.M., Gomez, K.M., Pickart, C.M. & Wolberger, C. Nat. Struct. Mol. Biol. 13, 915–920 (2006).
VanDemark, A.P., Hofmann, R.M., Tsui, C., Pickart, C.M. & Wolberger, C. Cell 105, 711–720 (2001).
Reverter, D. & Lima, C.D. Nature 435, 687–692 (2005).
Hofmann, R.M. & Pickart, C.M. Cell 96, 645–653 (1999).
Carlile, C.M., Pickart, C.M., Matunis, M.J. & Cohen, R.E. J. Biol. Chem. 284, 29326–29334 (2009).
Saha, A., Lewis, S., Kleiger, G., Kuhlman, B. & Deshaies, R.J. Mol. Cell 42, 75–83 (2011).
Wilkinson, K.D. & Rose, I.A. J. Biol. Chem. 256, 9890–9894 (1981).
Wilkinson, K.D. & Rose, I.A. J. Biol. Chem. 254, 12567–12572 (1979).
Ozkan, E., Yu, H. & Deisenhofer, J. Proc. Natl. Acad. Sci. USA 102, 18890–18895 (2005).
Bruice, T.C. & Pandit, U.K. Proc. Natl. Acad. Sci. USA 46, 402–404 (1960).
Jencks, W.P. Catalysis in Chemistry and Enzymology (Dover, New York, 1987).
Liu, H. & Naismith, J.H. BMC Biotechnol. 8, 91 (2008).
Berndsen, C.E. & Wolberger, C. Anal. Biochem. 418, 102–110 (2011).
Pickart, C.M. & Raasi, S. Methods Enzymol. 399, 21–36 (2005).
Gangavarapu, V. et al. Mol. Cell Biol. 26, 7783–7790 (2006).
McCoy, A.J. et al. J. Appl. Crystallogr. 40, 658–674 (2007).
Murshudov, G.N. et al. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Zwart, P. & Afonine, P. Methods Mol. Biol. 426, 419–435 (2008).
Afonine, P.V. et al. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Read, R.J. et al. Structure 19, 1395–1412 (2011).
Chen, V.B. et al. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Davis, I.W., Murray, L.W., Richardson, J.S. & Richardson, D.C. Nucleic Acids Res. 32, W615–W619 (2004).
Acknowledgements
We thank J. Hurley (National Institute of Diabetes and Digestive and Kidney Diseases) for the Ubc13–Mms2 coexpression plasmid and X. Zhang (Johns Hopkins University) for human E1 protein. We also thank J. Stivers, A. Hengge and L. Spyracopoulos for helpful discussions. This work was supported in part by a grant from the US National Science Foundation (MCB-0920082). C.E.B. was supported in part by a Ruth Kirchstein Fellowship from the National Institute of General Medical Science (F32GM089037). General Medical Sciences and Cancer Institutes Structural Biology Facility at the Advanced Photon Source has been funded in whole or in part with Federal funds from the National Cancer Institute (Y1-CO-1020) and the National Institute of General Medical Sciences (Y1-GM-1104). Use of the Advanced Photon Source was supported by the US Department of Energy, Basic Energy Sciences, Office of Science, under contract no. DE-AC02-06CH11357.
Author information
Authors and Affiliations
Contributions
C.E.B., R.W. and C.W. planned experiments; C.E.B., R.W., A.E.R. and I.W.Y. performed biochemical analysis of ubiquitin conjugation; C.E.B. and R.W. crystallized and determined the structure of Ubc13N79A; C.E.B. and C.W. wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results (PDF 982 kb)
Rights and permissions
About this article
Cite this article
Berndsen, C., Wiener, R., Yu, I. et al. A conserved asparagine has a structural role in ubiquitin-conjugating enzymes. Nat Chem Biol 9, 154–156 (2013). https://doi.org/10.1038/nchembio.1159
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1159
This article is cited by
-
UBE2A and UBE2B are recruited by an atypical E3 ligase module in UBR4
Nature Structural & Molecular Biology (2024)
-
Crystal structures of an E1–E2–ubiquitin thioester mimetic reveal molecular mechanisms of transthioesterification
Nature Communications (2021)
-
Linkage-specific ubiquitin chain formation depends on a lysine hydrocarbon ruler
Nature Chemical Biology (2021)
-
Structural basis for RING-Cys-Relay E3 ligase activity and its role in axon integrity
Nature Chemical Biology (2020)
-
A switch element in the autophagy E2 Atg3 mediates allosteric regulation across the lipidation cascade
Nature Communications (2019)