Abstract
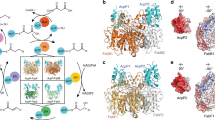
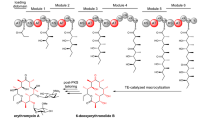
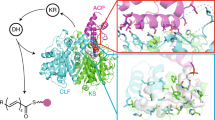
Type I polyketide synthases often use programmed β-branching, via enzymes of a 'hydroxymethylglutaryl-CoA synthase (HCS) cassette', to incorporate various side chains at the second carbon from the terminal carboxylic acid of growing polyketide backbones. We identified a strong sequence motif in acyl carrier proteins (ACPs) where β-branching is known to occur. Substituting ACPs confirmed a correlation of ACP type with β-branching specificity. Although these ACPs often occur in tandem, NMR analysis of tandem β-branching ACPs indicated no ACP-ACP synergistic effects and revealed that the conserved sequence motif forms an internal core rather than an exposed patch. Modeling and mutagenesis identified ACP helix III as a probable anchor point of the ACP–HCS complex whose position is determined by the core. Mutating the core affects ACP functionality, whereas ACP-HCS interface substitutions modulate system specificity. Our method for predicting β-carbon branching expands the potential for engineering new polyketides and lays a basis for determining specificity rules.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Staunton, J. & Weissman, K.J. Polyketide biosynthesis: a millennium review. Nat. Prod. Rep. 18, 380–416 (2001).
Butler, M.S. Natural products to drugs: natural product–derived compounds in clinical trials. Nat. Prod. Rep. 25, 475–516 (2008).
Piel, J. Biosynthesis of polyketides by trans-AT polyketide synthases. Nat. Prod. Rep. 27, 996–1047 (2010).
Calderone, C.T. Isoprenoid-like alkylations in polyketide biosynthesis. Nat. Prod. Rep. 25, 845–853 (2008).
Fuller, A.T. et al. Pseudomonic acid: an antibiotic produced by Pseudomonas fluorescens. Nature 234, 416–417 (1971).
Thomas, C.M., Hothersall, J., Willis, C.L. & Simpson, T.J. Resistance to and synthesis of the antibiotic mupirocin. Nat. Rev. Microbiol. 8, 281–289 (2010).
Fukuda, D. et al. A natural plasmid uniquely encodes two biosynthetic pathways creating a potent anti-MRSA antibiotic. PLoS ONE 6, e18031 (2011).
Rahman, A.S., Hothersall, J., Crosby, J., Simpson, T.J. & Thomas, C.M. Tandemly duplicated acyl carrier proteins, which increase polyketide antibiotic production, can apparently function either in parallel or in series. J. Biol. Chem. 280, 6399–6408 (2005).
Gu, L. et al. Tandem acyl carrier proteins in the curacin biosynthetic pathway promote consecutive multienzyme reactions with a synergistic effect. Angew. Chem. Int. Ed. Engl. 50, 2795–2798 (2011).
Busche, A. et al. Characterization of molecular interactions between ACP and halogenase domains in the curacin A polyketide synthase. ACS Chem. Biol. 7, 378–386 (2012).
El-Sayed, A.K. et al. Characterization of the mupirocin biosynthesis gene cluster from Pseudomonas fluorescens NCIMB 10586. Chem. Biol. 10, 419–430 (2003).
Chang, Z. et al. Biosynthetic pathway and gene cluster analysis of curacin A, an antitubulin natural product from the tropical marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 67, 1356–1367 (2004).
Edwards, D.J. et al. Structure and biosynthesis of the jamaicamides, new mixed polyketide-peptide neurotoxins from the marine cyanobacterium Lyngbya majuscula. Chem. Biol. 11, 817–833 (2004).
Piel, J., Wen, G.P., Platzer, M. & Hui, D.Q. Unprecedented diversity of catalytic domains in the first four modules of the putative pederin polyketide synthase. ChemBioChem 5, 93–98 (2004).
Calderone, C.T., Kowtoniuk, W.E., Kelleher, N.L., Walsh, C.T. & Dorrestein, P.C. Convergence of isoprene and polyketide biosynthetic machinery: isoprenyl-S-carrier proteins in the pksX pathway of Bacillus subtilis. Proc. Natl. Acad. Sci. USA 103, 8977–8982 (2006).
Chen, X.H. et al. Structural and functional characterization of three polyketide synthase gene clusters in Bacillus amyloliquefaciens FZB 42. J. Bacteriol. 188, 4024–4036 (2006).
Mattheus, W. et al. Isolation and purification of a new kalimantacin/batumin-related polyketide antibiotic and elucidation of its biosynthesis gene cluster. Chem. Biol. 17, 149–159 (2010).
Schneider, T.D. & Stephens, R.M. Sequence logos—a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Irschik, H. et al. Analysis of the sorangicin gene cluster reinforces the utility of a combined phylogenetic/retrobiosynthetic analysis for deciphering natural product assembly by trans-AT PKS. ChemBioChem 11, 1840–1849 (2010).
Nakano, M.M., Corbell, N., Besson, J. & Zuber, P. Isolation and characterization of sfp: a gene that functions in the production of the lipopeptide biosurfactant, surfactin, in Bacillus subtilis. Mol. Gen. Genet. 232, 313–321 (1992).
Shields, J.A. et al. Phosphopantetheinylation and specificity of acyl carrier proteins in the mupirocin biosynthetic cluster. ChemBioChem 11, 248–255 (2010).
Evans, S.E. et al. An ACP structural switch: conformational differences between the apo and holo forms of the actinorhodin polyketide synthase acyl carrier protein. ChemBioChem 9, 2424–2432 (2008).
Leibundgut, M., Jenni, S., Frick, C. & Ban, N. Structural basis for substrate delivery by acyl carrier protein in the yeast fatty acid synthase. Science 316, 288–290 (2007).
Alekseyev, V.Y., Liu, C.W., Cane, D.E., Puglisi, J.D. & Khosla, C. Solution structure and proposed domain-domain recognition interface of an acyl carrier protein domain from a modular polyketide synthase. Protein Sci. 16, 2093–2107 (2007).
Wattana-amorn, P. et al. Solution structure of an acyl carrier protein domain from a fungal Type I polyketide synthase. Biochemistry 49, 2186–2193 (2010).
Crump, M.P. et al. Solution structure of the actinorhodin polyketide synthase acyl carrier protein from Streptomyces coelicolor A3(2). Biochemistry 36, 6000–6008 (1997).
Evans, S.E. et al. Probing the interactions of early polyketide intermediates with the actinorhodin ACP from S. coelicolor A3(2). J. Mol. Biol. 389, 511–528 (2009).
El-Sayed, A.K., Hothersall, J. & Thomas, C.M. Quorum-sensing–dependent regulation of biosynthesis of the polyketide antibiotic mupirocin in Pseudomonas fluorescens NCIMB 10586. Microbiology 147, 2127–2139 (2001).
de Vries, S.J., van Dijk, M. & Bonvin, A.M.J.J. The HADDOCK web server for data-driven biomolecular docking. Nat. Protoc. 5, 883–897 (2010).
Mihalek, I., Res, I. & Lichtarge, O. A family of evolution-entropy hybrid methods for ranking protein residues by importance. J. Mol. Biol. 336, 1265–1282 (2004).
Kufareva, I., Budagyan, L., Raush, E., Totrov, M. & Abagyan, R. PIER: protein interface recognition for structural proteomics. Proteins 67, 400–417 (2007).
Valley, C.C. et al. The methionine-aromatic motif plays a unique role in stabilizing protein structure. J. Biol. Chem. 287, 34979–34991 (2012).
Calderone, C.T., Iwig, D.F., Dorrestein, P.C., Kelleher, N.L. & Walsh, C.T. Incorporation of nonmethyl branches by isoprenoid-like logic: multiple β-alkylation events in the biosynthesis of myxovirescin A1. Chem. Biol. 14, 835–846 (2007).
Khosla, C., Ebertkhosla, S. & Hopwood, D.A. Targeted gene replacements in a Streptomyces polyketide synthase gene-cluster—role for the acyl carrier protein. Mol. Microbiol. 6, 3237–3249 (1992).
Decker, H., Summers, R.G. & Hutchinson, C.R. Overproduction of the acyl carrier protein component of a Type II polyketide synthase stimulates production of tetracenomycin biosynthetic intermediates in Streptomyces glaucescens. J. Antibiot. (Tokyo) 47, 54–63 (1994).
Beltran-Alvarez, P., Cox, R.J., Crosby, J. & Simpson, T.J. Dissecting the component reactions catalyzed by the actinorhodin minimal polyketide synthase. Biochemistry 46, 14672–14681 (2007).
Daley, D.J. Queuing output processes. Adv. Appl. Prob. 8, 395–415 (1976).
Shiozawa, H. et al. Thiomarinol, a new hybrid antimicrobial antibiotic produced by a marine bacterium fermentation, isolation, structure and antimicrobial activity. J. Antibiot. (Tokyo) 46, 1834–1842 (1993).
Bertani, G. Studies on lysogenesis. 1. The mode of phage liberation by lysogenic Escherchia coli. J. Bacteriol. 62, 293–300 (1951).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual 2nd edn. (Cold Spring Harbor Press, New York, 1989).
Lepre, C.A. & Moore, J.M. Microdrop screening: a rapid method to optimize solvent conditions for NMR spectroscopy of proteins. J. Biomol. NMR 12, 493–499 (1998).
Vranken, W.F. et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59, 687–696 (2005).
Ottiger, M., Delaglio, F. & Bax, A. Measurement of J and dipolar couplings from simplified two-dimensional NMR spectra. J. Magn. Reson. 131, 373–378 (1998).
Dosset, P., Hus, J.C., Marion, D. & Blackledge, M. A novel interactive tool for rigid-body modeling of multi-domain macromolecules using residual dipolar couplings. J. Biomol. NMR 20, 223–231 (2001).
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999).
Habeck, M., Rieping, W., Linge, J.P. & Nilges, M. NOE assignment with ARIA 2.0: the nuts and bolts. Methods Mol. Biol. 278, 379–402 (2004).
Vriend, G. WHAT IF: a molecular modeling and drug design program. J. Mol. Graph. 8, 52–56 (1990).
Laskowski, R.A., Macarthur, M.W., Moss, D.S. & Thornton, J.M. Procheck—a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Spronk, C.A.E.M., Nabuurs, S.B., Krieger, E., Vriend, G. & Vuister, G.W. Validation of protein structures derived by NMR spectroscopy. Prog. Nucl. Magn. Reson. Spectrosc. 45, 315–337 (2004).
Bachmann, B.O. & Ravel, J. Methods for in silico prediction of microbial polyketide and non-ribosomal peptide biosynthetic pathways from DNA sequence data in. Complex Enzymes in Microbial Natural Product Biosynthesis, Part A: Overview Articles and Peptides Vol. 458 (ed. Hopwood, D.A.) 181–217 (Elsevier Inc., 2009).
Letunic, I., Doerks, T. & Bork, P. SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res. 40, D302–D305 (2012).
Gu, L. et al. Metabolic coupling of dehydration and decarboxylation in the curacin A pathway: Functional identification of a mechanistically diverse enzyme pair. J. Am. Chem. Soc. 128, 9014–9015 (2006).
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein. Sci. 5, 5.6 (2006).
Joosten, R.P. et al. A series of PDB related databases for everyday needs. Nucleic Acids Res. 39, D411–D419 (2011).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure—pattern-recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Galtier, N., Gouy, M. & Gautier, C. SEAVIEW and PHYLO_WIN: two graphic tools for sequence alignment and molecular phylogeny. Comput. Appl. Biosci. 12, 543–548 (1996).
Hornak, V. et al. Comparison of multiple amber force fields and development of improved protein backbone parameters. Proteins 65, 712–725 (2006).
Vriend, G. WHAT IF: a molecular modeling and drug design program. J. Mol. Graph. 8, 52–56 (1990).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958 (2010).
Acknowledgements
This work was supported by the Biotechnology and Biological Sciences Research Council and the Engineering and Physical Sciences Research Council (BB/F014570/1 to P.W-a. and X.D.; BB/I014373/1 to A.S.H., R.G. and J.H.; EP/F066104/1, BB/I003355/1 and BB/I014039/1 to Z.S. and for LC/MS equipment). E.P. was supported by a European Union studentship grant (FP6-mobility 504501). R.F. and Y.T. were supported by scholarships from the Darwin Trust of Edinburgh. J.M. is supported by a PhD scholarship from the FWO Vlaanderen. P.J.W. and R.F. thank A. Bonvin for technical assistance with HADDOCK.
Author information
Authors and Affiliations
Contributions
X.D. purified ACP-mupA3a and ACP-mupA3ab and calculated the NMR structures; P.W.-a. purified ACP mup-A3ab; E.P. purified wild-type ACP mup-A3ab; C.W. calculated NMR structures; C.M.T. and M.P.C. codesigned the study and wrote the paper; J.C. was critical for interpreting data on ACP structure and function; T.J.S. provided expertise in polyketide biosynthesis; A.S.H. discovered the conserved motif, carried out bioinformatic analysis and designed key elements of the experimental tests of our predictions; R.J.C. and C.L.W. designed the LC/MS analysis; Z.S. did the LC/MS analysis; R.F., Y.T. and P.J.W. did bioinformatic analysis and molecular modeling of the ACP mutants and ACP–HCS complex; R.L. and J.M. performed initial BatC complementation assays; and E.Y. and R.G. isolated and characterized the mutant ACP-mupA3a. E.R.S. and J.H. created and characterized the ACP and HCS mutants.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–7 and Supplementary Tables 1–5. (PDF 6776 kb)
Rights and permissions
About this article
Cite this article
Haines, A., Dong, X., Song, Z. et al. A conserved motif flags acyl carrier proteins for β-branching in polyketide synthesis. Nat Chem Biol 9, 685–692 (2013). https://doi.org/10.1038/nchembio.1342
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1342
This article is cited by
-
Insights into azalomycin F assembly-line contribute to evolution-guided polyketide synthase engineering and identification of intermodular recognition
Nature Communications (2023)
-
Initiating polyketide biosynthesis by on-line methyl esterification
Nature Communications (2021)
-
Structure and mechanism of a dehydratase/decarboxylase enzyme couple involved in polyketide β-methyl branch incorporation
Scientific Reports (2020)
-
The structural biology of biosynthetic megaenzymes
Nature Chemical Biology (2015)
-
Exploiting image registration for automated resonance assignment in NMR
Journal of Biomolecular NMR (2015)