Abstract
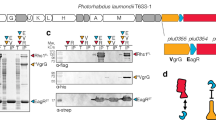
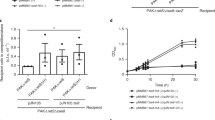
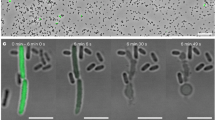
Among bacterial toxin-antitoxin systems, to date no antitoxin has been identified that functions by cleaving toxin mRNA. Here we show that YjdO (renamed GhoT) is a membrane lytic peptide that causes ghost cell formation (lysed cells with damaged membranes) and increases persistence (persister cells are tolerant to antibiotics without undergoing genetic change). GhoT is part of a new toxin-antitoxin system with YjdK (renamed GhoS) because in vitro RNA degradation studies, quantitative real-time reverse-transcription PCR and whole-transcriptome studies revealed that GhoS masks GhoT toxicity by cleaving specifically yjdO (ghoT) mRNA. Alanine substitutions showed that Arg28 is important for GhoS activity, and RNA sequencing indicated that the GhoS cleavage site is rich in U and A. The NMR structure of GhoS indicates it is related to the CRISPR-associated-2 RNase, and GhoS is a monomer. Hence, GhoT-GhoS is to our knowledge the first type V toxin-antitoxin system where a protein antitoxin inhibits the toxin by cleaving specifically its mRNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gerdes, K., Christensen, S.K. & Lobner-Olesen, A. Prokaryotic toxin-antitoxin stress response loci. Nat. Rev. Microbiol. 3, 371–382 (2005).
Hayes, F. & Van Melderen, L. Toxins-antitoxins: diversity, evolution and function. Crit. Rev. Biochem. Mol. Biol. 46, 386–408 (2011).
Masuda, H., Tan, Q., Awano, N., Wu, K.-P. & Inouye, M. YeeU enhances the bundling of cytoskeletal polymers of MreB and FtsZ, antagonizing the CbtA (YeeV) toxicity in Escherichia coli. Mol. Microbiol. 84, 979–989 (2012).
Ren, D., Bedzyk, L.A., Thomas, S.M., Ye, R.W. & Wood, T.K. Gene expression in Escherichia coli biofilms. Appl. Microbiol. Biotechnol. 64, 515–524 (2004).
Kim, Y., Wang, X., Qun, M., Zhang, X.-S. & Wood, T.K. Toxin-antitoxin systems in Escherichia coli influence biofilm formation through YjgK (TabA) and fimbriae. J. Bacteriol. 191, 1258–1267 (2009).
Kim, Y. & Wood, T.K. Toxins Hha and CspD and small RNA regulator Hfq are involved in persister cell formation through MqsR in Escherichia coli. Biochem. Biophys. Res. Commun. 391, 209–213 (2010).
Dörr, T., Vulić, M. & Lewis, K. Ciprofloxacin causes persister formation by inducing the TisB toxin in Escherichia coli. PLoS Biol. 8, e1000317 (2010).
Hu, Y., Benedik, M.J. & Wood, T.K. Antitoxin DinJ influences the general stress response through transcript stabilizer CspE. Environ. Microbiol. 14, 669–679 (2012).
Wang, X. et al. Antitoxin MqsA helps mediate the bacterial general stress response. Nat. Chem. Biol. 7, 359–366 (2011).
Wang, X. & Wood, T.K. Toxin/antitoxin systems influence biofilm and persister cell formation and the general stress response. Appl. Environ. Microbiol. 77, 5577–5583 (2011).
Amitai, S., Kolodkin-Gal, I., Hananya-Meltabashi, M., Sacher, A. & Engelberg-Kulka, H. Escherichia coli MazF leads to the simultaneous selective synthesis of both “death proteins” and “survival proteins”. PLoS Genet. 5, e1000390 (2009).
Belitsky, M. et al. The Escherichia coli extracellular death factor EDF induces the endoribonucleolytic activities of the toxins MazF and ChpBK. Mol. Cell 41, 625–635 (2011).
González Barrios, A.F. et al. Autoinducer 2 controls biofilm formation in Escherichia coli through a novel motility quorum-sensing regulator (MqsR, B3022). J. Bacteriol. 188, 305–316 (2006).
Kim, Y. et al. Escherichia coli toxin/antitoxin pair MqsR/MqsA regulate toxin CspD. Environ. Microbiol. 12, 1105–1121 (2010).
Yamaguchi, Y., Park, J.-H. & Inouye, M. MqsR, a crucial regulator for quorum sensing and biofilm formation, is a GCU-specific mRNA interferase in Escherichia coli. J. Biol. Chem. 284, 28746–28753 (2009).
Christensen-Dalsgaard, M., Jørgensen, M.G. & Gerdes, K. Three new RelE-homologous mRNA interferases of Escherichia coli differentially induced by environmental stresses. Mol. Microbiol. 75, 333–348 (2010).
Domka, J., Lee, J., Bansal, T. & Wood, T.K. Temporal gene-expression in Escherichia coli K-12 biofilms. Environ. Microbiol. 9, 332–346 (2007).
Lewis, K. Persister cells, dormancy and infectious disease. Nat. Rev. Microbiol. 5, 48–56 (2007).
Lewis, K. Persister cells. Annu. Rev. Microbiol. 64, 357–372 (2010).
Brown, B.L. et al. Three dimensional structure of the MqsR:MqsA complex: a novel TA pair comprised of a toxin homologous to RelE and an antitoxin with unique properties. PLoS Pathog. 5, e1000706 (2009).
Keren, I., Shah, D., Spoering, A., Kaldalu, N. & Lewis, K. Specialized persister cells and the mechanism of multidrug tolerance in Escherichia coli. J. Bacteriol. 186, 8172–8180 (2004).
Correia, F.F. et al. Kinase activity of overexpressed HipA is required for growth arrest and multidrug tolerance in Escherichia coli. J. Bacteriol. 188, 8360–8367 (2006).
Shah, D. et al. Persisters: a distinct physiological state of E. coli. BMC Microbiol. 6, 53 (2006).
Balaban, N.Q., Merrin, J., Chait, R., Kowalik, L. & Leibler, S. Bacterial persistence as a phenotypic switch. Science 305, 1622–1625 (2004).
Hofmann, K. & Stoffel, W. TMBASE—a database of membrane spanning protein segments. Biol. Chem. Hoppe-Seyler 374, 166 (1993).
Faridani, O.R., Nikravesh, A., Pandey, D.P., Gerdes, K. & Good, L. Competitive inhibition of natural antisense Sok-RNA interactions activates Hok-mediated cell killing in Escherichia coli. Nucleic Acids Res. 34, 5915–5922 (2006).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Van Melderen, L. & Saavedra De Bast, M. Bacterial toxin–antitoxin systems: more than selfish entities? PLoS Genet. 5, e1000437 (2009).
Güntert, P. Automated NMR structure calculation with CYANA. Methods Mol. Biol. 278, 353–378 (2004).
Herrmann, T., Guntert, P. & Wuthrich, K. Protein NMR structure determination with automated NOE-identification in the NOESY spectra using the new software ATNOS. J. Biomol. NMR 24, 171–189 (2002).
Herrmann, T., Guntert, P. & Wuthrich, K. Protein NMR structure determination with automated NOE assignment using the new software CANDID and the torsion angle dynamics algorithm DYANA. J. Mol. Biol. 319, 209–227 (2002).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Holm, L., Kaariainen, S., Rosenstrom, P. & Schenkel, A. Searching protein structure databases with DaliLite v.3. Bioinformatics 24, 2780–2781 (2008).
Orengo, C.A., Jones, D.T. & Thornton, J.M. Protein superfamilies and domain superfolds. Nature 372, 631–634 (1994).
Beloglazova, N. et al. A novel family of sequence-specific endoribonucleases associated with the clustered regularly interspaced short palindromic repeats. J. Biol. Chem. 283, 20361–20371 (2008).
Samai, P., Smith, P. & Shuman, S. Structure of a CRISPR-associated protein Cas2 from Desulfovibrio vulgaris. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 66, 1552–1556 (2010).
Beloglazova, N. et al. A novel family of sequence-specific endoribonucleases associated with the clustered regularly interspaced short palindromic repeats. J. Biol. Chem. 283, 20361–20371 (2008).
Unoson, C. & Wagner, E.G.H. A small SOS-induced toxin is targeted against the inner membrane in Escherichia coli. Mol. Microbiol. 70, 258–270 (2008).
de la Hoz, A.B. et al. Plasmid copy-number control and better-than-random segregation genes of pSM19035 share a common regulator. Proc. Natl. Acad. Sci. USA 97, 728–733 (2000).
Donegan, N.P. & Cheung, A.L. Regulation of the mazEF toxin-antitoxin module in Staphylococcus aureus and its impact on sigB expression. J. Bacteriol. 191, 2795–2805 (2009).
Jayaraman, R. Bacterial persistence: some new insights into an old phenomenon. J. Biosci. 33, 795–805 (2008).
Maisonneuve, E., Shakespeare, L.J., Jørgensen, M.G. & Gerdes, K. Bacterial persistence by RNA endonucleases. Proc. Natl. Acad. Sci. USA 108, 13206–13211 (2011).
Gerdes, K. The parB (hok/sok) locus of plasmid R1: a general purpose plasmid stabilization system. Nat. Biotechnol. 6, 1402–1405 (1988).
Pecota, D.C. & Wood, T.K. Exclusion of T4 phage by the hok/sok killer locus from plasmid R1. J. Bacteriol. 178, 2044–2050 (1996).
Pedersen, K. & Gerdes, K. Multiple hok genes on the chromosome of Escherichia coli. Mol. Microbiol. 32, 1090–1102 (1999).
Kwon, A.-R et al. Structural and biochemical characterization of HP0315 from Helicobacter pylori as a VapD protein with an endoribonuclease activity. Nucleic Acids Res. 40, 4216–4228 (2012).
Donegan, K., Matyac, C., Seidler, R. & Porteous, A. Evaluation of methods for sampling, recovery, and enumeration of bacteria applied to the phylloplane. Appl. Environ. Microbiol. 57, 51–56 (1991).
Fletcher, M. The effects of culture concentration and age, time, and temperature on bacterial attachment to polystyrene. Can. J. Microbiol. 23, 1–6 (1977).
Barrios, A.F., Zuo, R., Ren, D. & Wood, T.K. Hha, YbaJ, and OmpA regulate Escherichia coli K12 biofilm formation and conjugation plasmids abolish motility. Biotechnol. Bioeng. 93, 188–200 (2006).
Acknowledgements
This work was supported by the US National Institutes of Health (R01 GM089999 to T.K.W.) and the US National Science Foundation (CAREER award MCB 0952550 to R.P.). X.W. is partially supported by the 1000-Youth Elite Program from China. We are grateful for the Keio and ASKA strains provided by the Genome Analysis Project in Japan and for the initial growth studies conducted by X. Yan and T. Benefield. We also thank S. Vitha for assistance with microscope imaging and Y. Hu for assistance with western blotting. T.K.W. is the Biotechnology Endowed Professor at the Pennsylvania State University.
Author information
Authors and Affiliations
Contributions
T.K.W., X.W., R.P., W.P. and M.J.B. designed the experiments. X.W., H.-Y.C., D.M.L., S.H.H., D.O.O., V.S.-T. and C.Q. performed the in vivo and in vitro assays for the functional studies of GhoT and GhoS and for the regulation of the ghoST operon. K.Z. and D.M.L. purified GhoS, and D.M.L. and W.P. completed the NMR structure with help from T.H. with the structure calculations and refinement. T.K.W. and X.W. authored the nonstructural parts of the manuscript, and R.P. and W.P. wrote the structural sections. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 1554 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Lord, D., Cheng, HY. et al. A new type V toxin-antitoxin system where mRNA for toxin GhoT is cleaved by antitoxin GhoS. Nat Chem Biol 8, 855–861 (2012). https://doi.org/10.1038/nchembio.1062
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1062
This article is cited by
-
Role of PumB antitoxin as a transcriptional regulator of the PumAB type-II toxin–antitoxin system and its endoribonuclease activity on the PumA (toxin) transcript
Molecular Genetics and Genomics (2023)
-
Identification and Characterization of HEPN-MNT Type II TA System from Methanothermobacter thermautotrophicus ΔH
Journal of Microbiology (2023)
-
Biology and evolution of bacterial toxin–antitoxin systems
Nature Reviews Microbiology (2022)
-
The expression of type II TA system genes following persister cell formation in Pseudomonas aeruginosa isolates in the exponential and stationary phases
Archives of Microbiology (2022)
-
Toxin–antitoxin systems in pathogenic Vibrio species: a mini review from a structure perspective
3 Biotech (2022)