Abstract
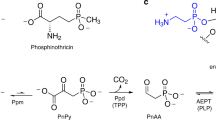
Modifications of the canonical structures of DNA and RNA play critical roles in cell physiology, DNA replication, transcription and translation in all organisms. We now report that bacterial dnd gene clusters incorporate sulfur into the DNA backbone as a sequence-selective, stereospecific phosphorothioate modification. To our knowledge, unlike any other DNA or RNA modification systems, DNA phosphorothioation by dnd gene clusters is the first physiological modification described on the DNA backbone.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
He, X. et al. Mol. Microbiol. 65, 1034–1048 (2007).
Zhou, X., Deng, Z., Firmin, J.L., Hopwood, D.A. & Kieser, T. Nucleic Acids Res. 16, 4341–4352 (1988).
Dyson, P. & Evans, M. Nucleic Acids Res. 26, 1248–1253 (1998).
Zhou, X. et al. Mol. Microbiol. 57, 1428–1438 (2005).
You, D., Wang, L., Yao, F., Zhou, X. & Deng, Z. Biochemistry 46, 6126–6133 (2007).
Lauhon, C.T. & Kambampati, R. J. Biol. Chem. 275, 20096–20103 (2000).
Mueller, E.G., Buck, C.J., Palenchar, P.M., Barnhart, L.E. & Paulson, J.L. Nucleic Acids Res. 26, 2606–2610 (1998).
Boybek, A., Ray, T.D., Evans, M.C. & Dyson, P.J. Nucleic Acids Res. 26, 3364–3371 (1998).
Liang, J. et al. Nucleic Acids Res. 35, 2944–2954 (2007).
Putney, S.D., Benkovic, S.J. & Schimmel, P.R. Proc. Natl. Acad. Sci. USA 78, 7350–7354 (1981).
Vosberg, H.P. & Eckstein, F. Biochemistry 16, 3633–3640 (1977).
Eckstein, F. & Gish, G. Trends Biochem. Sci. 14, 97–100 (1989).
Acknowledgements
We are very grateful to D. Hopwood for his continuous support and encouragement throughout our studies on the DNA S-modification system. We are grateful to L. Cui, B. Chen and J. Son (MIT) and J. Marr (Agilent) for expert assistance with mass spectrometry, to Agilent for access to the XCT Ultra ion-trap mass spectrometer, to T. Murase (Tottori University, Japan) for the generous gift of S. enterica serovar Cerro 87, and to J. Fleckenstein (University of Tennessee Health Sciences Center) for a generous gift of the E. coli strain B7A. Financial support for this work was provided by the Ministry of Science and Technology, the National Science Foundation of China, the Ministry of Education of China, the Shanghai Municipal Council of Science and Technology, the US National Cancer Institute and a Center Grant from the US National Institute of Environmental Health Sciences.
Author information
Authors and Affiliations
Contributions
L.W. and S.C. contributed equally to this work and were responsible for all of the studies related to the chemical and biochemical characterization of the phosphorothioate bonds. T.X. performed the Dnd phenotype studies depicted in Supplementary Figure 1. T.X., X.Z., D.Y. and Z.D. participated in the development of biochemical methods and the bacterial strains used in the studies. K.T. and J.S.W. developed analytical methods and assisted with performing chromatographic and mass spectrometric studies. P.C.D. was responsible for planning and oversight with the chemical and biochemical characterization of the phosphorothioate bonds. L.W., S.C., Z.D. and P.C.D. wrote the paper. All authors discussed the results and assisted with editing of the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1, Supplementary Data and Supplementary Methods (PDF 852 kb)
Rights and permissions
About this article
Cite this article
Wang, L., Chen, S., Xu, T. et al. Phosphorothioation of DNA in bacteria by dnd genes. Nat Chem Biol 3, 709–710 (2007). https://doi.org/10.1038/nchembio.2007.39
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2007.39
This article is cited by
-
Detection of low-frequency mutations in clinical samples by increasing mutation abundance via the excision of wild-type sequences
Nature Biomedical Engineering (2023)
-
Nicking mechanism underlying the DNA phosphorothioate-sensing antiphage defense by SspE
Nature Communications (2022)
-
The functional coupling between restriction and DNA phosphorothioate modification systems underlying the DndFGH restriction complex
Nature Catalysis (2022)
-
Nucleic acid binding by SAMHD1 contributes to the antiretroviral activity and is enhanced by the GpsN modification
Nature Communications (2021)
-
Phosphorothioate-DNA bacterial diet reduces the ROS levels in C. elegans while improving locomotion and longevity
Communications Biology (2021)