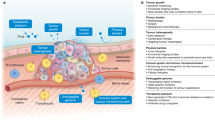
Abstract
Combinatorial control of biological processes, in which redundancy and multifunctionality are the norm, fundamentally limits the therapeutic index that can be achieved by even the most potent and highly selective drugs. Thus, it will almost certainly be necessary to use new 'targeted' pharmaceuticals in combinations. Multicomponent drugs are standard in cytotoxic chemotherapy, but their development has required arduous empirical testing. However, experimentally validated numerical models should greatly aid in the formulation of new combination therapies, particularly those tailored to the needs of specific patients. This perspective focuses on opportunities and challenges inherent in the application of mathematical modeling and systems approaches to pharmacology, specifically with respect to the idea of achieving combinatorial selectivity through use of multicomponent drugs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lipinski, C.A., Lombardo, F., Dominy, B.W. & Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 46, 3–26 (2001).
Wong, S. & Witte, O.N. The BCR-ABL story: bench to bedside and back. Annu. Rev. Immunol. 22, 247–306 (2004).
Saglio, G., Cilloni, D., Rancati, F. & Boano, L. Glivec and CML: a lucky date. J. Biol. Regul. Homeost. Agents 18, 246–251 (2004).
Ikeda, A. et al. Molecular targets and the treatment of myeloid leukemia. Mol. Genet. Metab. 88, 216–224 (2006).
Keith, C.T., Borisy, A.A. & Stockwell, B.R. Multicomponent therapeutics for networked systems. Nat. Rev. Drug Discov. 4, 71–78 (2005).
The Nobel lectures in immunology. The Nobel prize for physiology or medicine, 1908, awarded to Elie Metchnikoff & Paul Ehrlich “in recognition of their work on immunity.” Scand. J. Immunol. 31, 1–13 (1990).
Tortora, G., Bianco, R. & Daniele, G. Strategies for multiple signalling inhibition. J. Chemother. 16 Suppl 4, 41–43 (2004).
Clarke, R. et al. Antiestrogen resistance in breast cancer and the role of estrogen receptor signaling. Oncogene 22, 7316–7339 (2003).
Camirand, A., Zakikhani, M., Young, F. & Pollak, M. Inhibition of insulin-like growth factor-1 receptor signaling enhances growth-inhibitory and proapoptotic effects of gefitinib (Iressa) in human breast cancer cells. Breast Cancer Res. 7, R570–R579 (2005).
Chakravarti, A., Loeffler, J.S. & Dyson, N.J. Insulin-like growth factor receptor I mediates resistance to anti-epidermal growth factor receptor therapy in primary human glioblastoma cells through continued activation of phosphoinositide 3-kinase signaling. Cancer Res. 62, 200–207 (2002).
Scotlandi, K. et al. Prognostic and therapeutic relevance of HER2 expression in osteosarcoma and Ewing's sarcoma. Eur. J. Cancer 41, 1349–1361 (2005).
du Manoir, J.M. et al. Strategies for delaying or treating in vivo acquired resistance to trastuzumab in human breast cancer xenografts. Clin. Cancer Res. 12, 904–916 (2006).
Harrington, L.S. et al. The TSC1–2 tumor suppressor controls insulin-PI3K signaling via regulation of IRS proteins. J. Cell Biol. 166, 213–223 (2004).
Tremblay, F. & Marette, A. Amino acid and insulin signaling via the mTOR/p70 S6 kinase pathway. A negative feedback mechanism leading to insulin resistance in skeletal muscle cells. J. Biol. Chem. 276, 38052–38060 (2001).
Manning, B.D. Balancing Akt with S6K: implications for both metabolic diseases and tumorigenesis. J. Cell Biol. 167, 399–403 (2004).
Loewe, S. The problem of synergism and antagonism of combined drugs. Arzneimittelforschung 3, 285–290 (1953).
Chou, T.C. & Talalay, P. A simple generalized equation for the analysis of multiple inhibitions of Michaelis-Menten kinetic systems. J. Biol. Chem. 252, 6438–6442 (1977).
Chou, T.C. & Talalay, P. Quantitative analysis of dose-effect relationships: the combined effects of multiple drugs or enzyme inhibitors. Adv. Enzyme Regul. 22, 27–55 (1984).
Berenbaum, M.C. What is synergy? Pharmacol. Rev. 41, 93–141 (1989).
Bliss, C.I. The calculation of microbial assays. Bacteriol. Rev. 20, 243–258 (1956).
Greco, W.R., Bravo, G. & Parsons, J.C. The search for synergy: a critical review from a response surface perspective. Pharmacol. Rev. 47, 331–385 (1995).
Martinez-Irujo, J.J., Villahermosa, M.L., Mercapide, J., Cabodevilla, J.F. & Santiago, E. Analysis of the combined effect of two linear inhibitors on a single enzyme. Biochem. J. 329, 689–698 (1998).
Prichard, M.N. & Shipman, C. Jr. A three-dimensional model to analyze drug-drug interactions. Antiviral Res. 14, 181–205 (1990).
Martinez-Irujo, J.J., Villahermosa, M.L., Alberdi, E. & Santiago, E. A checkerboard method to evaluate interactions between drugs. Biochem. Pharmacol. 51, 635–644 (1996).
Tan, M., Fang, H.B., Tian, G.L. & Houghton, P.J. Experimental design and sample size determination for testing synergism in drug combination studies based on uniform measures. Stat. Med. 22, 2091–2100 (2003).
Tallarida, R.J., Stone, D.J. Jr. & Raffa, R.B. Efficient designs for studying synergistic drug combinations. Life Sci. 61, PL 417–425 (1997).
Mead, R. & Pike, D.J. A review of response surface methodology from a biometric viewpoint. Biometrics 31, 803–851 (1975).
Prichard, M.N., Prichard, L.E. & Shipman, C. Jr. Strategic design and three-dimensional analysis of antiviral drug combinations. Antimicrob. Agents Chemother. 37, 540–545 (1993).
Natarajan, M., Lin, K.M., Hsueh, R.C., Sternweis, P.C. & Ranganathan, R. A global analysis of cross-talk in a mammalian cellular signalling network. Nat. Cell Biol. 8, 571–580 (2006).
Yeh, P., Tschumi, A.I. & Kishony, R. Functional classification of drugs by properties of their pairwise interactions. Nat. Genet. 38, 489–494 (2006).
Boulikas, T. & Vougiouka, M. Recent clinical trials using cisplatin, carboplatin and their combination chemotherapy drugs (review). Oncol. Rep. 11, 559–595 (2004).
Peters, G.J. et al. Interaction between cisplatin and gemcitabine in vitro and in vivo. Semin. Oncol. 22, 72–79 (1995).
Chou, T.C., Motzer, R.J., Tong, Y. & Bosl, G.J. Computerized quantitation of synergism and antagonism of taxol, topotecan, and cisplatin against human teratocarcinoma cell growth: a rational approach to clinical protocol design. J. Natl. Cancer Inst. 86, 1517–1524 (1994).
Van Putte, B.P. et al. Combination chemotherapy with gemcitabine with isolated lung perfusion for the treatment of pulmonary metastases. J. Thorac. Cardiovasc. Surg. 130, 125–130 (2005).
Sirotnak, F.M., Zakowski, M.F., Miller, V.A., Scher, H.I. & Kris, M.G. Efficacy of cytotoxic agents against human tumor xenografts is markedly enhanced by coadministration of ZD1839 (Iressa), an inhibitor of EGFR tyrosine kinase. Clin. Cancer Res. 6, 4885–4892 (2000).
Thomas, H.D. et al. Randomized cross-over clinical trial to study potential pharmacokinetic interactions between cisplatin or carboplatin and etoposide. Br. J. Clin. Pharmacol. 53, 83–91 (2002).
Borisy, A.A. et al. Systematic discovery of multicomponent therapeutics. Proc. Natl. Acad. Sci. USA 100, 7977–7982 (2003).
Kholodenko, B.N., Demin, O.V., Moehren, G. & Hoek, J.B. Quantification of short term signaling by the epidermal growth factor receptor. J. Biol. Chem. 274, 30169–30181 (1999).
Schoeberl, B., Eichler-Jonsson, C., Gilles, E.D. & Muller, G. Computational modeling of the dynamics of the MAP kinase cascade activated by surface and internalized EGF receptors. Nat. Biotechnol. 20, 370–375 (2002).
Sasagawa, S., Ozaki, Y., Fujita, K. & Kuroda, S. Prediction and validation of the distinct dynamics of transient and sustained ERK activation. Nat. Cell Biol. 7, 365–373 (2005).
Park, C.S., Schneider, I.C. & Haugh, J.M. Kinetic analysis of platelet-derived growth factor receptor/phosphoinositide 3-kinase/Akt signaling in fibroblasts. J. Biol. Chem. 278, 37064–37072 (2003).
Christopher, R. et al. Data-driven computer simulation of human cancer cell. Ann. NY Acad. Sci. 1020, 132–153 (2004).
Hornberg, J.J. et al. Control of MAPK signalling: from complexity to what really matters. Oncogene 24, 5533–5542 (2005).
Nielsen, U.B. & Schoeberl, B. Using computational modeling to drive the development of targeted therapeutics. IDrugs 8, 822–826 (2005).
Angeli, D., Ferrell, J.E. Jr. & Sontag, E.D. Detection of multistability, bifurcations, and hysteresis in a large class of biological positive-feedback systems. Proc. Natl. Acad. Sci. USA 101, 1822–1827 (2004).
Jackson, R.C. Amphibolic drug combinations: the design of selective antimetabolite protocols based upon the kinetic properties of multienzyme systems. Cancer Res. 53, 3998–4003 (1993).
Lichtner, R.B., Menrad, A., Sommer, A., Klar, U. & Schneider, M.R. Signaling-inactive epidermal growth factor receptor/ligand complexes in intact carcinoma cells by quinazoline tyrosine kinase inhibitors. Cancer Res. 61, 5790–5795 (2001).
Anido, J. et al. ZD1839, a specific epidermal growth factor receptor (EGFR) tyrosine kinase inhibitor, induces the formation of inactive EGFR/HER2 and EGFR/HER3 heterodimers and prevents heregulin signaling in HER2-overexpressing breast cancer cells. Clin. Cancer Res. 9, 1274–1283 (2003).
Goldstein, N.I., Prewett, M., Zuklys, K., Rockwell, P. & Mendelsohn, J. Biological efficacy of a chimeric antibody to the epidermal growth factor receptor in a human tumor xenograft model. Clin. Cancer Res. 1, 1311–1318 (1995).
Matar, P. et al. Combined epidermal growth factor receptor targeting with the tyrosine kinase inhibitor gefitinib (ZD1839) and the monoclonal antibody cetuximab (IMC-C225): superiority over single-agent receptor targeting. Clin. Cancer Res. 10, 6487–6501 (2004).
Ferrell, J.E. Jr. & Machleder, E.M. The biochemical basis of an all-or-none cell fate switch in Xenopus oocytes. Science 280, 895–898 (1998).
Bhalla, U.S., Ram, P.T. & Lyengar, R. MAP kinase phosphatase as a locus of flexibility in a mitogen-activated protein kinase signaling network. Science 297, 1018–1023 (2002).
Kholodenko, B.N. Negative feedback and ultrasensitivity can bring about oscillations in the mitogen-activated protein kinase cascades. Eur. J. Biochem. 267, 1583–1588 (2000).
Asthagiri, A.R. & Lauffenburger, D.A. A computational study of feedback effects on signal dynamics in a mitogen-activated protein kinase (MAPK) pathway model. Biotechnol. Prog. 17, 227–239 (2001).
Sauro, H.M. & Kholodenko, B.N. Quantitative analysis of signaling networks. Prog. Biophys. Mol. Biol. 86, 5–43 (2004).
Manning, B.D. & Cantley, L.C. United at last: the tuberous sclerosis complex gene products connect the phosphoinositide 3-kinase/Akt pathway to mammalian target of rapamycin (mTOR) signalling. Biochem. Soc. Trans. 31, 573–578 (2003).
Bjornsti, M.A. & Houghton, P.J. The TOR pathway: a target for cancer therapy. Nat. Rev. Cancer 4, 335–348 (2004).
O'Reilly, K.E. et al. mTOR inhibition induces upstream receptor tyrosine kinase signaling and activates Akt. Cancer Res. 66, 1500–1508 (2006).
Alves, R., Antunes, F. & Salvador, A. Tools for kinetic modeling of biochemical networks. Nat. Biotechnol. 24, 667–672 (2006).
Kholodenko, B.N. Cell-signalling dynamics in time and space. Nat. Rev. Mol. Cell Biol. 7, 165–176 (2006).
Box, G.E.P. Robustness in the strategy of scientific model building. in Robustness in Statistics (Launer R.L & Wilkinson, G.N.) 202 (Academic Press, New York, 1979).
Loewe, S. Die quantitativen probleme der pharmakologie. Ergeb. Physiol. Biol. Chem. Exp. Pharmakol. 27, 47–187 (1928).
Chou, T.C. & Talalay, P. Generalized equations for the analysis of inhibitions of Michaelis-Menten and higher-order kinetic systems with two or more mutually exclusive and nonexclusive inhibitors. Eur. J. Biochem. 115, 207–216 (1981).
Webb, J.L. in Enzyme and Metabolic Inhibitors 55–79 (Academic, New York, 1963).
Berenbaum, M.C. Criteria for analyzing interactions between biologically active agents. Adv. Cancer Res. 35, 269–335 (1981).
Acknowledgements
We thank B. Harms for helpful suggestions and comments on the manuscript. This work was supported by National Institutes of Health grant GM68762 and by Merrimack Pharmaceuticals.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
P.K.S. is a co-founder of Merrimack Pharmaceuticals and chair of the Scientific Advisory Board.
Supplementary information
Supplementary Table 1
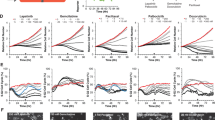
Species, parameters and reactions that can be used to recreate the mechanistic models for Figure 2 (two ligands, two inhibitors). (PDF 235 kb)
Supplementary Table 2
Species, parameters and reactions that can be used to recreate the mechanistic models for Figure 3 (exclusive versus nonexclusive), Figure 4 (linear and ultrasensitive), Figure 5 (negative feedback) and Figure 6 (diseased linear and ultrasensitive). (PDF 239 kb)
Rights and permissions
About this article
Cite this article
Fitzgerald, J., Schoeberl, B., Nielsen, U. et al. Systems biology and combination therapy in the quest for clinical efficacy. Nat Chem Biol 2, 458–466 (2006). https://doi.org/10.1038/nchembio817
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio817
This article is cited by
-
A Deep Neural Network for Predicting Synergistic Drug Combinations on Cancer
Interdisciplinary Sciences: Computational Life Sciences (2024)
-
Dual anti-angiogenic and anti-metastatic activity of myriocin synergistically enhances the anti-tumor activity of cisplatin
Cellular Oncology (2023)
-
Repurposing of approved antivirals against dengue virus serotypes: an in silico and in vitro mechanistic study
Molecular Diversity (2023)
-
Epigenetic derepression converts PPARγ into a druggable target in triple-negative and endocrine-resistant breast cancers
Cell Death Discovery (2021)
-
Sequential combinations of chemotherapeutic agents with BH3 mimetics to treat rhabdomyosarcoma and avoid resistance
Cell Death & Disease (2020)