Abstract
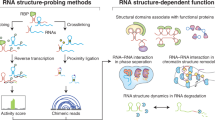
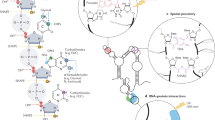
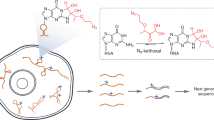
Proper gene expression is essential for the survival of every cell. Once thought to be a passive transporter of genetic information, RNA has recently emerged as a key player in nearly every pathway in the cell. A full description of its structure is critical to understanding RNA function. Decades of research have focused on utilizing chemical tools to interrogate the structures of RNAs, with recent focus shifting to performing experiments inside living cells. This Review will detail the design and utility of chemical reagents used in RNA structure probing. We also outline how these reagents have been used to gain a deeper understanding of RNA structure in vivo. We review the recent merger of chemical probing with deep sequencing. Finally, we outline some of the hurdles that remain in fully characterizing the structure of RNA inside living cells, and how chemical biology can uniquely tackle such challenges.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005).
Badis, G. et al. Diversity and complexity in DNA recognition by transcription factors. Science 324, 1720–1723 (2009).
Kruger, K. et al. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell 31, 147–157 (1982).
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983).
Noller, H.F., Hoffarth, V. & Zimniak, L. Unusual resistance of peptidyl transferase to protein extraction procedures. Science 256, 1416–1419 (1992).
Nissen, P., Hansen, J., Ban, N., Moore, P.B. & Steitz, T.A. The structural basis of ribosome activity in peptide bond synthesis. Science 289, 920–930 (2000).
Serganov, A. & Nudler, E. A decade of riboswitches. Cell 152, 17–24 (2013).
Rinn, J.L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
McHugh, C.A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Lunde, B.M., Moore, C. & Varani, G. RNA-binding proteins: modular design for efficient function. Nat. Rev. Mol. Cell Biol. 8, 479–490 (2007).
Stefl, R., Skrisovska, L. & Allain, F.H. RNA sequence- and shape-dependent recognition by proteins in the ribonucleoprotein particle. EMBO Rep. 6, 33–38 (2005).
Kedde, M. et al. A Pumilio-induced RNA structure switch in p27–3′ UTR controls miR-221 and miR-222 accessibility. Nat. Cell Biol. 12, 1014–1020 (2010).
Chartrand, P., Meng, X.H., Singer, R.H. & Long, R.M. Structural elements required for the localization of ASH1 mRNA and of a green fluorescent protein reporter particle in vivo. Curr. Biol. 9, 333–336 (1999).
Goodarzi, H. et al. Metastasis-suppressor transcript destabilization through TARBP2 binding of mRNA hairpins. Nature 513, 256–260 (2014).
Sobczak, K., de Mezer, M., Michlewski, G., Krol, J. & Krzyzosiak, W.J. RNA structure of trinucleotide repeats associated with human neurological diseases. Nucleic Acids Res. 31, 5469–5482 (2003).
Duan, R., Sharma, S., Xia, Q., Garber, K. & Jin, P. Towards understanding RNA-mediated neurological disorders. J. Genet. Genomics 41, 473–484 (2014).
Houck-Loomis, B. et al. An equilibrium-dependent retroviral mRNA switch regulates translational recoding. Nature 480, 561–564 (2011).
Kang, M., Eichhorn, C.D. & Feigon, J. Structural determinants for ligand capture by a class II preQ1 riboswitch. Proc. Natl. Acad. Sci. USA 111, E663–E671 (2014).
Jiang, J. et al. The architecture of Tetrahymena telomerase holoenzyme. Nature 496, 187–192 (2013).
Weeks, K.M. Advances in RNA structure analysis by chemical probing. Curr. Opin. Struct. Biol. 20, 295–304 (2010).
Ziehler, W.A. & Engelke, D.R. Probing RNA structure with chemical reagents and enzymes. Curr. Protoc. Nucleic Acid Chem. Ch. 6, 6.1.1–6.1.21 (2001).
Low, J.T. & Weeks, K.M. SHAPE-directed RNA secondary structure prediction. Methods 52, 150–158 (2010).
Mathews, D.H. et al. Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure. Proc. Natl. Acad. Sci. USA 101, 7287–7292 (2004).
Lempereur, L. et al. Conformation of yeast 18S rRNA. Direct chemical probing of the 5′ domain in ribosomal subunits and in deproteinized RNA by reverse transcriptase mapping of dimethyl sulfate-accessible. Nucleic Acids Res. 13, 8339–8357 (1985).
Mortimer, S.A., Johnson, J.S. & Weeks, K.M. Quantitative analysis of RNA solvent accessibility by N-silylation of guanosine. Biochemistry 48, 2109–2114 (2009).
Burgstaller, P., Kochoyan, M. & Famulok, M. Structural probing and damage selection of citrulline- and arginine-specific RNA aptamers identify base positions required for binding. Nucleic Acids Res. 23, 4769–4776 (1995).
Wilkinson, K.A., Merino, E.J. & Weeks, K.M. RNA SHAPE chemistry reveals nonhierarchical interactions dominate equilibrium structural transitions in tRNAAsp transcripts. J. Am. Chem. Soc. 127, 4659–4667 (2005).
Merino, E.J., Wilkinson, K.A., Coughlan, J.L. & Weeks, K.M. RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005).
Watts, J.M. et al. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460, 711–716 (2009).
McGinnis, J.L., Duncan, C.D. & Weeks, K.M. High-throughput SHAPE and hydroxyl radical analysis of RNA structure and ribonucleoprotein assembly. Methods Enzymol. 468, 67–89 (2009).
Wan, Y., Qu, K., Ouyang, Z. & Chang, H.Y. Genome-wide mapping of RNA structure using nuclease digestion and high-throughput sequencing. Nat. Protoc. 8, 849–869 (2013).
Peng, Y., Soper, T.J . Ouyang, Z. & Woodson, S.A. RNase footprinting of protein binding sites on an mRNA target of small RNAs. Methods Mol. Biol. 905, 213–224 (2012).
Ingle, S., Azad, R.N., Jain, S.S. & Tullius, T.D. Chemical probing of RNA with the hydroxyl radical at single-atom resolution. Nucleic Acids Res. 42, 12758–12767 (2014).
Costa, M. & Monachello, D. Probing RNA folding by hydroxyl radical footprinting. Methods Mol. Biol. 1086, 119–142 (2014).
Ding, F., Lavender, C.A., Weeks, K.M. & Dokholyan, N.V. Three-dimensional RNA structure refinement by hydroxyl radical probing. Nat. Methods 9, 603–608 (2012).
Regulski, E.E. & Breaker, R.R. In-line probing analysis of riboswitches. Methods Mol. Biol. 419, 53–67 (2008).
Heilman-Miller, S.L. & Woodson, S.A. Effect of transcription on folding of the Tetrahymena ribozyme. RNA 9, 722–733 (2003).
Perdrizet, G.A. II, Artsimovitch, I., Furman, R., Sosnick, T.R. & Pan, T. Transcriptional pausing coordinates folding of the aptamer domain and the expression platform of a riboswitch. Proc. Natl. Acad. Sci. USA 109, 3323–3328 (2012).
Mahen, E.M., Watson, P.Y., Cottrell, J.W. & Fedor, M.J. mRNA secondary structures fold sequentially but exchange rapidly in vivo. PLoS Biol. 8, e1000307 (2010).
Pan, T., Artsimovitch, I., Fang, X.W., Landick, R. & Sosnick, T.R. Folding of a large ribozyme during transcription and the effect of the elongation factor NusA. Proc. Natl. Acad. Sci. USA 96, 9545–9550 (1999).
Tyrrell, J., McGinnis, J.L., Weeks, K.M. & Pielak, G.J. The cellular environment stabilizes adenine riboswitch RNA structure. Biochemistry 52, 8777–8785 (2013).
Frieda, K.L. & Block, S.M. Direct observation of cotranscriptional folding in an adenine riboswitch. Science 338, 397–400 (2012).
Tijerina, P., Mohr, S. & Russell, R. DMS footprinting of structured RNAs and RNA-protein complexes. Nat. Protoc. 2, 2608–2623 (2007).
Waldsich, C., Masquida, B., Westhof, E. & Schroeder, R. Monitoring intermediate folding states of the td group I intron in vivo. EMBO J. 21, 5281–5291 (2002).
Waldsich, C., Grossberger, R. & Schroeder, R. RNA chaperone StpA loosens interactions of the tertiary structure in the td group I intron in vivo. Genes Dev. 16, 2300–2312 (2002).
Swiatkowska, A. et al. Kinetic analysis of pre-ribosome structure in vivo. RNA 18, 2187–2200 (2012).
McGinnis, J.L. & Weeks, K.M. Ribosome RNA assembly intermediates visualized in living cells. Biochemistry 53, 3237–3247 (2014).
McGinnis, J.L. et al. In-cell SHAPE reveals that free 30S ribosome subunits are in the inactive state. Proc. Natl. Acad. Sci. USA 112, 2425–2430 (2015).
Spitale, R.C. et al. RNA SHAPE analysis in living cells. Nat. Chem. Biol. 9, 18–20 (2013).
Kwok, C.K., Ding, Y., Tang, Y., Assmann, S.M. & Bevilacqua, P.C. Determination of in vivo RNA structure in low-abundance transcripts. Nat. Commun. 4, 2971 (2013).
Hector, R.D. et al. Snapshots of pre-rRNA structural flexibility reveal eukaryotic 40S assembly dynamics at nucleotide resolution. Nucleic Acids Res. 42, 12138–12154 (2014).
Latham, J.A. & Cech, T.R. Defining the inside and outside of a catalytic RNA molecule. Science 245, 276–282 (1989).
Sclavi, B., Woodson, S., Sullivan, M., Chance, M.R. & Brenowitz, M. Time-resolved synchrotron X-ray “footprinting”, a new approach to the study of nucleic acid structure and function: application to protein-DNA interactions and RNA folding. J. Mol. Biol. 266, 144–159 (1997).
Sclavi, B., Sullivan, M., Chance, M.R., Brenowitz, M. & Woodson, S.A. RNA folding at millisecond intervals by synchrotron hydroxyl radical footprinting. Science 279, 1940–1943 (1998).
Adilakshmi, T., Soper, S.F. & Woodson, S.A. Structural analysis of RNA in living cells by in vivo synchrotron X-ray footprinting. Methods Enzymol. 468, 239–258 (2009).
Kim, H. et al. Protein-guided RNA dynamics during early ribosome assembly. Nature 506, 334–338 (2014).
Clatterbuck Soper, S.F., Dator, R.P., Limbach, P.A. & Woodson, S.A. In vivo X-ray footprinting of pre-30S ribosomes reveals chaperone-dependent remodeling of late assembly intermediates. Mol. Cell 52, 506–516 (2013).
Blower, M.D. Molecular insights into intracellular RNA localization. Int. Rev. Cell. Mol. Biol. 302, 1–39 (2013).
Hogan, D.J., Riordan, D.P., Gerber, A.P., Herschlag, D. & Brown, P.O. Diverse RNA-binding proteins interact with functionally related sets of RNAs, suggesting an extensive regulatory system. PLoS Biol. 6, e255 (2008).
Mortimer, S.A., Kidwell, M.A. & Doudna, J.A. Insights into RNA structure and function from genome-wide studies. Nat. Rev. Genet. 15, 469–479 (2014).
Kwok, C.K., Tang, Y., Assmann, S.M. & Bevilacqua, P.C. The RNA structurome: transcriptome-wide structure probing with next-generation sequencing. Trends Biochem. Sci. 40, 221–232 (2015).
Lucks, J.B. et al. Multiplexed RNA structure characterization with selective 2′-hydroxyl acylation analyzed by primer extension sequencing (SHAPE-Seq). Proc. Natl. Acad. Sci. USA 108, 11063–11068 (2011).
Ding, Y. et al. In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 505, 696–700 (2014).
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J.S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705 (2014).
Spitale, R.C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–490 (2015).
Schwartz, S. et al. High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell 155, 1409–1421 (2013).
Liu, N. et al. N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518, 560–564 (2015).
Zarrinkar, P.P., Wang, J. & Williamson, J.R. Slow folding kinetics of RNase P RNA. RNA 2, 564–573 (1996).
Zemora, G. & Waldsich, C. RNA folding in living cells. RNA Biol. 7, 634–641 (2010).
Wong, T.N. & Pan, T. RNA folding during transcription: protocols and studies. Methods Enzymol. 468, 167–193 (2009).
Pan, T. & Sosnick, T. RNA folding during transcription. Annu. Rev. Biophys. Biomol. Struct. 35, 161–175 (2006).
Mahen, E.M., Harger, J.W., Calderon, E.M. & Fedor, M.J. Kinetics and thermodynamics make different contributions to RNA folding in vitro and in yeast. Mol. Cell 19, 27–37 (2005).
Danko, C.G. et al. Identification of active transcriptional regulatory elements from GRO-seq data. Nat. Methods 12, 433–438 (2015).
Beach, D.L. & Bloom, K. ASH1 mRNA localization in three acts. Mol. Biol. Cell 12, 2567–2577 (2001).
Lécuyer, E. et al. Global analysis of mRNA localization reveals a prominent role in organizing cellular architecture and function. Cell 131, 174–187 (2007).
Riley, K.J., Yario, T.A. & Steitz, J.A. Association of Argonaute proteins and microRNAs can occur after cell lysis. RNA 18, 1581–1585 (2012).
Mili, S. & Steitz, J.A. Evidence for reassociation of RNA-binding proteins after cell lysis: implications for the interpretation of immunoprecipitation analyses. RNA 10, 1692–1694 (2004).
Mayerle, M. & Woodson, S.A. Specific contacts between protein S4 and ribosomal RNA are required at multiple stages of ribosome assembly. RNA 19, 574–585 (2013).
Panja, S., Schu, D.J. & Woodson, S.A. Conserved arginines on the rim of Hfq catalyze base pair formation and exchange. Nucleic Acids Res. 41, 7536–7546 (2013).
Lucas, C.H., Calvez, M., Babu, R. & Brown, A. Altered subcellular localization of the NeuN/Rbfox3 RNA splicing factor in HIV-associated neurocognitive disorders (HAND). Neurosci. Lett. 558, 97–102 (2014).
Kudla, G., Granneman, S., Hahn, D., Beggs, J.D. & Tollervey, D. Cross-linking, ligation, and sequencing of hybrids reveals RNA-RNA interactions in yeast. Proc. Natl. Acad. Sci. USA 108, 10010–10015 (2011).
Belton, J.M. et al. Hi-C: a comprehensive technique to capture the conformation of genomes. Methods 58, 268–276 (2012).
Yaffe, E. & Tanay, A. Probabilistic modeling of Hi-C contact maps eliminates systematic biases to characterize global chromosomal architecture. Nat. Genet. 43, 1059–1065 (2011).
Sugimoto, Y. et al. hiCLIP reveals the in vivo atlas of mRNA secondary structures recognized by Staufen 1. Nature 519, 491–494 (2015).
Mathews, D.H. RNA secondary structure analysis using RNAstructure. Curr. Protoc. Bioinformatics 46, 12.6.1–12.6.25 (2014).
Markham, N.R. & Zuker, M. UNAFold: software for nucleic acid folding and hybridization. Methods Mol. Biol. 453, 3–31 (2008).
Gruber, A.R., Lorenz, R., Bernhart, S.H., Neubock, R. & Hofacker, I.L. The Vienna RNA websuite. Nucleic Acids Res. 36, W70–W74 (2008).
Deigan, K.E., Li, T.W., Mathews, D.H. & Weeks, K.M. Accurate SHAPE-directed RNA structure determination. Proc. Natl. Acad. Sci. USA 106, 97–102 (2009).
Washietl, S. et al. Computational analysis of noncoding RNAs. Wiley Interdiscip Rev. RNA 3, 759–778 (2012).
Andronescu, M., Condon, A., Hoos, H.H., Mathews, D.H. & Murphy, K.P. Computational approaches for RNA energy parameter estimation. RNA 16, 2304–2318 (2010).
Ouyang, Z., Snyder, M.P. & Chang, H.Y. SeqFold: genome-scale reconstruction of RNA secondary structure integrating high-throughput sequencing data. Genome Res. 23, 377–387 (2013).
Tang, Y. et al. StructureFold: genome-wide RNA secondary structure mapping and reconstruction in vivo. Bioinformatics 31, 2668–2675 (2015).
Corley, M., Solem, A., Qu, K., Chang, H.Y. & Laederach, A. Detecting riboSNitches with RNA folding algorithms: a genome-wide benchmark. Nucleic Acids Res. 43, 1859–1868 (2015).
Acknowledgements
We thank members of the Spitale laboratory and R. Flynn for helpful discussions. We also thank professors S. Woodson and A. Laederach for critical reading of the manuscript. The University of California, Irvine and the US National Institutes of Health Director's New Innovator Award (grant 1DP2GM119164-01) support RNA research in the Spitale lab.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kubota, M., Tran, C. & Spitale, R. Progress and challenges for chemical probing of RNA structure inside living cells. Nat Chem Biol 11, 933–941 (2015). https://doi.org/10.1038/nchembio.1958
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1958
This article is cited by
-
Probing the dynamic RNA structurome and its functions
Nature Reviews Genetics (2023)
-
Global mapping of RNA G-quadruplexes (G4-RNAs) using G4RP-seq
Nature Protocols (2022)
-
Keth-seq for transcriptome-wide RNA structure mapping
Nature Chemical Biology (2020)
-
Prediction and differential analysis of RNA secondary structure
Quantitative Biology (2020)
-
Functional 5′ UTR mRNA structures in eukaryotic translation regulation and how to find them
Nature Reviews Molecular Cell Biology (2018)