Abstract
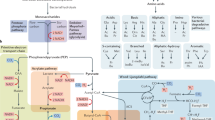
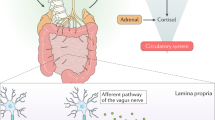
Gut microbiota is found in virtually any metazoan, from invertebrates to vertebrates. It has long been believed that gut microbiota, more specifically, the activity of the microbiome and its metabolic products, directly influence a variety of aspects in metazoan physiology. However, the exact molecular relationship among microbe-derived gut metabolites, host signaling pathways, and host physiology remains to be elucidated. Here we review recent discoveries regarding the molecular links between gut metabolites and host physiology in different invertebrate and vertebrate animal models. We describe the different roles of gut microbiome activity and their metabolites in regulating distinct host physiology and the molecular mechanisms by which gut metabolites cause physiological homeostasis via regulating specific host signaling pathways. Future studies in this direction using different animal models will provide the key concepts to understanding the evolutionarily conserved chemical dialogues between gut microbiota and metazoan cells and also human diseases associated with gut microbiota and metabolites.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lee, W.J. & Brey, P.T. How microbiomes influence metazoan development: insights from history and Drosophila modeling of gut-microbe interactions. Annu. Rev. Cell Dev. Biol. 29, 571–592 (2013).
Gusarov, I. et al. Bacterial nitric oxide extends the lifespan of C. elegans. Cell 152, 818–830 (2013).
Cabreiro, F. et al. Metformin retards aging in C. elegans by altering microbial folate and methionine metabolism. Cell 153, 228–239 (2013).
Buchon, N., Broderick, N.A. & Lemaitre, B. Gut homeostasis in a microbial world: insights from Drosophila melanogaster. Nat. Rev. Microbiol. 11, 615–626 (2013).
Lee, K.A. & Lee, W.J. Drosophila as a model for intestinal dysbiosis and chronic inflammatory diseases. Dev. Comp. Immunol. 42, 102–110 (2014).
Guo, L., Karpac, J., Tran, S.L. & Jasper, H. PGRP-SC2 promotes gut immune homeostasis to limit commensal dysbiosis and extend lifespan. Cell 156, 109–122 (2014).
Shin, S.C. et al. Drosophila microbiome modulates host developmental and metabolic homeostasis via insulin signaling. Science 334, 670–674 (2011).
Gems, D. & Partridge, L. Genetics of longevity in model organisms: debates and paradigm shifts. Annu. Rev. Physiol. 75, 621–644 (2013).
Abada, E.A. et al. C. elegans behavior of preference choice on bacterial food. Mol. Cells 28, 209–213 (2009).
Lenaerts, I., Walker, G.A., Van Hoorebeke, L., Gems, D. & Vanfleteren, J.R. Dietary restriction of Caenorhabditis elegans by axenic culture reflects nutritional requirement for constituents provided by metabolically active microbes. J. Gerontol. A Biol. Sci. Med. Sci. 63, 242–252 (2008).
Saiki, R. et al. Altered bacterial metabolism, not coenzyme Q content, is responsible for the lifespan extension in Caenorhabditis elegans fed an Escherichia coli diet lacking coenzyme Q. Aging Cell 7, 291–304 (2008).
Tang, W.H. et al. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N. Engl. J. Med. 368, 1575–1584 (2013).
Koeth, R.A. et al. Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat. Med. 19, 576–585 (2013).
Yoshimoto, S. et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 499, 97–101 (2013).
Johnson, C.H., Patterson, A.D., Idle, J.R. & Gonzalez, F.J. Xenobiotic metabolomics: major impact on the metabolome. Annu. Rev. Pharmacol. Toxicol. 52, 37–56 (2012).
Nicholson, J.K. & Wilson, I.D. Opinion: understanding 'global' systems biology: metabonomics and the continuum of metabolism. Nat. Rev. Drug Discov. 2, 668–676 (2003).
Mahmood, K., Naeem, M. & Rahimnajjad, N.A. Metformin: the hidden chronicles of a magic drug. Eur. J. Intern. Med. 24, 20–26 (2013).
Pyra, K.A., Saha, D.C. & Reimer, R.A. Prebiotic fiber increases hepatic acetyl CoA carboxylase phosphorylation and suppresses glucose-dependent insulinotropic polypeptide secretion more effectively when used with metformin in obese rats. J. Nutr. 142, 213–220 (2012).
Shin, N.R. et al. An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 63, 727–735 (2014).
Anisimov, V.N. et al. Metformin slows down aging and extends life span of female SHR mice. Cell Cycle 7, 2769–2773 (2008).
Onken, B. & Driscoll, M. Metformin induces a dietary restriction–like state and the oxidative stress response to extend C. elegans healthspan via AMPK, LKB1, and SKN-1. PLoS ONE 5, e8758 (2010).
Grandison, R.C., Piper, M.D. & Partridge, L. Amino-acid imbalance explains extension of lifespan by dietary restriction in Drosophila. Nature 462, 1061–1064 (2009).
Zimmerman, J.A., Malloy, V., Krajcik, R. & Orentreich, N. Nutritional control of aging. Exp. Gerontol. 38, 47–52 (2003).
Slack, C., Foley, A. & Partridge, L. Activation of AMPK by the putative dietary restriction mimetic metformin is insufficient to extend lifespan in Drosophila. PLoS ONE 7, e47699 (2012).
Ryu, J.H. et al. Innate immune homeostasis by the homeobox gene caudal and commensal-gut mutualism in Drosophila. Science 319, 777–782 (2008).
O'Hara, A.M. & Shanahan, F. The gut flora as a forgotten organ. EMBO Rep. 7, 688–693 (2006).
Packey, C.D. & Sartor, R.B. Commensal bacteria, traditional and opportunistic pathogens, dysbiosis and bacterial killing in inflammatory bowel diseases. Curr. Opin. Infect. Dis. 22, 292–301 (2009).
Kamada, N., Chen, G.Y., Inohara, N. & Nuñez, G. Control of pathogens and pathobionts by the gut microbiota. Nat. Immunol. 14, 685–690 (2013).
Lemaitre, B. & Hoffmann, J. The host defense of Drosophila melanogaster. Annu. Rev. Immunol. 25, 697–743 (2007).
Kim, S.H. & Lee, W.J. Role of DUOX in gut inflammation: lessons from Drosophila model of gut-microbiota interactions. Front. Cell. Infect. Microbiol. 3, 116 (2014).
Lee, K.-A. et al. Bacterial-derived uracil as a modulator of mucosal immunity and gut-microbe homeostasis in Drosophila. Cell 153, 797–811 (2013).
Bischoff, V. et al. Downregulation of the Drosophila immune response by peptidoglycan-recognition proteins SC1 and SC2. PLoS Pathog. 2, e14 (2006).
Paredes, J.C., Welchman, D.P., Poidevin, M. & Lemaitre, B. Negative regulation by amidase PGRPs shapes the Drosophila antibacterial response and protects the fly from innocuous infection. Immunity 35, 770–779 (2011).
Lhocine, N. et al. PIMS modulates immune tolerance by negatively regulating Drosophila innate immune signaling. Cell Host Microbe 4, 147–158 (2008).
Franzenburg, S. et al. Distinct antimicrobial peptide expression determines host species-specific bacterial associations. Proc. Natl. Acad. Sci. USA 110, E3730–E3738 (2013).
Cabreiro, F. & Gems, D. Worms need microbes too: microbiota, health and aging in Caenorhabditis elegans. EMBO Mol. Med. 5, 1300–1310 (2013).
Bosch, T.C. Cnidarian-microbe interactions and the origin of innate immunity in metazoans. Annu. Rev. Microbiol. 67, 499–518 (2013).
McFall-Ngai, M. et al. Animals in a bacterial world, a new imperative for the life sciences. Proc. Natl. Acad. Sci. USA 110, 3229–3236 (2013).
Chu, H. & Mazmanian, S.K. Innate immune recognition of the microbiota promotes host-microbial symbiosis. Nat. Immunol. 14, 668–675 (2013).
Storelli, G. et al. Lactobacillus plantarum promotes Drosophila systemic growth by modulating hormonal signals through TOR-dependent nutrient sensing. Cell Metab. 14, 403–414 (2011).
Ley, R.E., Peterson, D.A. & Gordon, J.I. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124, 837–848 (2006).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Nicholson, J.K. et al. Host-gut microbiota metabolic interactions. Science 336, 1262–1267 (2012).
Bäckhed, F. et al. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl. Acad. Sci. USA 101, 15718–15723 (2004).
Cummings, J.H. Fermentation in the human large intestine: evidence and implications for health. Lancet 1, 1206–1209 (1983).
Cummings, J.H., Pomare, E.W., Branch, W.J., Naylor, C.P. & Macfarlane, G.T. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 28, 1221–1227 (1987).
Donohoe, D.R. et al. The microbiome and butyrate regulate energy metabolism and autophagy in the mammalian colon. Cell Metab. 13, 517–526 (2011).
Kuma, A. et al. The role of autophagy during the early neonatal starvation period. Nature 432, 1032–1036 (2004).
Asanuma, N., Kawato, M., Ohkawara, S. & Hino, T. Characterization and transcription of the genes encoding enzymes involved in butyrate production in Butyrivibrio fibrisolvens. Curr. Microbiol. 47, 203–207 (2003).
Cho, I. et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 488, 621–626 (2012).
Ley, R.E. et al. Obesity alters gut microbial ecology. Proc. Natl. Acad. Sci. USA 102, 11070–11075 (2005).
Ley, R.E., Turnbaugh, P.J., Klein, S. & Gordon, J.I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Turnbaugh, P.J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Thangaraju, M. et al. GPR109A is a G protein–coupled receptor for the bacterial fermentation product butyrate and functions as a tumor suppressor in colon. Cancer Res. 69, 2826–2832 (2009).
Le Poul, E. et al. Functional characterization of human receptors for short chain fatty acids and their role in polymorphonuclear cell activation. J. Biol. Chem. 278, 25481–25489 (2003).
Singh, N. et al. Activation of gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 40, 128–139 (2014).
Tunaru, S. et al. PUMA-G and HM74 are receptors for nicotinic acid and mediate its anti-lipolytic effect. Nat. Med. 9, 352–355 (2003).
Hong, Y.-H. et al. Acetate and propionate short chain fatty acids stimulate adipogenesis via GPCR43. Endocrinology 146, 5092–5099 (2005).
Bjursell, M. et al. Improved glucose control and reduced body fat mass in free fatty acid receptor 2–deficient mice fed a high-fat diet. Am. J. Physiol. Endocrinol. Metab. 300, E211–E220 (2011).
Kimura, I. et al. The gut microbiota suppresses insulin-mediated fat accumulation via the short-chain fatty acid receptor GPR43. Nat. Commun. 4, 1829 (2013).
Tolhurst, G. et al. Short-chain fatty acids stimulate glucagon-like peptide-1 secretion via the G protein–coupled receptor FFAR2. Diabetes 61, 364–371 (2012).
Maslowski, K.M. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461, 1282–1286 (2009).
Garrett, W.S., Gordon, J.I. & Glimcher, L.H. Homeostasis and inflammation in the intestine. Cell 140, 859–870 (2010).
Del Rio, D., Borges, G. & Crozier, A. Berry flavonoids and phenolics: bioavailability and evidence of protective effects. Br. J. Nutr. 104 Suppl 3, S67–S90 (2010).
Possemiers, S., Bolca, S., Verstraete, W. & Heyerick, A. The intestinal microbiome: a separate organ inside the body with the metabolic potential to influence the bioactivity of botanicals. Fitoterapia 82, 53–66 (2011).
Brestoff, J.R. & Artis, D. Commensal bacteria at the interface of host metabolism and the immune system. Nat. Immunol. 14, 676–684 (2013).
Hooper, L.V. & Macpherson, A.J. Immune adaptations that maintain homeostasis with the intestinal microbiota. Nat. Rev. Immunol. 10, 159–169 (2010).
Bouskra, D. et al. Lymphoid tissue genesis induced by commensals through NOD1 regulates intestinal homeostasis. Nature 456, 507–510 (2008).
Moreau, M.C., Ducluzeau, R., Guy-Grand, D. & Muller, M.C. Increase in the population of duodenal immunoglobulin A plasmocytes in axenic mice associated with different living or dead bacterial strains of intestinal origin. Infect. Immun. 21, 532–539 (1978).
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331, 337–341 (2011).
Atarashi, K. et al. Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500, 232–236 (2013).
Mazmanian, S.K., Liu, C.H., Tzianabos, A.O. & Kasper, D.L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122, 107–118 (2005).
Cahenzli, J., Köller, Y., Wyss, M., Geuking, M.B. & McCoy, K.D. Intestinal microbial diversity during early-life colonization shapes long-term IgE levels. Cell Host Microbe 14, 559–570 (2013).
Umetsu, D.T., McIntire, J.J., Akbari, O., Macaubas, C. & DeKruyff, R.H. Asthma: an epidemic of dysregulated immunity. Nat. Immunol. 3, 715–720 (2002).
Mazmanian, S.K., Round, J.L. & Kasper, D.L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453, 620–625 (2008).
Round, J.L. et al. The Toll-like receptor 2 pathway establishes colonization by a commensal of the human microbiota. Science 332, 974–977 (2011).
Olszak, T. et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336, 489–493 (2012).
An, D. et al. Sphingolipids from a symbiotic microbe regulate homeostasis of host intestinal natural killer T cells. Cell 156, 123–133 (2014).
Kawano, T. et al. CD1d-restricted and TCR-mediated activation of va14 NKT cells by glycosylceramides. Science 278, 1626–1629 (1997).
Greer, R.L., Morgun, A. & Shulzhenko, N. Bridging immunity and lipid metabolism by gut microbiota. J. Allergy Clin. Immunol. 132, 253–262, quiz 263 (2013).
Rossi, M., Amaretti, A. & Raimondi, S. Folate production by probiotic bacteria. Nutrients 3, 118–134 (2011).
Kjer-Nielsen, L. et al. MR1 presents microbial vitamin B metabolites to MAIT cells. Nature 491, 717–723 (2012).
Bansal, T., Alaniz, R.C., Wood, T.K. & Jayaraman, A. The bacterial signal indole increases epithelial-cell tight-junction resistance and attenuates indicators of inflammation. Proc. Natl. Acad. Sci. USA 107, 228–233 (2010).
Shimada, Y. et al. Commensal bacteria-dependent indole production enhances epithelial barrier function in the colon. PLoS ONE 8, e80604 (2013).
Fukuda, S. et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543–547 (2011).
Inan, M.S. et al. The luminal short-chain fatty acid butyrate modulates NF-kB activity in a human colonic epithelial cell line. Gastroenterology 118, 724–734 (2000).
Smith, P.M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013).
Sakaguchi, S., Yamaguchi, T., Nomura, T. & Ono, M. Regulatory T cells and immune tolerance. Cell 133, 775–787 (2008).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Kim, S.V. et al. GPR15-mediated homing controls immune homeostasis in the large intestine mucosa. Science 340, 1456–1459 (2013).
Frank, D.N. et al. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl. Acad. Sci. USA 104, 13780–13785 (2007).
Trompette, A. et al. Gut microbiota metabolism of dietary fiber influences allergic airway disease and hematopoiesis. Nat. Med. 20, 159–166 (2014).
Blumberg, R. & Powrie, F. Microbiota, disease, and back to health: a metastable journey. Sci. Transl. Med. 4, 137rv137 (2012).
Vijay-Kumar, M. et al. Metabolic syndrome and altered gut microbiota in mice lacking Toll-like receptor 5. Science 328, 228–231 (2010).
Tillisch, K. et al. Consumption of fermented milk product with probiotic modulates brain activity. Gastroenterology 144, 1394–1401 (2013).
Hsiao, E.Y. et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155, 1451–1463 (2013).
Wang, L., Angley, M.T., Gerber, J.P. & Sorich, M.J. A review of candidate urinary biomarkers for autism spectrum disorder. Biomarkers 16, 537–552 (2011).
Donohoe, D.R. et al. The Warburg effect dictates the mechanism of butyrate-mediated histone acetylation and cell proliferation. Mol. Cell 48, 612–626 (2012).
Acknowledgements
This study was supported by the National Creative Research Initiative Program (no. 20120000231) from National Research Foundation of Korea to W.-J.L. and the Japan Society for the Promotion of Science (no. 24117723) and the Japan Science and Technology Agency (PRESTO) to K.H.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Lee, WJ., Hase, K. Gut microbiota–generated metabolites in animal health and disease. Nat Chem Biol 10, 416–424 (2014). https://doi.org/10.1038/nchembio.1535
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1535
This article is cited by
-
Effects of Radix dichroae extract supplementation on growth performance, oocysts output and gut microbiota in growing lambs with coccidiosis
Veterinary Research Communications (2024)
-
Intestine bacterial community affects the growth of the Pacific white shrimp (Litopenaeus vannamei)
Applied Microbiology and Biotechnology (2024)
-
Analysis of metabolites in human gut: illuminating the design of gut-targeted drugs
Journal of Cheminformatics (2023)
-
MetaPro: a scalable and reproducible data processing and analysis pipeline for metatranscriptomic investigation of microbial communities
Microbiome (2023)
-
Metagenomic analysis of gut microbiome and resistome of Whooper and Black Swans: a one health perspective
BMC Genomics (2023)