Abstract
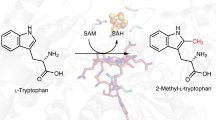
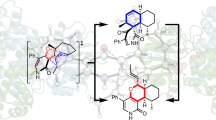
7-carboxy-7-deazaguanine synthase (QueE) catalyzes a key S-adenosyl-L-methionine (AdoMet)- and Mg2+-dependent radical-mediated ring contraction step, which is common to the biosynthetic pathways of all deazapurine-containing compounds. QueE is a member of the AdoMet radical superfamily, which employs the 5′-deoxyadenosyl radical from reductive cleavage of AdoMet to initiate chemistry. To provide a mechanistic rationale for this elaborate transformation, we present the crystal structure of a QueE along with structures of pre- and post-turnover states. We find that substrate binds perpendicular to the [4Fe-4S]-bound AdoMet, exposing its C6 hydrogen atom for abstraction and generating the binding site for Mg2+, which coordinates directly to the substrate. The Burkholderia multivorans structure reported here varies from all other previously characterized members of the AdoMet radical superfamily in that it contains a hypermodified (β6/α3) protein core and an expanded cluster-binding motif, CX14CX2C.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sofia, H.J., Chen, G., Hetzler, B.G., Reyes-Spindola, J.F. & Miller, N.E. Radical SAM, a novel protein superfamily linking unresolved steps in familiar biosynthetic pathways with radical mechanisms: functional characterization using new analysis and information visualization methods. Nucleic Acids Res. 29, 1097–1106 (2001).
Pegg, S.C.-H. et al. Leveraging enzyme structure-function relationships for functional inference and experimental design: the structure-function linkage database. Biochemistry 45, 2545–2555 (2006).
Shisler, K.A. & Broderick, J.B. Emerging themes in radical SAM chemistry. Curr. Opin. Struct. Biol. 22, 701–710 (2012).
Vey, J.L. & Drennan, C.L. Structural insights into radical generation by the radical SAM superfamily. Chem. Rev. 111, 2487–2506 (2011).
Dowling, D.P., Vey, J.L., Croft, A.K. & Drennan, C.L. Structural diversity in the AdoMet radical enzyme superfamily. Biochim. Biophys. Acta. 1824, 1178–1195 (2012).
McCarty, R.M., Somogyi, Á., Lin, G., Jacobsen, N.E. & Bandarian, V. The deazapurine biosynthetic pathway revealed: in vitro enzymatic synthesis of PreQ0 from guanosine 5′-triphosphate in four steps. Biochemistry 48, 3847–3852 (2009).
McCarty, R.M. & Bandarian, V. Biosynthesis of pyrrolopyrimidines. Bioorg. Chem. 43, 15–25 (2012).
Cheek, J. & Broderick, J.B. Direct H atom abstraction from spore photoproduct C-6 initiates DNA repair in the reaction catalyzed by spore photoproduct lyase: evidence for a reversibly generated adenosyl radical intermediate. J. Am. Chem. Soc. 124, 2860–2861 (2002).
Frey, P.A. & Reed, G.H. Pyridoxal-5′-phosphate as the catalyst for radical isomerization in reactions of PLP-dependent aminomutases. Biochim. Biophys. Acta. 1814, 1548–1557 (2011).
McCarty, R.M., Krebs, C. & Bandarian, V. Spectroscopic, steady-state kinetic, and mechanistic characterization of the radical SAM enzyme QueE, which catalyzes a complex cyclization reaction in the biosynthesis of 7-deazapurines. Biochemistry 52, 188–198 (2013).
Duschene, K.S., Veneziano, S.E., Silver, S.C. & Broderick, J.B. Control of radical chemistry in the AdoMet radical enzymes. Curr. Opin. Chem. Biol. 13, 74–83 (2009).
Layer, G., Moser, J., Heinz, D.W., Jahn, D. & Schubert, W.D. Crystal structure of coproporphyrinogen III oxidase reveals cofactor geometry of radical SAM enzymes. EMBO J. 22, 6214–6224 (2003).
Berkovitch, F., Nicolet, Y., Wan, J.T., Jarrett, J.T. & Drennan, C.L. Crystal structure of biotin synthase, an S-adenosylmethionine–dependent radical enzyme. Science 303, 76–79 (2004).
Vey, J.L. et al. Structural basis for glycyl radical formation by pyruvate formate-lyase activating enzyme. Proc. Natl. Acad. Sci. USA 105, 16137–16141 (2008).
Holm, L. & Rosenström, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
Farrar, C.E. & Jarrett, J.T. Protein residues that control the reaction trajectory in S-adenosylmethionine radical enzymes: mutagenesis of asparagine 153 and aspartate 155 in Escherichia coli biotin synthase. Biochemistry 48, 2448–2458 (2009).
Harding, M.M. Small revisions to predicted distances around metal sites in proteins. Acta Crystallogr. D Biol. Crystallogr. 62, 678–682 (2006).
Zheng, H., Chruszcz, M., Lasota, P., Lebioda, L. & Minor, W. Data mining of metal ion environments present in protein structures. J. Inorg. Biochem. 102, 1765–1776 (2008).
Walden, H. et al. Tiny TIM: a small, tetrameric, hyperthermostable triosephosphate isomerase. J. Mol. Biol. 306, 745–757 (2001).
Nagano, N., Orengo, C.A. & Thornton, J.M. One fold with many functions: the evolutionary relationships between TIM barrel families based on their sequences, structures and functions. J. Mol. Biol. 321, 741–765 (2002).
Blair, D.E. & van Aalten, D.M.F. Structures of Bacillus subtilis PdaA, a family 4 carbohydrate esterase, and a complex with N-acetyl-glucosamine. FEBS Lett. 570, 13–19 (2004).
Romier, C., Reuter, K., Suck, D. & Ficner, R. Crystal structure of tRNA-guanine transglycosylase: RNA modification by base exchange. EMBO J. 15, 2850–2857 (1996).
Huang, K., Li, Z., Jia, Y., Dunaway-Mariano, D. & Herzberg, O. Helix swapping between two α/β barrels: crystal structure of phosphoenolpyruvate mutase with bound Mg2+-oxalate. Structure 7, 539–548 (1999).
Takagi, H. et al. Crystal structure of the ribonuclease P protein Ph1877p from hyperthermophilic archaeon Pyrococcus horikoshii OT3. Biochem. Biophys. Res. Commun. 319, 787–794 (2004).
Teplyakov, A. et al. Crystal structure of the Escherichia coli YcdX protein reveals a trinuclear zinc active site. Proteins 51, 315–318 (2003).
Goldman, A., Ollis, D.L. & Steitz, T.A. Crystal structure of muconate lactonizing enzyme at 3 Å resolution. J. Mol. Biol. 194, 143–153 (1987).
Neidhart, D.J. et al. Mechanism of the reaction catalyzed by mandelate racemase. 2. Crystal structure of mandelate racemase at 2.5-Å resolution: identification of the active site and possible catalytic residues. Biochemistry 30, 9264–9273 (1991).
Gromiha, M.M., Pujadas, G., Magyar, C., Selvaraj, S. & Simon, I. Locating the stabilizing residues in (α/β)8 barrel proteins based on hydrophobicity, long-range interactions, and sequence conservation. Proteins 55, 316–329 (2004).
Chatterjee, A. et al. Reconstitution of ThiC in thiamine pyrimidine biosynthesis expands the radical SAM superfamily. Nat. Chem. Biol. 4, 758–765 (2008).
McGlynn, S.E. et al. Identification and characterization of a novel member of the radical AdoMet enzyme superfamily and implications for the biosynthesis of the Hmd hydrogenase active site cofactor. J. Bacteriol. 192, 595–598 (2010).
Paraskevopoulou, C., Fairhurst, S.A., Lowe, D.J., Brick, P. & Onesti, S. The elongator subunit Elp3 contains a Fe4S4 cluster and binds S-adenosylmethionine. Mol. Microbiol. 59, 795–806 (2006).
Marsh, E.N. & Melendez, G.D.R. Adenosylcobalamin enzymes: theory and experiment begin to converge. Biochim. Biophys. Acta 1824, 1154–1164 (2012).
Toraya, T., Honda, S. & Mori, K. Coenzyme B12–dependent diol dehydratase is a potassium ion–requiring calcium metalloenzyme: evidence that the substrate-coordinated metal ion is calcium. Biochemistry 49, 7210–7217 (2010).
Shibata, N. et al. Crystal structures of ethanolamine ammonia-lyase complexed with coenzyme B12 analogs and substrates. J. Biol. Chem. 285, 26484–26493 (2010).
Kamachi, T., Doitomi, K., Takahata, M., Toraya, T. & Yoshizawa, K. Catalytic roles of the metal ion in the substrate-binding site of coenzyme B12-dependent diol dehydratase. Inorg. Chem. 50, 2944–2952 (2011).
Frey, P.A. Lysine 2,3-aminomutase: is adenosylmethionine a poor man's adenosylcobalamin? FASEB J. 7, 662–670 (1993).
Goldman, P.J., Grove, T.L., Booker, S.J. & Drennan, C.L. X-ray analysis of butirosin biosynthetic enzyme BtrN redefines structural motifs for AdoMet radical chemistry. Proc. Natl. Acad. Sci. USA 110, 15949–15954 (2013).
Frazzon, J., Fick, J.R. & Dean, D.R. Biosynthesis of iron-sulphur clusters is a complex and highly conserved process. Biochem. Soc. Trans. 30, 680–685 (2002).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data in oscillation mode. Macromol. Crystallogr. A. 276, 307–326 (1997).
Terwilliger, T.C. et al. Decision-making in structure solution using Bayesian estimates of map quality: the PHENIX AutoSol wizard. Acta Crystallogr. D Biol. Crystallogr. 65, 582–601 (2009).
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Schröder, G.F., Levitt, M. & Brunger, A.T. Super-resolution biomolecular crystallography with low-resolution data. Nature 464, 1218–1222 (2010).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Brunger, A.T. Version 1.2 of the Crystallography and NMR system. Nat. Protoc. 2, 2728–2733 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
French, S. & Wilson, K. On the treatment of negative intensity observations. Acta Crystallogr. A 34, 517–525 (1978).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Painter, J. & Merritt, E.A. Optimal description of a protein structure in terms of multiple groups undergoing TLS motion. Acta Crystallogr. D Biol. Crystallogr. 62, 439–450 (2006).
Painter, J. & Merritt, E.A. TLSMD web server for the generation of multi-group TLS models. J. Appl. Crystallogr. 39, 109–111 (2006).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Bond, C.S. TopDraw: a sketchpad for protein structure topology cartoons. Bioinformatics 19, 311–312 (2003).
Lopes, C.T. et al. Cytoscape Web: an interactive web-based network browser. Bioinformatics 26, 2347–2348 (2010).
Acknowledgements
This work was supported by US National Institutes of Health grant GM72623 (V.B.) with Administrative Supplement GM72623 S01 to V.B. for the collaboration between V.B. and C.L.D.; a Career Award in Biomedical Sciences from the Burroughs Wellcome Fund (V.B.); and a Biological Chemistry Training Grant (GM008804) (R.M.M.). Additionally, C.L.D. is a Howard Hughes Medical Investigator. This work is based on research conducted at beamline X25 of the National Synchrotron Light Source (NSLS) and at the Advanced Photon Source (APS) on the Northeastern Collaborative Access Team (NE-CAT) beamlines. Financial support from NSLS comes from the Offices of Biological and Environmental Research and of Basic Energy Sciences of the US Department of Energy (DOE) and from the National Center for Research Resources (P41RR012408) and the National Institute of General Medical Sciences at the National Institutes of Health (P41GM103473). NE-CAT at APS is supported by grants from the National Center for Research Resources (5P41RR015301-10) and the National Institute of General Medical Sciences at the National Institutes of Health (8 P41 GM103403-10). APS is an Office of Science User Facility operated for the US DOE Office of Science by Argonne National Laboratory and is also supported by the US DOE under contract number DE-AC02-06CH11357. We thank G.L. Holliday and P. Babbitt (University of California San Francisco) and the Enzyme Function Initiative for their analysis of sequences in the AdoMet radical enzyme superfamily, which are available from the Structure Function Linkage Database (http://sfld.rbvi.ucsf.edu/django/superfamily/29/).
Author information
Authors and Affiliations
Contributions
D.P.D. and C.L.D. designed and performed the crystallography experiments. N.A.B., R.M.M., A.P.Y. and V.B. designed and carried out the biochemical experiments. D.P.D., N.A.B., V.B. and C.L.D. contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–12 and Supplementary Tables 1–4. (PDF 7398 kb)
Rights and permissions
About this article
Cite this article
Dowling, D., Bruender, N., Young, A. et al. Radical SAM enzyme QueE defines a new minimal core fold and metal-dependent mechanism. Nat Chem Biol 10, 106–112 (2014). https://doi.org/10.1038/nchembio.1426
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1426
This article is cited by
-
Structural and mechanistic basis for RiPP epimerization by a radical SAM enzyme
Nature Chemical Biology (2023)
-
Crystallographic snapshots of a B12-dependent radical SAM methyltransferase
Nature (2022)
-
QM/MM Study of the Mechanism of the Noncanonical S-Cγ Bond Scission in S-Adenosylmethionine Catalyzed by the CmnDph2 Radical Enzyme
Topics in Catalysis (2022)
-
Structure–function relationships of radical SAM enzymes
Nature Catalysis (2020)
-
Discovery and characterization of the tubercidin biosynthetic pathway from Streptomyces tubercidicus NBRC 13090
Microbial Cell Factories (2018)