Abstract
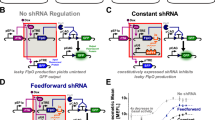
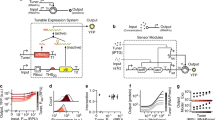
Transient delivery of gene circuits is required in many potential applications of synthetic biology, yet the pre-steady-state processes that dominate this delivery route pose major challenges for robust circuit deployment. Here we show that site-specific recombinases can rectify undesired effects by programmable timing of gene availability in multigene circuits. We exemplify the concept with a proportional sensor for endogenous microRNA (miRNA) and show a marked reduction in its ground state leakage due to desynchronization of the circuit's repressor components and their repression target. The new sensors display a dynamic range of up to 1,000-fold compared to 20-fold in the standard configuration. We applied the approach to classify cell types on the basis of miRNA expression profile and measured >200-fold output differential between positively and negatively identified cells. We also showed major improvements in specificity with cytotoxic output. Our study opens new venues in gene circuit design via judicious temporal control of circuits' genetic makeup.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Wodarz, A. & Nusse, R. Mechanisms of Wnt signaling in development. Annu. Rev. Cell Dev. Biol. 14, 59–88 (1998).
Shimizu, T.S., Tu, Y.H. & Berg, H.C. A modular gradient-sensing network for chemotaxis in Escherichia coli revealed by responses to time-varying stimuli. Mol. Syst. Biol. 6, 382 (2010).
Mansfield, J.H. et al. MicroRNA-responsive 'sensor' transgenes uncover Hox-like and other developmentally regulated patterns of vertebrate microRNA expression. Nat. Genet. 36, 1079–1083 (2004).
Zhang, F., Carothers, J.M. & Keasling, J.D. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat. Biotechnol. 30, 354–359 (2012).
Weber, W. et al. A synthetic mammalian gene circuit reveals antituberculosis compounds. Proc. Natl. Acad. Sci. USA 105, 9994–9998 (2008).
Callura, J.M., Dwyer, D.J., Isaacs, F.J., Cantor, C.R. & Collins, J.J. Tracking, tuning and terminating microbial physiology using synthetic riboregulators. Proc. Natl. Acad. Sci. USA 107, 15898–15903 (2010).
Gardner, T.S., Cantor, C.R. & Collins, J.J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Egbert, R.G. & Klavins, E. Fine-tuning gene networks using simple sequence repeats. Proc. Natl. Acad. Sci. USA 109, 16817–16822 (2012).
Basu, S., Gerchman, Y., Collins, C.H., Arnold, F.H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
de Las Heras, A., Carreno, C.A., Martinez-Garcia, E. & de Lorenzo, V. Engineering input/output nodes in prokaryotic regulatory circuits. FEMS Microbiol. Rev. 34, 842–865 (2010).
Liang, J.C., Chang, A.L., Kennedy, A.B. & Smolke, C.D. A high-throughput, quantitative cell-based screen for efficient tailoring of RNA device activity. Nucleic Acids Res. 40, e154 (2012).
Weber, W. et al. A synthetic time-delay circuit in mammalian cells and mice. Proc. Natl. Acad. Sci. USA 104, 2643–2648 (2007).
Elowitz, M.B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Prindle, A. et al. A sensing array of radically coupled genetic 'biopixels'. Nature 481, 39–44 (2012).
Tabor, J.J., Levskaya, A. & Voigt, C.A. Multichromatic control of gene expression in Escherichia coli. J. Mol. Biol. 405, 315–324 (2011).
Rinaudo, K. et al. A universal RNAi-based logic evaluator that operates in mammalian cells. Nat. Biotechnol. 25, 795–801 (2007).
Deans, T.L., Cantor, C.R. & Collins, J.J. A tunable genetic switch based on RNAi and repressor proteins for regulating gene expression in mammalian cells. Cell 130, 363–372 (2007).
Xie, Z., Wroblewska, L., Prochazka, L., Weiss, R. & Benenson, Y. Multi-input RNAi-based logic circuit for identification of specific cancer cells. Science 333, 1307–1311 (2011).
Haynes, K.A., Ceroni, F., Flicker, D., Younger, A. & Silver, P.A. A sensitive switch for visualizing natural gene silencing in single cells. ACS Synth. Biol. 1, 99–106 (2012).
Biffi, A. et al. Lentiviral hematopoietic stem cell gene therapy benefits metachromatic leukodystrophy. Science 341, 1233158 (2013).
Kota, J. et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 137, 1005–1017 (2009).
Mangan, S. & Alon, U. Structure and function of the feed-forward loop network motif. Proc. Natl. Acad. Sci. USA 100, 11980–11985 (2003).
Basu, S., Mehreja, R., Thiberge, S., Chen, M.T. & Weiss, R. Spatiotemporal control of gene expression with pulse-generating networks. Proc. Natl. Acad. Sci. USA 101, 6355–6360 (2004).
Sontag, E.D. Remarks on feedforward circuits, adaptation, and pulse memory. IET Syst. Biol. 4, 39–51 (2010).
Bleris, L. et al. Synthetic incoherent feedforward circuits show adaptation to the amount of their genetic template. Mol. Syst. Biol. 7, 519 (2011).
Weber, W., Kramer, B.P. & Fussenegger, M. A genetic time-delay circuitry in mammalian cells. Biotechnol. Bioeng. 98, 894–902 (2007).
Dunlop, M.J., Cox, R.S., Levine, J.H., Murray, R.M. & Elowitz, M.B. Regulatory activity revealed by dynamic correlations in gene expression noise. Nat. Genet. 40, 1493–1498 (2008).
Hausser, J. et al. Timescales and bottlenecks in miRNA-dependent gene regulation. Mol. Syst. Biol. 9, 711 (2013).
Hooshangi, S., Thiberge, S. & Weiss, R. Ultrasensitivity and noise propagation in a synthetic transcriptional cascade. Proc. Natl. Acad. Sci. USA 102, 3581–3586 (2005).
Schnütgen, F. et al. A directional strategy for monitoring Cre-mediated recombination at the cellular level in the mouse. Nat. Biotechnol. 21, 562–565 (2003).
Weber, W. et al. Macrolide-based transgene control in mammalian cells and mice. Nat. Biotechnol. 20, 901–907 (2002).
Matsuda, T. & Cepko, C.L. Controlled expression of transgenes introduced by in vivo electroporation. Proc. Natl. Acad. Sci. USA 104, 1027–1032 (2007).
Singh, J. & Padgett, R.A. Rates of in situ transcription and splicing in large human genes. Nat. Struct. Mol. Biol. 16, 1128–1133 (2009).
Leisner, M., Bleris, L., Lohmueller, J., Xie, Z. & Benenson, Y. Rationally designed logic integration of regulatory signals in mammalian cells. Nat. Nanotechnol. 5, 666–670 (2010).
Kanegae, Y. et al. High-level expression by tissue/cancer-specific promoter with strict specificity using a single-adenoviral vector. Nucleic Acids Res. 39, e7 (2011).
Andrade-Rozental, A.F. et al. Gap junctions: the “kiss of death” and the “kiss of life”. Brain Res. Brain Res. Rev. 32, 308–315 (2000).
Mesnil, M., Piccoli, C., Tiraby, G., Willecke, K. & Yamasaki, H. Bystander killing of cancer cells by herpes simplex virus thymidine kinase gene is mediated by connexins. Proc. Natl. Acad. Sci. USA 93, 1831–1835 (1996).
Mohr, L. et al. Gene therapy of hepatocellular carcinoma in vitro and in vivo in nude mice by adenoviral transfer of the Escherichia coli purine nucleoside phosphorylase gene. Hepatology 31, 606–614 (2000).
Mercer, K.E., Ahn, C.E., Coke, A., Compadre, C.M. & Drake, R.R. Mutation of herpesvirus thymidine kinase to generate ganciclovir-specific kinases for use in cancer gene therapies. Protein Eng. 15, 903–911 (2002).
Backman, K., Oconnor, M.J., Maruya, A. & Erfle, M. Use of synchronous site-specific recombination in vivo to regulate gene-expression. Bio/Technology 2, 1045–1049 (1984).
Dale, E.C. & Ow, D.W. Gene-transfer with subsequent removal of the selection gene from the host genome. Proc. Natl. Acad. Sci. USA 88, 10558–10562 (1991).
Benenson, Y. Biomolecular computing systems: principles, progress and potential. Nat. Rev. Genet. 13, 455–468 (2012).
Ham, T.S., Lee, S.K., Keasling, J.D. & Arkin, A.P. Design and construction of a double inversion recombination switch for heritable sequential genetic memory. PLoS ONE 3, e2815 (2008).
Friedland, A.E. et al. Synthetic gene networks that count. Science 324, 1199–1202 (2009).
Siuti, P., Yazbek, J. & Lu, T.K. Synthetic circuits integrating logic and memory in living cells. Nat. Biotechnol. 31, 448–452 (2013).
Bonnet, J., Yin, P., Ortiz, M.E., Subsoontorn, P. & Endy, D. Amplifying genetic logic gates. Science 340, 599–603 (2013).
Prochazka, L., Angelici, B., Haefliger, B. & Benenson, Y. Highly modular bow-tie gene circuits with programmable dynamic behaviour. Nat. Commun. 5, 4729 (2014).
Weber, W., Kramer, B.P., Fux, C., Keller, B. & Fussenegger, M. Novel promoter/transactivator configurations for macrolide- and streptogramin-responsive transgene expression in mammalian cells. J. Gene Med. 4, 676–686 (2002).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Acknowledgements
The work was funded the US National Institutes of Health and National Cancer Institute grant 5R01CA155320 and a European Research Council starting grant CellControl. We wish to thank the M. Fussenegger lab in our department for the ET-ETR inducible system, K. Hoelle and L. Prochazka for plasmids and Benenson lab members for discussions. We thank M. Dessing and V. Jaeggin for assistance with flow cytometry and T. Horn for help with imaging. We thank R. Kellogg from S. Tay's laboratory in our department for help with viability assays.
Author information
Authors and Affiliations
Contributions
N.L. conceived of the project, designed experiments, performed all the experiments, analyzed data and wrote the paper. Y.B. conceived of and supervised the project, designed experiments, analyzed data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
Declaration: The results of this manuscript have been submitted as a priority filing to the European Patent office under filing number EP 14001960.5 with the title `Near-perfect digital switching in a synthetic biosensor circuit achieved through temporal control of circuit's genetic makeup.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–19 and Supplementary Tables 1–16. (PDF 26817 kb)
Rights and permissions
About this article
Cite this article
Lapique, N., Benenson, Y. Digital switching in a biosensor circuit via programmable timing of gene availability. Nat Chem Biol 10, 1020–1027 (2014). https://doi.org/10.1038/nchembio.1680
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1680
This article is cited by
-
Engineering intelligent chassis cells via recombinase-based MEMORY circuits
Nature Communications (2024)
-
Next generation synthetic memory via intercepting recombinase function
Nature Communications (2023)
-
Oncolytic adenovirus programmed by synthetic gene circuit for cancer immunotherapy
Nature Communications (2019)
-
Genetic programs can be compressed and autonomously decompressed in live cells
Nature Nanotechnology (2018)
-
Synthetic RNA-based logic computation in mammalian cells
Nature Communications (2018)