Abstract
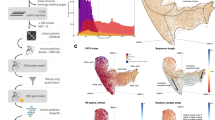
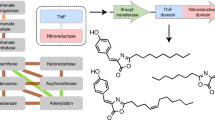
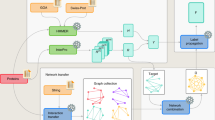
Millions of protein database entries are not assigned reliable functions, preventing the full understanding of chemical diversity in living organisms. Here, we describe an integrated strategy for the discovery of various enzymatic activities catalyzed within protein families of unknown or little known function. This approach relies on the definition of a generic reaction conserved within the family, high-throughput enzymatic screening on representatives, structural and modeling investigations and analysis of genomic and metabolic context. As a proof of principle, we investigated the DUF849 Pfam family and unearthed 14 potential new enzymatic activities, leading to the designation of these proteins as β-keto acid cleavage enzymes. We propose an in vivo role for four enzymatic activities and suggest key residues for guiding further functional annotation. Our results show that the functional diversity within a family may be largely underestimated. The extension of this strategy to other families will improve our knowledge of the enzymatic landscape.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Galperin, M.Y. & Koonin, E.V. From complete genome sequence to 'complete' understanding? Trends Biotechnol. 28, 398–406 (2010).
Roberts, R.J. Identifying protein function–a call for community action. PLoS Biol. 2, E42 (2004).
Karp, P.D. Call for an enzyme genomics initiative. Genome Biol. 5, 401 (2004).
Hanson, A.D., Pribat, A., Waller, J.C. & de Crecy-Lagard, V. 'Unknown' proteins and 'orphan' enzymes: the missing half of the engineering parts list–and how to find it. Biochem. J. 425, 1–11 (2010).
Gifford, L.K., Carter, L.G., Gabanyi, M.J., Berman, H.M. & Adams, P.D. The Protein Structure Initiative Structural Biology Knowledgebase Technology Portal: a structural biology web resource. J. Struct. Funct. Genomics 13, 57–62 (2012).
Gerlt, J.A. et al. The Enzyme Function Initiative. Biochemistry 50, 9950–9962 (2011).
Lukk, T. et al. Homology models guide discovery of diverse enzyme specificities among dipeptide epimerases in the enolase superfamily. Proc. Natl. Acad. Sci. USA 109, 4122–4127 (2012).
Finn, R.D. et al. The Pfam protein families database. Nucleic Acids Res. 38, D211–D222 (2010).
Furnham, N., Garavelli, J.S., Apweiler, R. & Thornton, J.M. Missing in action: enzyme functional annotations in biological databases. Nat. Chem. Biol. 5, 521–525 (2009).
Huang, H. et al. Divergence of structure and function in the haloacid dehalogenase enzyme superfamily: Bacteroides thetaiotaomicron BT2127 is an inorganic pyrophosphatase. Biochemistry 50, 8937–8949 (2011).
Schnoes, A.M., Brown, S.D., Dodevski, I. & Babbitt, P.C. Annotation error in public databases: misannotation of molecular function in enzyme superfamilies. PLoS Comput. Biol. 5, e1000605 (2009).
Lespinet, O. & Labedan, B. Orphan enzymes? Science 307, 42 (2005).
Kreimeyer, A. et al. Identification of the last unknown genes in the fermentation pathway of lysine. J. Biol. Chem. 282, 7191–7197 (2007).
Bellinzoni, M. et al. 3-Keto-5-aminohexanoate cleavage enzyme: a common fold for an uncommon Claisen-type condensation. J. Biol. Chem. 286, 27399–27405 (2011).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 40, D742–D753 (2012).
Deniélou, Y.P., Sagot, M.F., Boyer, F. & Viari, A. Bacterial syntenies: an exact approach with gene quorum. BMC Bioinformatics 12, 193 (2011).
de Melo-Minardi, R.C., Bastard, K. & Artiguenave, F. Identification of subfamily-specific sites based on active sites modeling and clustering. Bioinformatics 26, 3075–3082 (2010).
Strehl, A. & Ghosh, J. Cluster ensembles—a knowledge reuse framework for combining partitionings. J. Mach. Learn. Res. 3, 583–617 (2002).
Pan, H., Bao, W., Xie, Z., Zhang, J. & Li, Y. Molecular cloning and characterization of a cis-epoxysuccinate hydrolase from Bordetella sp. BK-52. J. Microbiol. Biotechnol. 20, 659–665 (2010).
Bao, W. et al. Analysis of essential amino acid residues for catalytic activity of cis-epoxysuccinate hydrolase from Bordetella sp. BK-52. Appl. Microbiol. Biotechnol. http://dx.doi.org/10.1007/s00253-013-5019-2 (2013).
Pelletier, E. et al. “Candidatus Cloacamonas acidaminovorans”: genome sequence reconstruction provides a first glimpse of a new bacterial division. J. Bacteriol. 190, 2572–2579 (2008).
Uanschou, C., Frieht, R. & Pittner, F. What to learn from a comparative genomic sequence analysis of L-carnitine dehydrogenase. Monatsh. Chem. 136, 1365–1381 (2005).
Wargo, M.J. & Hogan, D.A. Identification of genes required for Pseudomonas aeruginosa carnitine catabolism. Microbiology 155, 2411–2419 (2009).
Bar-Even, A. et al. The moderately efficient enzyme: evolutionary and physicochemical trends shaping enzyme parameters. Biochemistry 50, 4402–4410 (2011).
Kleber, H.P. Bacterial carnitine metabolism. FEMS Microbiol. Lett. 147, 1–9 (1997).
Collier, L.S., Gaines, G.L. III & Neidle, E.L. Regulation of benzoate degradation in Acinetobacter sp. strain ADP1 by BenM, a LysR-type transcriptional activator. J. Bacteriol. 180, 2493–2501 (1998).
Yalpani, M., Willecke, K. & Lynen, F. Triacetic acid lactone, a derailment product of fatty acid biosynthesis. Eur. J. Biochem. 8, 495–502 (1969).
Xie, D. et al. Microbial synthesis of triacetic acid lactone. Biotechnol. Bioeng. 93, 727–736 (2006).
Monticello, D.J. & Costilow, R.N. Interconversion of valine and leucine by Clostridium sporogenes. J. Bacteriol. 152, 946–949 (1982).
Magrane, M. & Consortium, U. UniProt Knowledgebase: a hub of integrated protein data. Database (Oxford) 2011, bar009 (2011).
Howe, K., Bateman, A. & Durbin, R. QuickTree: building huge neighbour-joining trees of protein sequences. Bioinformatics 18, 1546–1547 (2002).
Katoh, K., Kuma, K., Toh, H. & Miyata, T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–518 (2005).
Brown, D.P., Krishnamurthy, N. & Sjolander, K. Automated protein subfamily identification and classification. PLoS Comput. Biol. 3, e160 (2007).
Fisher, D. Knowledge acquisition via incremental conceptual clustering. Mach. Learn. 2, 139–172 (1987).
Puranen, J.S., Vainio, M.J. & Johnson, M.S. Accurate conformation-dependent molecular electrostatic potentials for high-throughput in silico drug discovery. J. Comput. Chem. 31, 1722–1732 (2010).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Trott, O. & Olson, A.J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386 (2000).
Ralser, M. et al. An efficient and economic enhancer mix for PCR. Biochem. Biophys. Res. Commun. 347, 747–751 (2006).
Aslanidis, C. & de Jong, P.J. Ligation-independent cloning of PCR products (LIC-PCR). Nucleic Acids Res. 18, 6069–6074 (1990).
Moriyama, T. & Srere, P.A. Purification of rat heart and rat liver citrate synthases. Physical, kinetic, and immunological studies. J. Biol. Chem. 246, 3217–3223 (1971).
Swart, M., Snijders, J.G. & van Duijnenb, Th.P. Polarizabilities of amino acid residues. J. Comp. Meth. Sci. Eng. 4, 419–425 (2004).
Acknowledgements
We are grateful to M. Besnard-Gonnet, D. Baud and A. Fossey for excellent technical assistance and to S. Tricot and L. Stuani for MS analyses. We also thank P.L. Saaidi, G. Cohen and M. Stam for helpful discussions and D. Roche, P. Bowe and A. Tolonen for useful comments on the manuscript. This work was supported by grants from Commissariat à l'énergie atomique et aux énergies alternatives (CEA), the CNRS and the University of Evry. A.A.T. Smith has been supported by the MICROME project, European Union Framework Program 7 Collaborative Project (222886-2).
Author information
Authors and Affiliations
Contributions
D.V. and M.S. designed and supervised this study. K.B. and A.A.T.S. conducted all of the bioinformatics analyses. F.A. and R.D.M.-M. initiated structural bioinformatics. C.V.-V. and A.Z. contributed to the chemical sections. K.B., A.A.T.S., D.V. and M.S. wrote the manuscript. C.M. and J.W. contributed to the design of this study. A.K. gave support to metabolic analysis. A.P. and V.D.B. supervised experiments, and J.-L.P., V.P., A.D., A.M., N.P., M.B., C.L. and C.P. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Data Set descriptions, Supplementary Results, Supplementary Notes 1–4, Supplementary Tables 1–6 and Supplementary Figures 1–7. (PDF 3235 kb)
Supplementary Data Set 1
Multiple sequence alignment of the 725 sequences of the DUF849 family. (TXT 278 kb)
Supplementary Data Set 2
The different clusters of the 725 sequences depending on active site composition, genomic context, phylogeny and sequence similarity. (XLSX 65 kb)
Supplementary Data Set 3
Primer sequences of the 322 candidates proteins. (XLSX 49 kb)
Supplementary Data Set 4
Pre-process data of the enzymatic screening. (XLSX 78 kb)
Supplementary Data Set 5
The “active site” hierarchical tree of the 725 sequences of the DUF849 family in nexus format. (TXT 10 kb)
Rights and permissions
About this article
Cite this article
Bastard, K., Smith, A., Vergne-Vaxelaire, C. et al. Revealing the hidden functional diversity of an enzyme family. Nat Chem Biol 10, 42–49 (2014). https://doi.org/10.1038/nchembio.1387
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1387
This article is cited by
-
Identification of putative essential protein domains from high-density transposon insertion sequencing
Scientific Reports (2022)
-
Role of carnitine in adaptation of Chromohalobacter salexigens DSM 3043 and its mutants to osmotic and temperature stress in defined medium
Extremophiles (2022)
-
XszenFHal, a novel tryptophan 5-halogenase from Xenorhabdus szentirmaii
AMB Express (2019)
-
Exploring natural biodiversity to expand access to microbial terpene synthesis
Microbial Cell Factories (2019)
-
A family of native amine dehydrogenases for the asymmetric reductive amination of ketones
Nature Catalysis (2019)