Abstract
Many asymmetrically dividing cells unequally partition cellular structures according to age. Yet, it is unclear how cells differentiate pre-existing from newly synthesized material. Yeast cells segregate the spindle pole body (SPB, centrosome equivalent) inherited from the previous mitosis to the bud, while keeping the new one in the mother cell. Here, we show that the SPB inheritance network (SPIN), comprising the kinases Swe1 (also known as Wee1) and Kin3 (also known as Nek2) and the acetyltransferase NuA4 (also known as Tip60), distinguishes pre-existing from new SPBs. Swe1 phosphorylated Nud1 (orthologous to Centriolin) on young SPBs as they turned into pre-existing ones. The subsequent inactivation of Swe1 protected newly assembling SPBs from being marked. Kin3 and NuA4 maintained age marks on SPBs through following divisions. Downstream of SPIN, the Hippo regulator Bfa1–Bub2 bound the marked SPB, directed the spindle-positioning protein Kar9 towards it and drove its partition to the bud. Thus, coordination of SPIN activity and SPB assembly encodes age onto SPBs to enable their age-dependent segregation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Inaba, M. & Yamashita, Y. M. Asymmetric stem cell division: precision for robustness. Cell Stem Cell 11, 461–469 (2012).
Yadlapalli, S. & Yamashita, Y. M. Chromosome-specific nonrandom sister chromatid segregation during stem-cell division. Nature 498, 251–254 (2013).
Shinin, V., Gayraud-Morel, B., Gomes, D. & Tajbakhsh, S. Asymmetric division and cosegregation of template DNA strands in adult muscle satellite cells. Nat. Cell Biol. 8, 677–687 (2006).
Cairns, J. Mutation selection and the natural history of cancer. Nature 255, 197–200 (1975).
Tran, V., Lim, C., Xie, J. & Chen, X. Asymmetric division of Drosophila male germline stem cell shows asymmetric histone distribution. Science 338, 679–682 (2012).
Xie, J. et al. Histone H3 threonine phosphorylation regulates asymmetric histone inheritance in the Drosophila male germline. Cell 163, 920–933 (2015).
Katajisto, P. et al. Stem cells. Asymmetric apportioning of aged mitochondria between daughter cells is required for stemness. Science 348, 340–343 (2015).
McFaline-Figueroa, J. R. et al. Mitochondrial quality control during inheritance is associated with lifespan and mother–daughter age asymmetry in budding yeast. Aging Cell 10, 885–895 (2011).
Wang, X. et al. Asymmetric centrosome inheritance maintains neural progenitors in the neocortex. Nature 461, 947–955 (2009).
Yamashita, Y. M., Mahowald, A. P., Perlin, J. R. & Fuller, M. T. Asymmetric inheritance of mother versus daughter centrosome in stem cell division. Science 315, 518–521 (2007).
Salzmann, V. et al. Centrosome-dependent asymmetric inheritance of the midbody ring in Drosophila germline stem cell division. Mol. Biol. Cell 25, 267–275 (2014).
Januschke, J., Llamazares, S., Reina, J. & Gonzalez, C. Drosophila neuroblasts retain the daughter centrosome. Nat. Commun. 2, 243 (2011).
Conduit, P. T. & Raff, J. W. Cnn dynamics drive centrosome size asymmetry to ensure daughter centriole retention in Drosophila neuroblasts. Curr. Biol. 20, 2187–2192 (2010).
Paridaen, J. T., Wilsch-Brauninger, M. & Huttner, W. B. Asymmetric inheritance of centrosome-associated primary cilium membrane directs ciliogenesis after cell division. Cell 155, 333–344 (2013).
Azimzadeh, J. & Bornens, M. Structure and duplication of the centrosome. J. Cell Sci. 120, 2139–2142 (2007).
Fu, J., Hagan, I. M. & Glover, D. M. The centrosome and its duplication cycle. Cold Spring Harb. Perspect. Biol. 7, a015800 (2015).
Vorobjev, I. A. & Chentsov, Yu. S. Centrioles in the cell cycle. I. Epithelial cells. J. Cell Biol. 93, 938–949 (1982).
Paintrand, M., Moudjou, M., Delacroix, H. & Bornens, M. Centrosome organization and centriole architecture: their sensitivity to divalent cations. J. Struct. Biol. 108, 107–128 (1992).
Winey, M. & O’Toole, E. Centriole structure. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, 20130457 (2014).
Yoder, T. J., Pearson, C. G., Bloom, K. & Davis, T. N. The Saccharomyces cerevisiae spindle pole body is a dynamic structure. Mol. Biol. Cell 14, 3494–3505 (2003).
Pereira, G., Tanaka, T. U., Nasmyth, K. & Schiebel, E. Modes of spindle pole body inheritance and segregation of the Bfa1p-Bub2p checkpoint protein complex. EMBO J. 20, 6359–6370 (2001).
Hotz, M. et al. Spindle pole bodies exploit the mitotic exit network in metaphase to drive their age-dependent segregation. Cell 148, 958–972 (2012).
Liakopoulos, D., Kusch, J., Grava, S., Vogel, J. & Barral, Y. Asymmetric loading of Kar9 onto spindle poles and microtubules ensures proper spindle alignment. Cell 112, 561–574 (2003).
Maekawa, H., Usui, T., Knop, M. & Schiebel, E. Yeast Cdk1 translocates to the plus end of cytoplasmic microtubules to regulate bud cortex interactions. EMBO J. 22, 438–449 (2003).
Kusch, J., Liakopoulos, D. & Barral, Y. Spindle asymmetry: a compass for the cell. Trends Cell Biol. 13, 562–569 (2003).
Siller, K. H. & Doe, C. Q. Spindle orientation during asymmetric cell division. Nat. Cell Biol. 11, 365–374 (2009).
Masuda, H., Fong, C. S., Ohtsuki, C., Haraguchi, T. & Hiraoka, Y. Spatiotemporal regulations of Wee1 at the G2/M transition. Mol. Biol. Cell 22, 555–569 (2011).
Mitchell, L. et al. mChIP-KAT-MS, a method to map protein interactions and acetylation sites for lysine acetyltransferases. Proc. Natl Acad. Sci. USA 110, E1641–E1650 (2013).
Grallert, A., Krapp, A., Bagley, S., Simanis, V. & Hagan, I. M. Recruitment of NIMA kinase shows that maturation of the S. pombe spindle-pole body occurs over consecutive cell cycles and reveals a role for NIMA in modulating SIN activity. Genes Dev. 18, 1007–1021 (2004).
Cid, V. J., Shulewitz, M. J., McDonald, K. L. & Thorner, J. Dynamic localization of the Swe1 regulator Hsl7 during the Saccharomyces cerevisiae cell cycle. Mol. Biol. Cell 12, 1645–1669 (2001).
Doyon, Y., Selleck, W., Lane, W. S., Tan, S. & Cote, J. Structural and functional conservation of the NuA4 histone acetyltransferase complex from yeast to humans. Mol. Cell. Biol. 24, 1884–1896 (2004).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Longtine, M. S. et al. Septin-dependent assembly of a cell cycle-regulatory module in Saccharomyces cerevisiae. Mol. Cell. Biol. 20, 4049–4061 (2000).
Sia, R. A., Herald, H. A. & Lew, D. J. Cdc28 tyrosine phosphorylation and the morphogenesis checkpoint in budding yeast. Mol. Biol. Cell 7, 1657–1666 (1996).
McMillan, J. N. et al. The morphogenesis checkpoint in Saccharomyces cerevisiae: cell cycle control of Swe1p degradation by Hsl1p and Hsl7p. Mol. Cell. Biol. 19, 6929–6939 (1999).
Keaton, M. A. et al. Differential susceptibility of yeast S and M phase CDK complexes to inhibitory tyrosine phosphorylation. Curr. Biol. 17, 1181–1189 (2007).
Booher, R. N., Deshaies, R. J. & Kirschner, M. W. Properties of Saccharomyces cerevisiae wee1 and its differential regulation of p34CDC28 in response to G1 and G2 cyclins. EMBO J. 12, 3417–3426 (1993).
Hu, F., Gan, Y. & Aparicio, O. M. Identification of Clb2 residues required for Swe1 regulation of Clb2-Cdc28 in Saccharomyces cerevisiae. Genetics 179, 863–874 (2008).
Hu, F. & Aparicio, O. M. Swe1 regulation and transcriptional control restrict the activity of mitotic cyclins toward replication proteins in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 102, 8910–8915 (2005).
Amon, A., Surana, U., Muroff, I. & Nasmyth, K. Regulation of p34CDC28 tyrosine phosphorylation is not required for entry into mitosis in S. cerevisiae. Nature 355, 368–371 (1992).
Sorger, P. K. & Murray, A. W. S-phase feedback control in budding yeast independent of tyrosine phosphorylation of p34cdc28. Nature 355, 365–368 (1992).
Bartholomew, C. R., Woo, S. H., Chung, Y. S., Jones, C. & Hardy, C. F. Cdc5 interacts with the Wee1 kinase in budding yeast. Mol. Cell. Biol. 21, 4949–4959 (2001).
Keck, J. M. et al. A cell cycle phosphoproteome of the yeast centrosome. Science 332, 1557–1561 (2011).
Espinoza, F. H. et al. Cak1 is required for Kin28 phosphorylation and activation in vivo. Mol. Cell. Biol. 18, 6365–6373 (1998).
Remy, I. & Michnick, S. W. Application of protein-fragment complementation assays in cell biology. Biotechniques 42, 137–145 (2007).
Barral, Y., Parra, M., Bidlingmaier, S. & Snyder, M. Nim1-related kinases coordinate cell cycle progression with the organization of the peripheral cytoskeleton in yeast. Genes Dev. 13, 176–187 (1999).
Verzijlbergen, K. F. et al. Recombination-induced tag exchange to track old and new proteins. Proc. Natl Acad. Sci. USA 107, 64–68 (2010).
Burns, S. et al. Structured illumination with particle averaging reveals novel roles for yeast centrosome components during duplication. eLife 4, e08586 (2015).
Monje-Casas, F. & Amon, A. Cell polarity determinants establish asymmetry in MEN signaling. Dev. Cell 16, 132–145 (2009).
Pereira, G., Hofken, T., Grindlay, J., Manson, C. & Schiebel, E. The Bub2p spindle checkpoint links nuclear migration with mitotic exit. Mol. Cell 6, 1–10 (2000).
Caydasi, A. K. & Pereira, G. Spindle alignment regulates the dynamic association of checkpoint proteins with yeast spindle pole bodies. Dev. Cell 16, 146–156 (2009).
Scarfone, I. et al. Asymmetry of the budding yeast Tem1 GTPase at spindle poles is required for spindle positioning but not for mitotic exit. PLoS Genet. 11, e1004938 (2015).
Bardin, A. J. & Amon, A. Men and sin: what’s the difference? Nat. Rev. Mol. Cell Biol. 2, 815–826 (2001).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Avena, J. S. et al. Licensing of yeast centrosome duplication requires phosphoregulation of sfi1. PLoS Genet. 10, e1004666 (2014).
Elserafy, M. et al. Molecular mechanisms that restrict yeast centrosome duplication to one event per cell cycle. Curr. Biol. 24, 1456–1466 (2014).
Bardin, A. J., Visintin, R. & Amon, A. A mechanism for coupling exit from mitosis to partitioning of the nucleus. Cell 102, 21–31 (2000).
Ro, H. S., Song, S. & Lee, K. S. Bfa1 can regulate Tem1 function independently of Bub2 in the mitotic exit network of Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 99, 5436–5441 (2002).
Januschke, J. & Gonzalez, C. The interphase microtubule aster is a determinant of asymmetric division orientation in Drosophila neuroblasts. J. Cell Biol. 188, 693–706 (2010).
Januschke, J. et al. Centrobin controls mother-daughter centriole asymmetry in Drosophila neuroblasts. Nat. Cell Biol. 15, 241–248 (2013).
Stumpff, J., Kellogg, D. R., Krohne, K. A. & Su, T. T. Drosophila Wee1 interacts with members of the gammaTURC and is required for proper mitotic-spindle morphogenesis and positioning. Curr. Biol. 15, 1525–1534 (2005).
Boutros, R. & Ducommun, B. Asymmetric localization of the CDC25B phosphatase to the mother centrosome during interphase. Cell Cycle 7, 401–406 (2008).
Zhang, S. M. et al. HIV-1 Tat impairs cell cycle control by targeting the Tip60, Plk1 and cyclin B1 ternary complex. Cell Cycle 11, 1217–1234 (2012).
Jeong, Y., Lee, J., Kim, K., Yoo, J. C. & Rhee, K. Characterization of NIP2/centrobin, a novel substrate of Nek2, and its potential role in microtubule stabilization. J. Cell Sci. 120, 2106–2116 (2007).
Zou, C. et al. Centrobin: a novel daughter centriole-associated protein that is required for centriole duplication. J. Cell Biol. 171, 437–445 (2005).
Mardin, B. R. et al. Components of the Hippo pathway cooperate with Nek2 kinase to regulate centrosome disjunction. Nat. Cell Biol. 12, 1166–1176 (2010).
Davidow, L. S., Goetsch, L. & Byers, B. Preferential occurrence of nonsister spores in two-spored asci of Saccharomyces cerevisiae: evidence for regulation of spore-wall formation by the spindle pole body. Genetics 94, 581–595 (1980).
Gordon, O. et al. Nud1p, the yeast homolog of Centriolin, regulates spindle pole body inheritance in meiosis. EMBO J. 25, 3856–3868 (2006).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Boettcher, B., Marquez-Lago, T. T., Bayer, M., Weiss, E. L. & Barral, Y. Nuclear envelope morphology constrains diffusion and promotes asymmetric protein segregation in closed mitosis. J. Cell Biol. 197, 921–937 (2012).
Acknowledgements
We thank J. Thorner, M. Winey, V. Simanis, M. Aebi, C. Weirich, F. Caudron, J. Vogel and her laboratory members, and the former and current members of the Barral laboratory for discussions and comments on the manuscript, R. Dechant, A. Lehmann and ScopeM for technical help and L. Pillus (University of California, USA), S. Piatti (CNRS, France), J. Vogel (McGill University, Canada), J. Thorner (UC Berkeley, USA), S. Michnick (Université de Montréal, Canada) and F. Winston (Harvard Medical School, USA) for strains and reagents. M.R. was supported by a NSERC Canadian Graduate Scholarship, K.B. by a grant from the Canadian Institutes of Health Research (http://www.cihr-irsc.gc.ca/e/193.html; MOP-142403) and J.L. and Y.B. were supported by ETHZ and grants from the Swiss National Science Foundation and the European Research Council to Y.B.
Author information
Authors and Affiliations
Contributions
J.L., M.H. and M.R. conducted the experiments and analysed the data, J.L., K.B. and Y.B. designed the experiments, and J.L. and Y.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Inactivation of modifying enzymes that localise to or post-translationally modify SPBs or centrosomes and analysis of effect on SPB inheritance.
(a) Asymmetry index of Spc42 tagged with mCherry in meta- and anaphase (metaphase n = 139, anaphase n = 100 cells pooled from three independent experiments). Scale bars, 2 μm. (b) Quantification of anaphase cells of indicated genotype segregating the new SPB into the bud (%) (n = 3 except WT n = 4 and sir2Δ n = 1, independent experiments with a total of >40 cells analysed per experiment). Student t-test was performed to test significance. For all panels: ∗∗∗∗P < 0.0001, ∗∗∗P < 0.001, ∗∗P < 0.01, NS = non-significant. All error bars represent mean ± s.d.
Supplementary Figure 2 The SPIN does not impact metaphase progression, spindle alignment, astral microtubule length and occupancy on pre-existing SPBs.
(a,b) Quantification of time from SPB separation to spindle alignment (a) or to early anaphase (b) in cells of indicated genotype segregating the pre-existing SPB into the bud (bright colour) or the new SPB into the bud (dim colour). Spc42-mCherry was used as SPB marker and CFP-Tub1 as spindle marker (n = 3 independent experiments with a total of >60 cells per genotype analysed). (c) Quantification of microtubule length over time after SPB separation (μm) in cells of indicated genotype segregating the pre-existing SPB into the bud (bright colour) or the new SPB into the bud (dim colour). Spc42-mCherry was used as SPB marker, Bik1-3xGFP as microtubule marker and CFP-Tub1 as spindle marker (n = 3 independent experiments with a total of >60 cells per genotype analysed) (d) Quantification of anaphase cells of indicated genotype segregating the new SPB into the bud (%) (n = 3 independent experiments with a total of >120 cells per genotype analysed). For all panels: All statistical significances were calculated using two-tailed Student t-tests, NS = non-significant. All error bars represent mean ± s.d.
Supplementary Figure 3 NuA4 promotes SPB inheritance likely by modifying Nud1 and Spc72
(Quantification of anaphase cells of indicated genotype segregating the new SPB into the bud (%) (n = 3 independent experiments, except WT, SPC72-TAP, nud1-44, yaf9Δ nud1-44, NUD1-TAP and NUD1K35R yaf9Δ n = 4, with a total of >120 cells per genotype analysed). One way ANOVA was performed to test significance, ∗∗∗P < 0.001, ∗P < 0.05, NS = non-significant. All error bars represent mean ± s.d.
Supplementary Figure 4 The specification of young and old SPBs involves different mechanisms.
(a) Quantification of haploid and diploid anaphase cells of indicated genotype with the new SPB in the bud of snapshots or long-term imaging conditions (see material and methods) (n = 3 independent experiments with a total of >120 cells per genotype analysed). (b) Swe1-6xHA-AID detection before and after auxin addition by immunoblotting using a α-HA antibody. (c) Quantification of time from bud emergence to anaphase (1) and from anaphase to cytokinesis (2) in haploid cells of indicated genotype (n = 3 independent experiments with a total of >60 cells per genotype analysed). (d) Quantification of diploid anaphase cells of indicated genotype with the new SPB segregated into the bud in the young SPB lineage (green), old SPB lineage (red) and average of both SPB lineages (blue) (%) (n = 3 independent experiments with a total of >120 cells per genotype analysed). (e) Quantification of haploid anaphase cells of indicated genotype with the new SPB segregated into the bud in the young SPB lineage (green), old SPB lineage (red) and average of both SPB lineages (blue) (%) (n = 3 independent experiments, except WT and swe1Δn = 4 with a total of >120 cells per genotype analysed). For all panels: All statistical significances were calculated using one-way ANOVA, ∗∗∗∗P < 0.0001, ∗∗P < 0.01, ∗P < 0.05, NS = non-significant. All error bars represent mean ± s.d.
Supplementary Figure 5 Swe1 promotes SPB specification by modifying Nud1 on the pre-existing SPB.
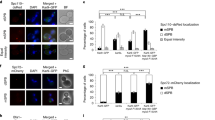
(a) Quantification of anaphase cells of indicated genotype segregating the new SPB into the bud (%) (n = 3, except WT and NUD1-TAP n = 4 independent experiments with a total of >120 cells per genotype analysed). (b) Quantification of haploid anaphase cells of indicated genotype with the new SPB segregated into the bud in the young SPB lineage (green), old SPB lineage (red) and average of both SPB lineages (blue) (%) (n = 3 independent, except WT and swe1Δ n = 4 experiments with a total of >120 cells per genotype analysed). (c) Fraction of anaphase cells of indicated genotype in an asynchronous population (%) (n = 3 independent experiments with a total of >120 cells per genotype analysed). (d) Representative images and quantification of SPOC integrity. Cells of indicated phenotype were grown at 25 °C and shifted to 30 °C for 3 h. SPOC deficiency was quantified as the fraction of cells with misaligned spindles (blue bar) and multi-nucleated phenotypes (black bar) (n = 2 independent experiments with a total of >250 cells per genotype analysed). (e) Genetic interactions between NUD1-FF and Swe1 depletion with deletions of the spindle positioning factors KAR9 and DYN1. Serial dilutions of stationary phase cultures of the indicated strains were spotted on YEPD plates supplemented with 500 μM IAA for SWE1-AID mutant cells and incubated at 30 °C (n = 2). (f) Protein-fragment complementation assay (PCA); Representative images of metaphase cells with the C-terminally tagged Swe1-VF1 and C-terminally Nud1-VF2 or N-terminally tagged VF2-Nud1 fragments. SPB-age marker is Spc42-mCherry and arrow marks the new SPB. Representative images of Nud1-sfGFP and Spc42-mCherry localization in metaphase. For all panels: All statistical significances were calculated using one-way ANOVA, ∗∗∗∗P < 0.0001, ∗∗∗P < 0.001, ∗P < 0.05, NS = non-significant. All error bars represent mean ± s.d. Scale bars, 2 μm.
Supplementary Figure 6 Swe1 is loaded onto the pre-existing SPB in G1 phase, that is, before full assembly of the new SPB.
(a) Categorization of the orientation of metaphase spindles with (i) pre-existing SPB proximal (ii) tilted or (iii) new SPB proximal (%) in wild type, Swe1-VF1 + VF2-Nud1 coexpressing and myo2-16 mutant (positive control) cells. Two-tailed Student t-test was performed to test significance, ∗∗P < 0.01, NS = non-significant. (n = 3 independent experiments with >120 cells analysed, mean ± s.d.) (b) Bleaching of SPBs in early G1 phase (left column) or metaphase (right column) and quantification of fluorescence intensity (%) of Swe1-VF1 + VF2-Nud1 during G1 phase for 50 min (SPB before separation and bud emerge, left column) and after SPB separation for 50 min (right column) (n = 10 cells pooled from two independent experiments, mean ± s.e.m.). Spc42-mCherry was used as an age marker. (c) Schematic overview of Swe1-dependent SPB specification and Swe1 during the cell cycle. (d) Representative image of Nud1-yeGFP during a FRAP experiment. Scale bars, 2 μm. (e) Quantification of fluorescence intensity (%) of Nud1-yeGFP of the bleached (orange; distal or proximal), unbleached (black) Nud1-yeGFP and of unbleached cells (dark red) (n = 30 cells pooled from three independent experiments, mean ± s.e.m.).
Supplementary Figure 7 SPIN recruits Bfa1 to pre-existing SPBs in metaphase.
(a,b) Quantification of fluorescence intensity (AU) of Bub2-yeGFP or Bfa1-yeGFP at the new and pre-existing SPB of correct oriented and inverted spindles in metaphase or anaphase (metaphase Bub2-yeGFP n = 65, anaphase Bfa1-yeGFP n = 60) and representative images. SPB age was determined by fluorescence asymmetry of Spc42-mCherry. (c) Quantification of fluorescence intensity (AU) of Bfa1-yeGFP at pre-existing or new SPB of wild type or mutant cells in the young and old SPB lineage of anaphase cells (%) (young SPB lineage, pre-existing SPB: WT n = 103, SWE1-AID n = 108; young SPB lineage, new SPB: WT n = 103, SWE1-AID n = 107; old SPB lineage, pre-existing SPB: WT n = 89, SWE1-AID n = 100; old SPB lineage, new SPB: WT n = 89, SWE1-AID n = 100) (d) Quantification of Bfa1-yeGFP and Spc42-mCherry intensities (AU) of the pre-existing SPBs in the young or old SPB lineage of wild type and mutant cells (WT young SPB lineage n = 98, old SPB lineage n = 98; SWE1-AID young SPB lineage n = 104, old SPB lineage n = 112). (e) Quantification of Bfa1-yeGFP and Spc42-mCherry intensities (AU) of the pre-existing SPBs of wild type and mutant cells (WT n = 168, SWE1-AID n = 148). For all panels: All statistical significances were calculated using two-tailed Student t-tests, ∗∗∗∗P < 0.0001, NS = non-significant.n represents number of cells pooled from three independent experiments. All error bars represent mean ± s.d. Scale bars 2 μm.
Supplementary Figure 8 Bfa1 and Bub2 direct Kar9 asymmetry towards the pre-existing SPB.
(a) Quantification of metaphase cells of indicated genotype with Kar9-YFP on the new SPB and anaphase cells with the new SPB segregated into the bud (%) (n = 3 independent experiments with a total of >120 cells per genotype analysed). All error bars represent mean ± s.d. Scale bars, 2 μm. SPB-age marker is Spc42-mCherry and arrow marks the new SPB.
Supplementary Figure 9 Uncropped scans of immunoblots.
Cropped images for figures are indicated by a red box.
Supplementary information
Supplementary Information
Supplementary Information (PDF 3976 kb)
Supplementary Information
Supplementary Information (PDF 68 kb)
Supplementary Table 1
Supplementary Information (XLSX 78 kb)
Supplementary Table 2
Supplementary Information (XLSX 26 kb)
Rights and permissions
About this article
Cite this article
Lengefeld, J., Hotz, M., Rollins, M. et al. Budding yeast Wee1 distinguishes spindle pole bodies to guide their pattern of age-dependent segregation. Nat Cell Biol 19, 941–951 (2017). https://doi.org/10.1038/ncb3576
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3576
This article is cited by
-
An updated view on the centrosome as a cell cycle regulator
Cell Division (2022)
-
Kar9 symmetry breaking alone is insufficient to ensure spindle alignment
Scientific Reports (2021)
-
Spindle pole power in health and disease
Current Genetics (2019)
-
Interrogation of γ-tubulin alleles using high-resolution fitness measurements reveals a distinct cytoplasmic function in spindle alignment
Scientific Reports (2017)