Abstract
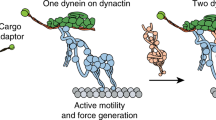
Kinesin and dynein motors transport intracellular cargos bidirectionally by pulling them in opposite directions along microtubules, through a process frequently described as a ‘tug of war’1. While kinesin produces 6 pN of force, mammalian dynein was found to be a surprisingly weak motor (0.5–1.5 pN) in vitro, suggesting that many dyneins are required to counteract the pull of a single kinesin2. Mammalian dynein’s association with dynactin and Bicaudal-D2 (BICD2) activates its processive motility3,4,5,6, but it was unknown how this affects dynein’s force output. Here, we show that formation of the dynein–dynactin–BICD2 (DDB) complex increases human dynein’s force production to 4.3 pN. An in vitro tug-of-war assay revealed that a single DDB successfully resists a single kinesin. Contrary to previous reports, the clustering of many dyneins is not required to win the tug of war. Our work reveals the key role of dynactin and a cargo adaptor protein in shifting the balance of forces between dynein and kinesin motors during intracellular transport.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hancock, W. O. Bidirectional cargo transport: moving beyond tug of war. Nat. Rev. Mol. Cell Biol. 15, 615–628 (2014).
Kunwar, A. et al. Mechanical stochastic tug-of-war models cannot explain bidirectional lipid-droplet transport. Proc. Natl Acad. Sci. USA 108, 18960–18965 (2011).
Schlager, M. A., Hoang, H. T., Urnavicius, L., Bullock, S. L. & Carter, A. P. In vitro reconstitution of a highly processive recombinant human dynein complex. EMBO J. 33, 1855–1868 (2014).
McKenney, R. J., Huynh, W., Tanenbaum, M. E., Bhabha, G. & Vale, R. D. Activation of cytoplasmic dynein motility by dynactin-cargo adapter complexes. Science 345, 337–341 (2014).
Urnavicius, L. et al. The structure of the dynactin complex and its interaction with dynein. Science 347, 1441–1446 (2015).
Splinter, D. et al. BICD2, dynactin, and LIS1 cooperate in regulating dynein recruitment to cellular structures. Mol. Biol. Cell 23, 4226–4241 (2012).
Roberts, A. J., Kon, T., Knight, P. J., Sutoh, K. & Burgess, S. A. Functions and mechanics of dynein motor proteins. Nat. Rev. Mol. Cell Biol. 14, 713–726 (2013).
Burgess, S. A., Walker, M. L., Sakakibara, H., Knight, P. J. & Oiwa, K. Dynein structure and power stroke. Nature 421, 715–718 (2003).
Torisawa, T. et al. Autoinhibition and cooperative activation mechanisms of cytoplasmic dynein. Nat. Cell Biol. 16, 1118–1124 (2014).
Moughamian, A. J., Osborn, G. E., Lazarus, J. E., Maday, S. & Holzbaur, E. L. F. Ordered recruitment of dynactin to the microtubule plus-end is required for efficient initiation of retrograde axonal transport. J. Neurosci. 33, 13190–13203 (2013).
Wang, Z. & Sheetz, M. P. One-dimensional diffusion on microtubules of particles coated with cytoplasmic dynein and immunoglobulins. Cell Struct. Funct. 24, 373–383 (1999).
Ayloo, S. et al. Dynactin functions as both a dynamic tether and brake during dynein-driven motility. Nat. Commun. 5, 4807 (2014).
McKenney, R. J., Vershinin, M., Kunwar, A., Vallee, R. B. & Gross, S. P. LIS1 and NudE induce a persistent dynein force-producing state. Cell 141, 304–314 (2010).
Rai, A. K., Ramaiya, A. J., Jha, R. & Mallik, R. Molecular adaptations allow dynein to generate large collective forces inside cells. Cell 152, 172–182 (2013).
Mallik, R., Carter, B. C., Lex, S. A., King, S. J. & Gross, S. P. Cytoplasmic dynein functions as a gear in response to load. Nature 427, 649–652 (2004).
Tripathy, S. K. et al. Autoregulatory mechanism for dynactin control of processive and diffusive dynein transport. Nat. Cell Biol. 16, 1192–1201 (2014).
Ori-McKenney, K. M., Xu, J., Gross, S. P. & Vallee, R. B. A cytoplasmic dynein tail mutation impairs motor processivity. Nat. Cell Biol. 12, 1228–1234 (2010).
Visscher, K., Schnitzer, M. J. & Block, S. M. Single kinesin molecules studied with a molecular force clamp. Nature 400, 184–189 (1999).
Rai, A. et al. Dynein clusters into lipid microdomains on phagosomes to drive rapid transport toward lysosomes. Cell 164, 722–734 (2016).
Hendricks, A. G. et al. Motor coordination via a tug-of-war mechanism drives bidirectional vesicle transport. Curr. Biol. 20, 697–702 (2010).
Thirumurugan, K., Sakamoto, T., Hammer, J. A. III, Sellers, J. R. & Knight, P. J. The cargo-binding domain regulates structure and activity of myosin 5. Nature 442, 212–215 (2006).
Kaan, H. Y. K., Hackney, D. D. & Kozielski, F. The structure of the kinesin-1 motor-tail complex reveals the mechanism of autoinhibition. Science 333, 883–885 (2011).
King, S. J. & Schroer, T. Dynactin increases the processivity of the cytoplasmic dynein motor. Nat. Cell Biol. 2, 20–24 (2000).
Chowdhury, S., Ketcham, S. A., Schroer, T. A. & Lander, G. C. Structural organization of the dynein–dynactin complex bound to microtubules. Nat. Struct. Mol. Biol. 22, 345–347 (2015).
Nicholas, M. P. et al. Control of cytoplasmic dynein force production and processivity by its C-terminal domain. Nat. Commun. 6, 6206 (2015).
DeWitt, M. A., Chang, A. Y., Combs, P. A. & Yildiz, A. Cytoplasmic dynein moves through uncoordinated stepping of the AAA + ring domains. Science 335, 221–225 (2012).
Cleary, F. B. et al. Tension on the linker gates the ATP-dependent release of dynein from microtubules. Nat. Commun. 5, 4587 (2014).
Belyy, V., Hendel, N. L., Chien, A. & Yildiz, A. Cytoplasmic dynein transports cargos via load-sharing between the heads. Nat. Commun. 5, 5544 (2014).
Gennerich, A., Carter, A. P., Reck-Peterson, S. L. & Vale, R. D. Force-induced bidirectional stepping of cytoplasmic dynein. Cell 131, 952–965 (2007).
Svoboda, K. & Block, S. M. Force and velocity measured for single kinesin molecules. Cell 77, 773–784 (1994).
Derr, N. D. et al. Tug-of-war in motor protein ensembles revealed with a programmable DNA origami scaffold. Science 338, 662–665 (2012).
Diehl, M. R., Zhang, K., Lee, H. J. & Tirrell, D. A. Engineering cooperativity in biomotor-protein assemblies. Science 311, 1468–1471 (2006).
Fu, M. M. & Holzbaur, E. L. F. JIP1 regulates the directionality of APP axonal transport by coordinating kinesin and dynein motors. J. Cell Biol. 202, 495–508 (2013).
Soppina, V., Rai, A. K., Ramaiya, A. J., Barak, P. & Mallik, R. Tug-of-war between dissimilar teams of microtubule motors regulates transport and fission of endosomes. Proc. Natl Acad. Sci. USA 106, 19381–19386 (2009).
Leidel, C., Longoria, R. a., Gutierrez, F. M. & Shubeita, G. T. Measuring molecular motor forces in vivo: implications for tug-of-war models of bidirectional transport. Biophys. J. 103, 492–500 (2012).
Shubeita, G. T. et al. Consequences of motor copy number on the intracellular transport of kinesin-1-driven lipid droplets. Cell 135, 1098–1107 (2008).
Fu, M. & Holzbaur, E. L. F. Integrated regulation of motor-driven organelle transport by scaffolding proteins. Trends Cell Biol. 24, 564–574 (2014).
Klumpp, S. & Lipowsky, R. Cooperative cargo transport by several molecular motors. Proc. Natl Acad. Sci. USA 102, 17284–17289 (2005).
Reck-Peterson, S. L. et al. Single-molecule analysis of dynein processivity and stepping behavior. Cell 126, 335–348 (2006).
Castoldi, M. & Popov, A. V. Purification of brain tubulin through two cycles of polymerization–depolymerization in a high-molarity buffer. Protein Expr. Purif. 32, 83–88 (2003).
Acknowledgements
We are grateful to S. Can, L. S. Ferro, A. Chien and J. Bandaria for helpful discussions. This work was supported by the NIH (GM094522 (A.Y.)), an NSF CAREER Award (MCB-1055017 (A.Y.)), the Medical Research Council, UK (MC_UP_A025_1011 (A.P.C.)), a Wellcome Trust New Investigator Award (WT100387) and an EMBO Young Investigator Award (both to A.P.C.), a Marie Curie Intra European Fellowship (M.A.S.), and an NSF Graduate Research Fellowship (DGE 1106400 (V.B.)).
Author information
Authors and Affiliations
Contributions
V.B., A.P.C. and A.Y. designed the study. M.A.S. and H.F. prepared the constructs and purified the protein. V.B. and A.E.R. performed the optical trapping experiments. V.B. labelled dynein and kinesin with DNA, performed the fluorescence motility experiments, and analysed the data. V.B., A.P.C. and A.Y. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Temporally color-coded projections of free dyneins and DDB complexes on microtubules.
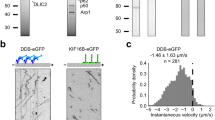
(a) (Top) Interactions of single free dyneins with microtubules are visualized by color-coding a 2D projection of Supplementary Video 1. Scale bar: 5 μm. (Bottom) Fluorescence kymograph along one of the three visible axonemes in Supplementary Video 1. Motile dyneins walked at 79 ± 11 nm s−1 (s.e.m.,n = 15). At 1 nM motor, the density of fluorescent spots on the axoneme was 0.19 per micron length of axoneme per minute, n = 93 (b). (Top) Processive motility of DDB complexes on microtubules is visualized by color-coding a 2D projection of Supplementary Video 2. (Bottom) Fluorescence kymograph along the axoneme in Supplementary Video 3. At 1 nM motor, the density of fluorescent spots on the axoneme was 0.66 per micron length of axoneme per minute, n = 98.
Supplementary Figure 2 Individual steps in trapping traces and Gaussian mixture model fitting of stall distributions.
The kinesin (a) and dynein (b) stall traces are zoomed-in looks on the traces shown in Fig. 2e and Fig. 2c, respectively. Red arrowheads represent the detachment of the motor from the microtubule after the stall. (c,d) Comparison of 2-component and 3-component Gaussian mixture model fits to the DDB stall force distribution shown in Fig. 2e. The fitting is carried out on the underlying distribution and is thus independent of the binning of the histogram. Blue curves are the scaled results of 2-component (c) and 3-component (d) Gaussian fits. Numbers represent center values ± standard errors of each peak. The Bayes Information Criterion (BIC) is lower for the 2-Gaussian fit, indicating that adding a third Gaussian peak is not statistically warranted.
Supplementary Figure 3 Stall force measurements of full-length yeast cytoplasmic dynein.
(a) Full-length yeast cytoplasmic dynein with an N-terminal GFP (GFP-Dyn471kD-DHA) is attached to an optically trapped bead via anti-GFP antibody. (b) Trace showing a typical stall of a bead driven by full-length yeast dynein in a fixed trap assay. Red arrowhead represents the detachment of the motor from the microtubule after the stall. (c) The histogram of observed stalls reveals the mean stall force (mean ± s.e.m.,n = 21 stalls from 5 beads).
Supplementary Figure 4 Examples of dynein-kinesin colocalizers in the presence of dynactin and BICD2N.
Dynein is tagged with Alexa647 (red) and kinesin is tagged with TMR (cyan). The red and cyan channels are laterally offset by five pixels to enhance the visibility of the colocalizers. Individual channels are shown in grayscale. All microtubules are arranged with the plus ends facing towards the right. Scale bars are identical in all images.
Supplementary Figure 5 Full images of all gels shown in main figures.
(a) Denaturing SDS–PAGE gel of purified dynein fractions, stained with Instant Blue. Bands corresponding to all dynein subunits can be observed. (b) Denaturing SDS–PAGE gel of purified BICD2N, stained with Instant Blue. (c) Denaturing SDS-PAGE gel of kinesin labeled with DNA. Excess DNA is removed from labeled kinesin by microtubule bind and release (MTBR). The MTBR fractions shown in lanes 10 through 14 are Pre (kinesin pre-mixed with microtubules), SN1 (supernatant from the first spin), P1 (pellet from the first spin), SN2 (supernatant from the second spin), and P2 (pellet from the second spin). (d,e) Denaturing SDS-PAGE gel of dynein labeled with DNA and Alexa647. Dynein was first labeled with ssDNA-BG-GLA-NHS at the indicated molar ratios, then labeled with an excess of Alexa647-BG-GLA-NHS. (d) Silver stain reveals the total amount of dynein, regardless of what fraction of it is labeled with ssDNA. (e) When imaged specifically for Alexa647 fluorescence, the intensities of individual bands on the same gel reveal the fraction of dyneins whose SNAPf sites are occupied by ssDNA and therefore not accessible to labeling by Alexa647-BG-GLA-NHS.
Supplementary information
Supplementary Information
Supplementary Information (PDF 880 kb)
Human dynein exhibits minimal motility in the absence of tail-binding proteins.
GFP-labeled recombinant dynein motors interacting with surface-immobilized axonemes in the presence of 2 mM ATP. Scale bar: 5 μm. (AVI 8996 kb)
Addition of dynactin and BICD2N to dynein leads to fast processive movement.
GFP-labeled dynein motors walking on surface-immobilized axonemes in the presence of a 5× molar excess of dynactin, 2× molar excess of BICD2N, and 2 mM ATP. Scale bar: 5 μm. (AVI 8996 kb)
Fast motility is activated by the binding of large cargos to dynein’s tail.
Bright-field recording of dynein-driven motility of a 200 nm polystyrene bead on a surface-immobilized axoneme in the presence of 2 mM ATP. The position of the motile bead is marked with a yellow arrow for clarity. Scale bar: 5 μm. (AVI 8996 kb)
Motility of a dynein-kinesin colocalizer in the absence of dynactin and BICD2N.
Two-color TIRF imaging of kinesin-TMR (shown in cyan) and dynein-Alexa647 (shown in red) on a microtubule. Colocalizers are marked with white arrows. (AVI 2354 kb)
Motility of a dynein-kinesin colocalizer in the presence of dynactin and BICD2N.
Two-color TIRF imaging of kinesin-TMR (shown in cyan) and dynein-Alexa647 (shown in red) on a microtubule. The colocalizer is marked with white arrows. (AVI 769 kb)
Rights and permissions
About this article
Cite this article
Belyy, V., Schlager, M., Foster, H. et al. The mammalian dynein–dynactin complex is a strong opponent to kinesin in a tug-of-war competition. Nat Cell Biol 18, 1018–1024 (2016). https://doi.org/10.1038/ncb3393
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3393
This article is cited by
-
Lis1 slows force-induced detachment of cytoplasmic dynein from microtubules
Nature Chemical Biology (2024)
-
Microtubule acetylation dyshomeostasis in Parkinson’s disease
Translational Neurodegeneration (2023)
-
TRAK adaptors regulate the recruitment and activation of dynein and kinesin in mitochondrial transport
Nature Communications (2023)
-
CENP-E activation by Aurora A and B controls kinetochore fibrous corona disassembly
Nature Communications (2023)
-
The regulatory function of the AAA4 ATPase domain of cytoplasmic dynein
Nature Communications (2020)