Abstract
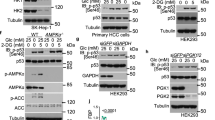
Cancer cells consume large quantities of glucose and primarily use glycolysis for ATP production, even in the presence of adequate oxygen1,2. This metabolic signature (aerobic glycolysis or the Warburg effect) enables cancer cells to direct glucose to biosynthesis, supporting their rapid growth and proliferation3,4. However, both causes of the Warburg effect and its connection to biosynthesis are not well understood. Here we show that the tumour suppressor p53, the most frequently mutated gene in human tumours, inhibits the pentose phosphate pathway5 (PPP). Through the PPP, p53 suppresses glucose consumption, NADPH production and biosynthesis. The p53 protein binds to glucose-6-phosphate dehydrogenase (G6PD), the first and rate-limiting enzyme of the PPP, and prevents the formation of the active dimer. Tumour-associated p53 mutants lack the G6PD-inhibitory activity. Therefore, enhanced PPP glucose flux due to p53 inactivation may increase glucose consumption and direct glucose towards biosynthesis in tumour cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Warburg, O., Posener, K. & Negelein, E. Ueber den Stoffwechsel der Tumoren. Biochem. Z. 152, 319–344 (1924).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
DeBerardinis, R. J., Lum, J. J., Hatzivassiliou, G. & Thompson, C. B. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 7, 11–20 (2008).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Berg, J. M., Tymoczko, J. L. & Stryer, L. Biochemistry 6th edn 577–589 (W. H. Freeman, 2006).
Vogelstein, B., Lane, D. & Levine, A. J. Surfing the p53 network. Nature 408, 307–310 (2000).
Vousden, K. H. & Prives, C. Blinded by the light: the growing complexity of p53. Cell 137, 413–431 (2009).
Kondoh, H. et al. Glycolytic enzymes can modulate cellular life span. Cancer Res. 65, 177–185 (2005).
Matoba, S. et al. p53 regulates mitochondrial respiration. Science 312, 1650–1653 (2006).
Vousden, K. H. & Ryan, K. M. p53 and metabolism. Nat. Rev. Cancer 9, 691–700 (2009).
Bunz, F. et al. Requirement for p53 and p21 to sustain G2 arrest after DNA damage. Science 282, 1497–1501 (1998).
Tseng, Y. H. et al. New role of bone morphogenetic protein 7 in brown adipogenesis and energy expenditure. Nature 454, 1000–1004 (2008).
Komarov, P. G. et al. A chemical inhibitor of p53 that protects mice from the side effects of cancer therapy. Science 285, 1733–1737 (1999).
Inga, A. & Resnick, M. A. Novel human p53 mutations that are toxic to yeast can enhance transactivation of specific promoters and reactivate tumor p53 mutants. Oncogene 20, 3409–3419 (2001).
Freedman, D. A. & Levine, A. J. Nuclear export is required for degradation of endogenous p53 by MDM2 and human papillomavirus E6. Mol. Cell Biol. 18, 7288–7293 (1998).
Stommel, J. M. et al. A leucine-rich nuclear export signal in the p53 tetramerization domain: Regulation of subcellular localization and p53 activity by NES masking. EMBO J. 18, 1660–1672 (1999).
Yan, W. & Chen, X. Characterization of functional domains necessary for mutant p53 gain of function. J. Biol. Chem. 285, 14229–14238 (2010).
Nikolova, P. V., Henckel, J., Lane, D. P. & Fersht, A. R. Semirational design of active tumor suppressor p53 DNA binding domain with enhanced stability. Proc. Natl Acad. Sci. USA 95, 14675–14680 (1998).
Au, S. W., Gover, S., Lam, V. M. & Adams, M. J. Human glucose-6-phosphate dehydrogenase: The crystal structure reveals a structural NADP(+) molecule and provides insights into enzyme deficiency. Structure 8, 293–303 (2000).
Roos, D. et al. Molecular basis and enzymatic properties of glucose 6-phosphate dehydrogenase volendam, leading to chronic nonspherocytic anemia, granulocyte dysfunction, and increased susceptibility to infections. Blood 94, 2955–2962 (1999).
Green, D. R. & Kroemer, G. Cytoplasmic functions of the tumour suppressor p53. Nature 458, 1127–1130 (2009).
Bensaad, K. et al. TIGAR, a p53-inducible regulator of glycolysis and apoptosis. Cell 126, 107–120 (2006).
Tang, J. et al. Critical role for Daxx in regulating Mdm2. Nat. Cell Biol. 8, 855–862 (2006).
Mancuso, A., Sharfstein, S. T., Tucker, S. N., Clark, D. S. & Blanch, H. W. Examination of primary metabolic pathways in a murine hybridoma with carbon-13 nuclear magnetic resonance spectroscopy. Biotechnol. Bioeng. 44, 563–585 (1994).
Brummelkamp, T. R., Bernards, R. & Agami, R. A system for stableexpression of short interfering RNAs in mammalian cells. Science 296, 550–553 (2002).
Tian, W. N. et al. Importance of glucose-6-phosphate dehydrogenase activity for cell growth. J. Biol. Chem. 273, 10609–10617 (1998).
Adorno, M. et al. A mutant-p53/Smad complex opposes p63 to empower TGF β-induced metastasis. Cell 137, 87–98 (2009).
Du, W. et al. Suppression of p53 activity by Siva1. Cell Death Differ. 16, 1493–1504 (2009).
Tang, J. et al. A novel transcription regulatory complex containing death domain-associated protein and the ATR-X syndrome protein. J. Biol. Chem. 279, 20369–20377 (2004).
Zhang, Z., Yu, J. & Stanton, R. C. A method for determination of pyridine nucleotides using a single extract. Anal. Biochem. 285, 163–167 (2000).
Acknowledgements
We thank W. Xie for isolating p53+/+ and p53−/− MEF cells; X. Chen for SW480 cells; B. Vogelstein and W. El-Deiry for HCT116 cells; J. Cross, N. Li, J. Wu, Y. Mei, A. Stonestrom, W. Tan, H. Liu, Y. Hao, X. Zhao and Z. Lou for technical assistance; C. B. Thompson and J. Delikatny for helpful comments; and A. Stonestrom and E. Thompson for help with manuscript preparation. Supported by grants from the China National Natural Science Foundation (31030046), the Ministry of Science and Technology (2010CB912804 and 2011CB966302) and the Chinese Academy of Sciences (KSCX1-YW-R-57) to M.W. and the US National Institutes of Health (CA088868 and GM060911) and the Department of Defense (W81XWH-07-1-0336 and W81XWH-10-1-0468) to X.Y.
Author information
Authors and Affiliations
Contributions
P.J., W.D., M.W. and X.Y. designed the experiments and interpreted results. P.J. and W.D. carried out all the experiments, except those mentioned below. X.W. carried out the experiments on G6PD activity in yeast, the surface plasmon resonance, and lipid droplets in mouse liver. A.M. and P.J. analysed the oxidative PPP flux. X.G. supplied the p53 wild-type and knockout mice. X.Y. wrote the manuscript with the help of P.J. and W.D.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 805 kb)
Rights and permissions
About this article
Cite this article
Jiang, P., Du, W., Wang, X. et al. p53 regulates biosynthesis through direct inactivation of glucose-6-phosphate dehydrogenase. Nat Cell Biol 13, 310–316 (2011). https://doi.org/10.1038/ncb2172
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2172
This article is cited by
-
G6PD maintains the VSMC synthetic phenotype and accelerates vascular neointimal hyperplasia by inhibiting the VDAC1–Bax-mediated mitochondrial apoptosis pathway
Cellular & Molecular Biology Letters (2024)
-
Cyclosporine A alleviates colitis by inhibiting the formation of neutrophil extracellular traps via the regulating pentose phosphate pathway
Molecular Medicine (2023)
-
Metabolic reprogramming in colorectal cancer: regulatory networks and therapy
Cell & Bioscience (2023)
-
Lipid metabolic reprogramming in tumor microenvironment: from mechanisms to therapeutics
Journal of Hematology & Oncology (2023)
-
Cuproptosis: p53-regulated metabolic cell death?
Cell Death & Differentiation (2023)