Abstract
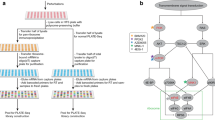
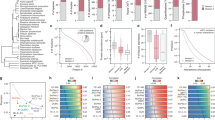
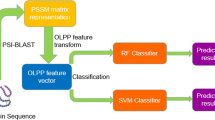
Identifying physical interactions between proteins and other molecules is a critical aspect of biological analysis. Here we describe PLATO, an in vitro method for mapping such interactions by affinity enrichment of a library of full-length open reading frames displayed on ribosomes, followed by massively parallel analysis using DNA sequencing. We demonstrate the broad utility of the method for human proteins by identifying known and previously unidentified interacting partners of LYN kinase, patient autoantibodies, and the small-molecules gefitinib and dasatinib.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Faix, P.H. et al. Biotechniques 36, 1018–1022, 1024, 1026–1029 (2004).
Zhu, H., Bilgin, M. & Snyder, M. Annu. Rev. Biochem. 72, 783–812 (2003).
Jeong, J.S. et al. Mol. Cell Proteomics 11, O111.016253 (2012).
Lamesch, P. et al. Genomics 89, 307–315 (2007).
Amstutz, P., Binz, H.K., Zahnd, C. & Plückthun,, A. Ribosome display: in vitro selection of protein-protein interactions in Cell Biology—A Laboratory Handbook (ed. Celis, J.) Vol. 1, 3rd Ed., 497–509 (Elsevier Academic Press, 2006).
Boggon, T.J. & Eck, M.J. Oncogene 23, 7918–7927 (2004).
Weng, Z. et al. Mol. Cell Biol. 14, 4509–4521 (1994).
Samuels, A.L., Klinken, S.P. & Ingley, E. Blood 113, 3845–3856 (2009).
Larman, H.B. et al. Nat. Biotechnol. 29, 535–541 (2011).
Carthagena, L. et al. PLoS ONE 4, e4894 (2009).
Brehmer, D. et al. Cancer Res. 65, 379–382 (2005).
Acknowledgements
We would like to thank K. Waraska, M. Cicero, S. Alian and A. Gagne for assistance with Illumina sequencing, and J. Laserson for statistical advice. Thanks to D. Zhu for help with synthesis of biotin-dasatinib, which was partially supported by US National Institutes of Health U54 CA156734 to the University of Massachusetts Boston–Dana-Farber Harvard Cancer Center U54 Comprehensive Partnership (Project 3, Co-PIs: N.S. Gray, W. Zhang, and P.L. Yang). We also thank N. Gray at Harvard Medical School for valuable advice, and H. Varmus at the National Cancer Institute for providing gefitinib reagents and advice. This work was supported in part by NIH grant 3P30CA023100-25S8 to S.K. S.J.E. is an investigator with the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
S.J.E. and H.B.L. conceived and supervised the project. pRD human ORFeome library was constructed by J.Z., and characterized by J.Z. and H.B.L. The PLATO protocol was developed by H.B.L. and J.Z. Clinical evaluations and patient sample acquisitions were performed by S.K. Statistical analysis was performed by U.L. under the supervision of G.C. R.S. provided gefitinib-conjugated beads. PLATO candidates were confirmed by J.Z. and G.G. A.C. provided support for the validation of LYN binding candidates. N.P. provided support for the validation of PND autoantigen candidates. Z.Z. and W.Z. provided biotin-dasatinib. The manuscript was prepared by H.B.L. and J.Z., and edited by S.J.E.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–6 and Supplementary Methods (PDF 3363 kb)
Supplementary Tables 1-4
ORFeome enrichment by GST-LYN, ORFeome enrichment by PND patient autoantibodies, ORFeome enrichment by dasatinib, Sequences of DNA primers used in this study (XLSX 101 kb)
Rights and permissions
About this article
Cite this article
Zhu, J., Larman, H., Gao, G. et al. Protein interaction discovery using parallel analysis of translated ORFs (PLATO). Nat Biotechnol 31, 331–334 (2013). https://doi.org/10.1038/nbt.2539
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2539
This article is cited by
-
HSP90 inhibitor NVP-BEP800 affects stability of SRC kinases and growth of T-cell and B-cell acute lymphoblastic leukemias
Blood Cancer Journal (2021)
-
PhIP-Seq characterization of serum antibodies using oligonucleotide-encoded peptidomes
Nature Protocols (2018)
-
High throughput protease profiling comprehensively defines active site specificity for thrombin and ADAMTS13
Scientific Reports (2018)
-
Association study between copy number variation and beef fatty acid profile of Nellore cattle
Journal of Applied Genetics (2018)
-
Genetic engineering: Lassoing genomic libraries
Nature Biomedical Engineering (2017)