Abstract
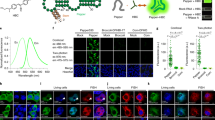
The functional state of a cell is largely determined by the spatiotemporal organization of its proteome. Technologies exist for measuring particular aspects of protein turnover and localization, but comprehensive analysis of protein dynamics across different scales is possible only by combining several methods. Here we describe tandem fluorescent protein timers (tFTs), fusions of two single-color fluorescent proteins that mature with different kinetics, which we use to analyze protein turnover and mobility in living cells. We fuse tFTs to proteins in yeast to study the longevity, segregation and inheritance of cellular components and the mobility of proteins between subcellular compartments; to measure protein degradation kinetics without the need for time-course measurements; and to conduct high-throughput screens for regulators of protein turnover. Our experiments reveal the stable nature and asymmetric inheritance of nuclear pore complexes and identify regulators of N-end rule–mediated protein degradation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schoenheimer, R. The Dynamic State of Body Constituents (Harvard University Press, Cambridge, MA, 1942).
Nakayama, K.I. & Nakayama, K. Ubiquitin ligases: cell-cycle control and cancer. Nat. Rev. Cancer 6, 369–381 (2006).
Johnston, J.A., Ward, C.L. & Kopito, R.R. Aggresomes: a cellular response to misfolded proteins. J. Cell Biol. 143, 1883–1898 (1998).
Morimoto, R.I. Proteotoxic stress and inducible chaperone networks in neurodegenerative disease and aging. Genes Dev. 22, 1427–1438 (2008).
Balch, W.E., Morimoto, R.I., Dillin, A. & Kelly, J.W. Adapting proteostasis for disease intervention. Science 319, 916–919 (2008).
Pratt, J.M. et al. Dynamics of protein turnover, a missing dimension in proteomics. Mol. Cell. Proteomics 1, 579–591 (2002).
Eden, E. et al. Proteome half-life dynamics in living human cells. Science 331, 764–768 (2011).
Zhang, L. et al. Method for real-time monitoring of protein degradation at the single cell level. Biotechniques 42, 446–450 (2007).
Yen, H.-C.S., Xu, Q., Chou, D.M., Zhao, Z. & Elledge, S.J. Global protein stability profiling in mammalian cells. Science 322, 918–923 (2008).
Gordon, A. et al. Single-cell quantification of molecules and rates using open-source microscope-based cytometry. Nat. Methods 4, 175–181 (2007).
Yen, H.-C.S. & Elledge, S.J. Identification of SCF ubiquitin ligase substrates by global protein stability profiling. Science 322, 923–929 (2008).
Newman, R.H., Fosbrink, M.D. & Zhang, J. Genetically encodable fluorescent biosensors for tracking signaling dynamics in living cells. Chem. Rev. 111, 3614–3666 (2011).
Wu, B., Piatkevich, K.D., Lionnet, T., Singer, R.H. & Verkhusha, V.V. Modern fluorescent proteins and imaging technologies to study gene expression, nuclear localization, and dynamics. Curr. Opin. Cell Biol. 23, 310–317 (2011).
Terskikh, A. et al. “Fluorescent timer”: protein that changes color with time. Science 290, 1585–1588 (2000).
Subach, F.V. et al. Monomeric fluorescent timers that change color from blue to red report on cellular trafficking. Nat. Chem. Biol. 5, 118–126 (2009).
Tsuboi, T., Kitaguchi, T., Karasawa, S., Fukuda, M. & Miyawaki, A. Age-dependent preferential dense-core vesicle exocytosis in neuroendocrine cells revealed by newly developed monomeric fluorescent timer protein. Mol. Biol. Cell 21, 87–94 (2010).
Merzlyak, E.M. et al. Bright monomeric red fluorescent protein with an extended fluorescence lifetime. Nat. Methods 4, 555–557 (2007).
Pédelacq, J.-D., Cabantous, S., Tran, T., Terwilliger, T.C. & Waldo, G.S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Pereira, G., Tanaka, T.U., Nasmyth, K. & Schiebel, E. Modes of spindle pole body inheritance and segregation of the Bfa1p-Bub2p checkpoint protein complex. EMBO J. 20, 6359–6370 (2001).
Malínská, K., Malínský, J., Opekarová, M. & Tanner, W. Visualization of protein compartmentation within the plasma membrane of living yeast cells. Mol. Biol. Cell 14, 4427–4436 (2003).
Takizawa, P.A., DeRisi, J.L., Wilhelm, J.E. & Vale, R.D. Plasma membrane compartmentalization in yeast by messenger RNA transport and a septin diffusion barrier. Science 290, 341–344 (2000).
Chen, T. et al. Multigenerational cortical inheritance of the Rax2 protein in orienting polarity and division in yeast. Science 290, 1975–1978 (2000).
Strambio-De-Castillia, C., Niepel, M. & Rout, M.P. The nuclear pore complex: bridging nuclear transport and gene regulation. Nat. Rev. Mol. Cell Biol. 11, 490–501 (2010).
D'Angelo, M.A., Raices, M., Panowski, S.H. & Hetzer, M.W. Age-dependent deterioration of nuclear pore complexes causes a loss of nuclear integrity in postmitotic cells. Cell 136, 284–295 (2009).
Zabel, U. et al. Nic96p is required for nuclear pore formation and functionally interacts with a novel nucleoporin, Nup188p. J. Cell Biol. 133, 1141–1152 (1996).
Makio, T. et al. The nucleoporins Nup170p and Nup157p are essential for nuclear pore complex assembly. J. Cell Biol. 185, 459–473 (2009).
Steinkraus, K.A., Kaeberlein, M. & Kennedy, B.K. Replicative aging in yeast: the means to the end. Annu. Rev. Cell Dev. Biol. 24, 29–54 (2008).
Winey, M., Yarar, D., Giddings, T.H. & Mastronarde, D.N. Nuclear pore complex number and distribution throughout the Saccharomyces cerevisiae cell cycle by three-dimensional reconstruction from electron micrographs of nuclear envelopes. Mol. Biol. Cell 8, 2119–2132 (1997).
Khmelinskii, A., Keller, P.J., Lorenz, H., Schiebel, E. & Knop, M. Segregation of yeast nuclear pores. Nature 466, E1 (2010).
Campbell, R.E. et al. A monomeric red fluorescent protein. Proc. Natl. Acad. Sci. USA 99, 7877–7882 (2002).
Verzijlbergen, K.F. et al. Recombination-induced tag exchange to track old and new proteins. Proc. Natl. Acad. Sci. USA 107, 64–68 (2010).
Ouellet, J. & Barral, Y. Organelle segregation during mitosis: Lessons from asymmetrically dividing cells. J. Cell Biol. 196, 305–313 (2012).
Varshavsky, A. The N-end rule pathway and regulation by proteolysis. Protein Sci. 20, 1298–1345 (2011).
Belle, A., Tanay, A., Bitincka, L., Shamir, R. & O'Shea, E.K. Quantification of protein half-lives in the budding yeast proteome. Proc. Natl. Acad. Sci. USA 103, 13004–13009 (2006).
Winzeler, E.A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Tong, A.H.Y. & Boone, C. High-throughput strain construction and systematic synthetic lethal screening in Saccharomyces cerevisiae. Methods Microbiol. 36, 369–386 (2007).
Hwang, C.-S., Shemorry, A. & Varshavsky, A. N-terminal acetylation of cellular proteins creates specific degradation signals. Science 327, 973–977 (2010).
Swanson, R., Locher, M. & Hochstrasser, M. A conserved ubiquitin ligase of the nuclear envelope/endoplasmic reticulum that functions in both ER-associated and Matalpha2 repressor degradation. Genes Dev. 15, 2660–2674 (2001).
Leggett, D.S. et al. Multiple associated proteins regulate proteasome structure and function. Mol. Cell 10, 495–507 (2002).
Hwang, C.-S., Shemorry, A., Auerbach, D. & Varshavsky, A. The N-end rule pathway is mediated by a complex of the RING-type Ubr1 and HECT-type Ufd4 ubiquitin ligases. Nat. Cell Biol. 12, 1177–1185 (2010).
Shaner, N.C., Patterson, G.H. & Davidson, M.W. Advances in fluorescent protein technology. J. Cell Sci. 120, 4247–4260 (2007).
Hardy, S., Legagneux, V., Audic, Y. & Paillard, L. Reverse genetics in eukaryotes. Biol. Cell 102, 561–580 (2010).
Skarnes, W.C. et al. A conditional knockout resource for the genome-wide study of mouse gene function. Nature 474, 337–342 (2011).
Bogdanove, A.J. & Voytas, D.F. TAL effectors: customizable proteins for DNA targeting. Science 333, 1843–1846 (2011).
Yewdell, J.W., Lacsina, J.R., Rechsteiner, M.C. & Nicchitta, C.V. Out with the old, in with the new? Comparing methods for measuring protein degradation. Cell Biol. Int. 35, 457–462 (2011).
Alber, F. et al. The molecular architecture of the nuclear pore complex. Nature 450, 695–701 (2007).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Shaner, N.C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Zacharias, D.A., Violin, J.D., Newton, A.C. & Tsien, R.Y. Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 296, 913–916 (2002).
Bachmair, A., Finley, D. & Varshavsky, A. In vivo half-life of a protein is a function of its amino-terminal residue. Science 234, 179–186 (1986).
Pau, G., Fuchs, F., Sklyar, O., Boutros, M. & Huber, W. EBImage–an R package for image processing with applications to cellular phenotypes. Bioinformatics 26, 979–981 (2010).
Boutros, M., Brás, L.P. & Huber, W. Analysis of cell-based RNAi screens. Genome Biol. 7, R66 (2006).
Malo, N., Hanley, J.A., Cerquozzi, S., Pelletier, J. & Nadon, R. Statistical practice in high-throughput screening data analysis. Nat. Biotechnol. 24, 167–175 (2006).
Acknowledgements
We are grateful to Y. Belyaev, S. Terjung and the Advanced Light Microscopy Facility of EMBL for support with microscopy; A. Riddell, A.P. Gonzalez and the FACS Facility of EMBL for flow cytometry analyses; the CellNetworks cluster for funding to M.K., the German Research Foundation for funding to E.S. (SFB638), the Howard Hughes Medical Institute for funding to P.J.K., the European Molecular Biology Organization for funding to A.Kh. (EMBO ALTF 1124-2010), the EU-FP7 Network of Excellence in Systems Microscopy for funding to J.D.B. and W.H., and the Novartis Stiftung for funding to A.Ka. We thank P.I. Bastiaens, D. Gilmour and A. Kinkhabwala for discussions.
Author information
Authors and Affiliations
Contributions
M.K. conceived and, together with E.S., A.Kh. and P.J.K., designed the project. A.Kh., M.M., A.B., A.Ka., S.T. and P.J.K. did all yeast experiments. B.R.M. did mammalian cell work. G.P. contributed reagents. P.J.K., J.D.B., A.Kh., W.H., M.W. and M.K. developed theory. P.J.K., J.D.B. and A.Kh. developed analytical tools. M.K., A.Kh. and P.J.K. wrote the manuscript with input from E.S. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Notes 1–6, Supplementary Theory, Supplementary Tables 1,2 and Supplementary Figures 1–16 (PDF 2971 kb)
Supplementary Table 3
Results of the screens for components of the N-end rule pathway (XLS 866 kb)
Supplementary Movie 1
Movie corresponding to Figure 3d (i, ii) (MOV 556 kb)
Supplementary Movie 2
Movie corresponding to Figure 3d (iii) (MOV 1374 kb)
Rights and permissions
About this article
Cite this article
Khmelinskii, A., Keller, P., Bartosik, A. et al. Tandem fluorescent protein timers for in vivo analysis of protein dynamics. Nat Biotechnol 30, 708–714 (2012). https://doi.org/10.1038/nbt.2281
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2281
This article is cited by
-
Quantifying label enrichment from two mass isotopomers increases proteome coverage for in vivo protein turnover using heavy water metabolic labeling
Communications Chemistry (2023)
-
Orphan quality control by an SCF ubiquitin ligase directed to pervasive C-degrons
Nature Communications (2023)
-
Temporal segregation of biosynthetic processes is responsible for metabolic oscillations during the budding yeast cell cycle
Nature Metabolism (2023)
-
Learning perturbation-inducible cell states from observability analysis of transcriptome dynamics
Nature Communications (2023)
-
Morphogen gradient scaling by recycling of intracellular Dpp
Nature (2022)