Abstract
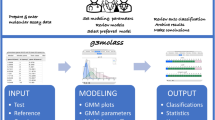
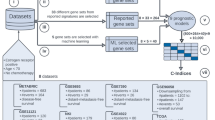
Gene expression data from microarrays are being applied to predict preclinical and clinical endpoints, but the reliability of these predictions has not been established. In the MAQC-II project, 36 independent teams analyzed six microarray data sets to generate predictive models for classifying a sample with respect to one of 13 endpoints indicative of lung or liver toxicity in rodents, or of breast cancer, multiple myeloma or neuroblastoma in humans. In total, >30,000 models were built using many combinations of analytical methods. The teams generated predictive models without knowing the biological meaning of some of the endpoints and, to mimic clinical reality, tested the models on data that had not been used for training. We found that model performance depended largely on the endpoint and team proficiency and that different approaches generated models of similar performance. The conclusions and recommendations from MAQC-II should be useful for regulatory agencies, study committees and independent investigators that evaluate methods for global gene expression analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Marshall, E. Getting the noise out of gene arrays. Science 306, 630–631 (2004).
Frantz, S. An array of problems. Nat. Rev. Drug Discov. 4, 362–363 (2005).
Michiels, S., Koscielny, S. & Hill, C. Prediction of cancer outcome with microarrays: a multiple random validation strategy. Lancet 365, 488–492 (2005).
Ntzani, E.E. & Ioannidis, J.P. Predictive ability of DNA microarrays for cancer outcomes and correlates: an empirical assessment. Lancet 362, 1439–1444 (2003).
Ioannidis, J.P. Microarrays and molecular research: noise discovery? Lancet 365, 454–455 (2005).
Ein-Dor, L., Kela, I., Getz, G., Givol, D. & Domany, E. Outcome signature genes in breast cancer: is there a unique set? Bioinformatics 21, 171–178 (2005).
Ein-Dor, L., Zuk, O. & Domany, E. Thousands of samples are needed to generate a robust gene list for predicting outcome in cancer. Proc. Natl. Acad. Sci. USA 103, 5923–5928 (2006).
Shi, L. et al. QA/QC: challenges and pitfalls facing the microarray community and regulatory agencies. Expert Rev. Mol. Diagn. 4, 761–777 (2004).
Shi, L. et al. Cross-platform comparability of microarray technology: intra-platform consistency and appropriate data analysis procedures are essential. BMC Bioinformatics 6 Suppl 2, S12 (2005).
Shi, L. et al. The MicroArray Quality Control (MAQC) project shows inter- and intraplatform reproducibility of gene expression measurements. Nat. Biotechnol. 24, 1151–1161 (2006).
Guo, L. et al. Rat toxicogenomic study reveals analytical consistency across microarray platforms. Nat. Biotechnol. 24, 1162–1169 (2006).
Canales, R.D. et al. Evaluation of DNA microarray results with quantitative gene expression platforms. Nat. Biotechnol. 24, 1115–1122 (2006).
Patterson, T.A. et al. Performance comparison of one-color and two-color platforms within the MicroArray Quality Control (MAQC) project. Nat. Biotechnol. 24, 1140–1150 (2006).
Shippy, R. et al. Using RNA sample titrations to assess microarray platform performance and normalization techniques. Nat. Biotechnol. 24, 1123–1131 (2006).
Tong, W. et al. Evaluation of external RNA controls for the assessment of microarray performance. Nat. Biotechnol. 24, 1132–1139 (2006).
Irizarry, R.A. et al. Multiple-laboratory comparison of microarray platforms. Nat. Methods 2, 345–350 (2005).
Strauss, E. Arrays of hope. Cell 127, 657–659 (2006).
Shi, L., Perkins, R.G., Fang, H. & Tong, W. Reproducible and reliable microarray results through quality control: good laboratory proficiency and appropriate data analysis practices are essential. Curr. Opin. Biotechnol. 19, 10–18 (2008).
Dudoit, S., Fridlyand, J. & Speed, T.P. Comparison of discrimination methods for the classification of tumors using gene expression data. J. Am. Stat. Assoc. 97, 77–87 (2002).
Goodsaid, F.M. et al. Voluntary exploratory data submissions to the US FDA and the EMA: experience and impact. Nat. Rev. Drug Discov. 9, 435–445 (2010).
van 't Veer, L.J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002).
Buyse, M. et al. Validation and clinical utility of a 70-gene prognostic signature for women with node-negative breast cancer. J. Natl. Cancer Inst. 98, 1183–1192 (2006).
Dumur, C.I. et al. Interlaboratory performance of a microarray-based gene expression test to determine tissue of origin in poorly differentiated and undifferentiated cancers. J. Mol. Diagn. 10, 67–77 (2008).
Deng, M.C. et al. Noninvasive discrimination of rejection in cardiac allograft recipients using gene expression profiling. Am. J. Transplant. 6, 150–160 (2006).
Coombes, K.R., Wang, J. & Baggerly, K.A. Microarrays: retracing steps. Nat. Med. 13, 1276–1277, author reply 1277–1278 (2007).
Ioannidis, J.P.A. et al. Repeatability of published microarray gene expression analyses. Nat. Genet. 41, 149–155 (2009).
Baggerly, K.A., Edmonson, S.R., Morris, J.S. & Coombes, K.R. High-resolution serum proteomic patterns for ovarian cancer detection. Endocr. Relat. Cancer 11, 583–584, author reply 585–587 (2004).
Ambroise, C. & McLachlan, G.J. Selection bias in gene extraction on the basis of microarray gene-expression data. Proc. Natl. Acad. Sci. USA 99, 6562–6566 (2002).
Simon, R. Using DNA microarrays for diagnostic and prognostic prediction. Expert Rev. Mol. Diagn. 3, 587–595 (2003).
Dobbin, K.K. et al. Interlaboratory comparability study of cancer gene expression analysis using oligonucleotide microarrays. Clin. Cancer Res. 11, 565–572 (2005).
Shedden, K. et al. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat. Med. 14, 822–827 (2008).
Parry, R.M. et al. K-nearest neighbors (KNN) models for microarray gene-expression analysis and reliable clinical outcome prediction. Pharmacogenomics J. 10, 292–309 (2010).
Dupuy, A. & Simon, R.M. Critical review of published microarray studies for cancer outcome and guidelines on statistical analysis and reporting. J. Natl. Cancer Inst. 99, 147–157 (2007).
Dave, S.S. et al. Prediction of survival in follicular lymphoma based on molecular features of tumor-infiltrating immune cells. N. Engl. J. Med. 351, 2159–2169 (2004).
Tibshirani, R. Immune signatures in follicular lymphoma. N. Engl. J. Med. 352, 1496–1497, author reply 1496–1497 (2005).
Shi, W. et al. Functional analysis of multiple genomic signatures demonstrates that classification algorithms choose phenotype-related genes. Pharmacogenomics J. 10, 310–323 (2010).
Robinson, G.K. That BLUP is a good thing: the estimation of random effects. Stat. Sci. 6, 15–32 (1991).
Hothorn, T., Hornik, K. & Zeileis, A. Unbiased recursive partitioning: a conditional inference framework. J. Comput. Graph. Statist. 15, 651–674 (2006).
Boutros, P.C. et al. Prognostic gene signatures for non-small-cell lung cancer. Proc. Natl. Acad. Sci. USA 106, 2824–2828 (2009).
Popovici, V. et al. Effect of training sample size and classification difficulty on the accuracy of genomic predictors. Breast Cancer Res. 12, R5 (2010).
Yousef, W.A., Wagner, R.F. & Loew, M.H. Assessing classifiers from two independent data sets using ROC analysis: a nonparametric approach. IEEE Trans. Pattern Anal. Mach. Intell. 28, 1809–1817 (2006).
Gur, D., Wagner, R.F. & Chan, H.P. On the repeated use of databases for testing incremental improvement of computer-aided detection schemes. Acad. Radiol. 11, 103–105 (2004).
Allison, D.B., Cui, X., Page, G.P. & Sabripour, M. Microarray data analysis: from disarray to consolidation and consensus. Nat. Rev. Genet. 7, 55–65 (2006).
Wood, I.A., Visscher, P.M. & Mengersen, K.L. Classification based upon gene expression data: bias and precision of error rates. Bioinformatics 23, 1363–1370 (2007).
Luo, J. et al. A comparison of batch effect removal methods for enhancement of prediction performance using MAQC-II microarray gene expression data. Pharmacogenomics J. 10, 278–291 (2010).
Fan, X. et al. Consistency of predictive signature genes and classifiers generated using different microarray platforms. Pharmacogenomics J. 10, 247–257 (2010).
Huang, J. et al. Genomic indicators in the blood predict drug-induced liver injury. Pharmacogenomics J. 10, 267–277 (2010).
Oberthuer, A. et al. Comparison of performance of one-color and two-color gene-expression analyses in predicting clinical endpoints of neuroblastoma patients. Pharmacogenomics J. 10, 258–266 (2010).
Hong, H. et al. Assessing sources of inconsistencies in genotypes and their effects on genome-wide association studies with HapMap samples. Pharmacogenomics J. 10, 364–374 (2010).
Thomas, R.S., Pluta, L., Yang, L. & Halsey, T.A. Application of genomic biomarkers to predict increased lung tumor incidence in 2-year rodent cancer bioassays. Toxicol. Sci. 97, 55–64 (2007).
Fielden, M.R., Brennan, R. & Gollub, J. A gene expression biomarker provides early prediction and mechanistic assessment of hepatic tumor induction by nongenotoxic chemicals. Toxicol. Sci. 99, 90–100 (2007).
Ganter, B. et al. Development of a large-scale chemogenomics database to improve drug candidate selection and to understand mechanisms of chemical toxicity and action. J. Biotechnol. 119, 219–244 (2005).
Lobenhofer, E.K. et al. Gene expression response in target organ and whole blood varies as a function of target organ injury phenotype. Genome Biol. 9, R100 (2008).
Symmans, W.F. et al. Total RNA yield and microarray gene expression profiles from fine-needle aspiration biopsy and core-needle biopsy samples of breast carcinoma. Cancer 97, 2960–2971 (2003).
Gong, Y. et al. Determination of oestrogen-receptor status and ERBB2 status of breast carcinoma: a gene-expression profiling study. Lancet Oncol. 8, 203–211 (2007).
Hess, K.R. et al. Pharmacogenomic predictor of sensitivity to preoperative chemotherapy with paclitaxel and fluorouracil, doxorubicin, and cyclophosphamide in breast cancer. J. Clin. Oncol. 24, 4236–4244 (2006).
Zhan, F. et al. The molecular classification of multiple myeloma. Blood 108, 2020–2028 (2006).
Shaughnessy, J.D. Jr. et al. A validated gene expression model of high-risk multiple myeloma is defined by deregulated expression of genes mapping to chromosome 1. Blood 109, 2276–2284 (2007).
Barlogie, B. et al. Thalidomide and hematopoietic-cell transplantation for multiple myeloma. N. Engl. J. Med. 354, 1021–1030 (2006).
Zhan, F., Barlogie, B., Mulligan, G., Shaughnessy, J.D. Jr. & Bryant, B. High-risk myeloma: a gene expression based risk-stratification model for newly diagnosed multiple myeloma treated with high-dose therapy is predictive of outcome in relapsed disease treated with single-agent bortezomib or high-dose dexamethasone. Blood 111, 968–969 (2008).
Chng, W.J., Kuehl, W.M., Bergsagel, P.L. & Fonseca, R. Translocation t(4;14) retains prognostic significance even in the setting of high-risk molecular signature. Leukemia 22, 459–461 (2008).
Decaux, O. et al. Prediction of survival in multiple myeloma based on gene expression profiles reveals cell cycle and chromosomal instability signatures in high-risk patients and hyperdiploid signatures in low-risk patients: a study of the Intergroupe Francophone du Myelome. J. Clin. Oncol. 26, 4798–4805 (2008).
Oberthuer, A. et al. Customized oligonucleotide microarray gene expression-based classification of neuroblastoma patients outperforms current clinical risk stratification. J. Clin. Oncol. 24, 5070–5078 (2006).
Acknowledgements
The MAQC-II project was funded in part by the FDA's Office of Critical Path Programs (to L.S.). Participants from the National Institutes of Health (NIH) were supported by the Intramural Research Program of NIH, Bethesda, Maryland or the Intramural Research Program of the NIH, National Institute of Environmental Health Sciences (NIEHS), Research Triangle Park, North Carolina. J.F. was supported by the Division of Intramural Research of the NIEHS under contract HHSN273200700046U. Participants from the Johns Hopkins University were supported by grants from the NIH (1R01GM083084-01 and 1R01RR021967-01A2 to R.A.I. and T32GM074906 to M.M.). Participants from the Weill Medical College of Cornell University were partially supported by the Biomedical Informatics Core of the Institutional Clinical and Translational Science Award RFA-RM-07-002. F.C. acknowledges resources from The HRH Prince Alwaleed Bin Talal Bin Abdulaziz Alsaud Institute for Computational Biomedicine and from the David A. Cofrin Center for Biomedical Information at Weill Cornell. The data set from The Hamner Institutes for Health Sciences was supported by a grant from the American Chemistry Council's Long Range Research Initiative. The breast cancer data set was generated with support of grants from NIH (R-01 to L.P.), The Breast Cancer Research Foundation (to L.P. and W.F.S.) and the Faculty Incentive Funds of the University of Texas MD Anderson Cancer Center (to W.F.S.). The data set from the University of Arkansas for Medical Sciences was supported by National Cancer Institute (NCI) PO1 grant CA55819-01A1, NCI R33 Grant CA97513-01, Donna D. and Donald M. Lambert Lebow Fund to Cure Myeloma and Nancy and Steven Grand Foundation. We are grateful to the individuals whose gene expression data were used in this study. All MAQC-II participants freely donated their time and reagents for the completion and analyses of the MAQC-II project. The MAQC-II consortium also thanks R. O'Neill for his encouragement and coordination among FDA Centers on the formation of the RBWG. The MAQC-II consortium gratefully dedicates this work in memory of R.F. Wagner who enthusiastically worked on the MAQC-II project and inspired many of us until he unexpectedly passed away in June 2008.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
Many of the MAQC-II participants are employed by companies that manufacture gene expression products and/or perform testing services.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 3–8, Supplementary Data and Supplementary Figs. 1–13 (PDF 4568 kb)
Supplementary Table 1
UniqueModels19779_PerformanceMetrics (XLS 14906 kb)
Supplementary Table 2
Swap_UniqueModels13287_PerformanceMetrics (XLS 12587 kb)
Rights and permissions
About this article
Cite this article
MAQC Consortium. The MicroArray Quality Control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models. Nat Biotechnol 28, 827–838 (2010). https://doi.org/10.1038/nbt.1665
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1665
This article is cited by
-
AF1q is a universal marker of neuroblastoma that sustains N-Myc expression and drives tumorigenesis
Oncogene (2024)
-
Basal–epithelial subpopulations underlie and predict chemotherapy resistance in triple-negative breast cancer
EMBO Molecular Medicine (2024)
-
Enhancing prognostic power in multiple myeloma using a plasma cell signature derived from single-cell RNA sequencing
Blood Cancer Journal (2024)
-
The age-specific comorbidity burden of mild cognitive impairment: a US claims database study
Alzheimer's Research & Therapy (2023)
-
Notch-based gene signature for predicting the response to neoadjuvant chemotherapy in triple-negative breast cancer
Journal of Translational Medicine (2023)