Abstract
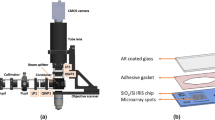
DNA microarrays are used for gene-expression profiling, single-nucleotide polymorphism detection and disease diagnosis1,2,3. A persistent challenge in this area is the lack of microarray screening technology suitable for integration into routine clinical care4,5. Here, we describe a method for sensitive and label-free electrostatic readout of DNA or RNA hybridization on microarrays. The electrostatic properties of the microarray are measured from the position and motion of charged microspheres randomly dispersed over the surface. We demonstrate nondestructive electrostatic imaging with 10-μm lateral resolution over centimeter-length scales, which is four-orders of magnitude larger than that achievable with conventional scanning electrostatic force microscopy. Changes in surface charge density as a result of specific hybridization can be detected and quantified with 50-pM sensitivity, single base-pair mismatch selectivity and in the presence of complex background. Because the naked eye is sufficient to read out hybridization, this approach may facilitate broad application of multiplexed assays.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
McGuire, A., Cho, M., McGuire, S. & Caulfield, T. Medicine: the future of personal genomics. Science 317, 1687 (2007).
Cobb, J.P. et al. Application of genome-wide expression analysis to human health and disease. Proc. Natl. Acad. Sci. USA 102, 4801–4806 (2005).
Barken, K., Haagensen, J.J. & Tolker-Nielsen, T. Advances in nucleic acid-based diagnostics of bacterial infections. Clin. Chim. Acta 384, 1–11 (2007).
Aitman, T.J. Science, medicine, and the future: DNA microarrays in medical practice. Br. Med. J. 323, 611–615 (2001).
Abdullah-Sayani, A., Bueno-de-Mesquita, J.M. & van de Vijver, Marc J. Technology insight: tuning into the genetic orchestra using microarrays—limitations of DNA microarrays in clinical practice. Nat. Clin. Pract. Oncol. 3, 501–516 (2006).
Global Health Diagnostics Forum. The right tools can save lives. Nature 444, 681 (2006).
Mabey, D., Peeling, R.W., Ustianowski, A. & Perkins, M.D. Diagnostics for the developing world. Nat. Rev. Microbiol. 2, 231–240 (2004).
Steinmetz, L.M. & Davis, R.W. Maximizing the potential of functional genomics. Nat. Rev. Genet. 5, 190–201 (2004).
Fang, S., Lee, H.J., Wark, A.W. & Corn, R.M. Attomole microarray detection of microRNAs by nanoparticle-amplified SPR imaging measurements of surface polyadenylation reactions. J. Am. Chem. Soc. 128, 14044–14046 (2006).
Cooper, M.A. Optical biosensors in drug discovery. Nat. Rev. Drug Discov. 1, 515–528 (2002).
Fan, C., Plaxco, K.W. & Heeger, A.J. Electrochemical interrogation of conformational changes as a reagentless method for the sequence-specific detection of DNA. Proc. Natl. Acad. Sci. USA 100, 9134–9137 (2003).
Drummond, T.G., Hill, M.G. & Barton, J.K. Electrochemical DNA sensors. Nat. Biotechnol. 21, 1192–1199 (2003).
Nilsson, K.P. & Inganas, O. Chip and solution detection of DNA hybridization using a luminescent zwitterionic polythiophene derivative. Nat. Mater. 2, 419–424 (2003).
Zhou, D., Sinniah, K., Abell, C. & Rayment, T. Label-free detection of DNA hybridization at the nanoscale: a highly sensitive and selective approach using atomic-force microscopy. Angew. Chem. Int. Ed. 42, 4934–4937 (2003).
Sinensky, A.K. & Belcher, A.M. Label-free and high-resolution protein//DNA nanoarray analysis using Kelvin probe force microscopy. Nat. Nanotechnol. 2, 653–659 (2007).
Zhang, J. et al. Rapid and label-free nanomechanical detection of biomarker transcripts in human RNA. Nat. Nanotechnol. 1, 214–220 (2006).
Hahm, J.-i. & Lieber, C.M. Direct ultrasensitive electrical detection of DNA and DNA sequence variations using nanowire nanosensors. Nano Lett. 4, 51–54 (2004).
Fritz, J., Cooper, E., Gaudet, S., Sorger, P.K. & Manalis, S.R. Electronic detection of DNA by its intrinsic molecular charge. Proc. Natl. Acad. Sci. USA 99, 14142 (2002).
Hood, L., Heath, J., Phelps, M. & Lin, B. Systems biology and new technologies enable predictive and preventative medicine. Science 306, 640–643 (2004).
Drummond, T.G., Hill, M.G. & Barton, J.K. Electrochemical DNA sensors. Nat. Biotechnol. 21, 1192–1199 (2003).
't Hoen, P.C., deKort, F., vanOmmen, G.J.B. & denDunnen, J. Fluorescent labelling of cRNA for microarray applications. Nucleic Acids Res. 31, e20 (2003).
Dove, A. Business office feature: from morgan to microarrays: gene mapping hits the big time. Science 318, 473–478 (2007).
Butt, H.J., Capella, B. & Kappl, M. Force measurements with the atomic force microscope: technique, interpretation and applications. Surf. Sci. Rep. 59, 1–152 (2005).
Despont, M., Drechsler, U., Yu, R., Pogge, H.B. & Vettiger, P. Wafer-scale microdevice transfer/interconnect: its application in an AFM-based data-storage system. J. Mem. S. 13, 895–901 (2004).
Russel, W.B., Saville, D.A. & Schowalter, W.R. Electrostatics. in Colloidal Dispersions (ed. Batchelor, G. K.) (Cambridge University Press, Cambridge, UK, 1991).
Schilling, J., Sengupta, K., Goennenwein, S., Bausch, A.R. & Sackmann, E. Absolute interfacial distance measurements by dual-wavelength reflection interference contrast microscopy. Phys. Rev. E. 69, 021901 (2004).
Clack, N.G. & Groves, J.T. Many-particle tracking with nanometer resolution in three dimensions by reflection interference contrast microscopy. Langmuir 21, 6430–6435 (2005).
Peterson, A., Heaton, R. & Georgiadis, R. The effect of surface probe density on DNA hybridization. Nucleic Acids Res. 29, 5163–5168 (2001).
Heyries, K.A., Marquette, C.A. & Blum, L.J. Straightforward protein immobilization on Sylgard 184 PDMS microarray surface. Langmuir 23, 4523–4527 (2007).
Yu, A.A. et al. High resolution printing of DNA feature on poly(methyl methacrylate) substrates using supramolecular nano-stamping. J. Am. Chem. Soc. 127, 16774–16775 (2005).
Salaita, K., Wang, Y.H. & Mirkin, C.A. Applications of dip-pen nanolithography. Nat. Nano. 2, 145–155 (2007).
MacBeath, G. Protein microarrays and proteomics. Nat. Genet. 32, 526–532 (2002).
Winssinger, N., Pianowski, Z. & Debaene, F. Probing biology with small molecule microarrays (SMM). Top. Curr. Chem. 278, 311–342 (2007).
de Gans, B.-J. & Schubert, U.S. Inkjet printing of polymer micro-arrays and libraries: instrumentation, requirements, and perspective. Macromol. Rapid Commun. 24, 659–666 (2003).
Cassell, A.M., Verma, S., Delzeit, L., Meyyappan, M. & Han, J. Combinatorial optimization of heterogeneous catalysts used in the growth of carbon nanotubes. Langmuir 17, 260–264 (2001).
Acknowledgements
We thank Lance Kizer and the UC Berkeley Functional Genomics Laboratory for assistance in generating microarrays. This work was supported by the Director, Office of Science, Office of Basic Energy Sciences of the US Department of Energy under Contract No. DE-AC03-76SF00098. N.G.C. also received partial support from the National Science Foundation through The Center on Polymer Interfaces and Macromolecular Assemblies.
Author information
Authors and Affiliations
Contributions
N.G.C. and K.S. designed and implemented all experiments. N.G.C., K.S. and J.T.G. conceived of ideas and wrote the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Figures 1–4, Table 1, Methods, Note (PDF 948 kb)
Rights and permissions
About this article
Cite this article
Clack, N., Salaita, K. & Groves, J. Electrostatic readout of DNA microarrays with charged microspheres. Nat Biotechnol 26, 825–830 (2008). https://doi.org/10.1038/nbt1416
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1416
This article is cited by
-
Submillimetre Network Formation by Light-induced Hybridization of Zeptomole-level DNA
Scientific Reports (2016)
-
Visualizing mechanical tension across membrane receptors with a fluorescent sensor
Nature Methods (2012)
-
An ultrasensitive universal detector based on neutralizer displacement
Nature Chemistry (2012)
-
Engineering supported membranes for cell biology
Medical & Biological Engineering & Computing (2010)
-
DNA nanomechanics allows direct digital detection of complementary DNA and microRNA targets
Nature (2009)