Abstract
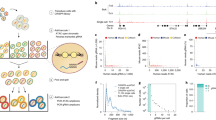
Our understanding of how chromatin structure influences cellular processes such as transcription and replication has been limited by a lack of nucleosome-positioning data in human cells. We describe a high-resolution microarray approach combined with an analysis algorithm to examine nucleosome positioning in 3,692 promoters within seven human cell lines. Unlike unexpressed genes without transcription-preinitiation complexes at their promoters, expressed genes or genes containing preinitiation complexes exhibit characteristic nucleosome-free regions at their transcription start sites. The combination of these nucleosome data with chromatin immunoprecipitation–chip analyses reveals that the melanocyte master regulator microphthalmia-associated transcription factor (MITF) predominantly binds nucleosome-free regions, supporting the model that nucleosomes limit sequence accessibility. This study presents a global view of human nucleosome positioning and provides a high-throughput tool for analyzing chromatin structure in development and disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Lu, Q., Wallrath, L.L. & Elgin, S.C. Nucleosome positioning and gene regulation. J. Cell. Biochem. 55, 83–92 (1994).
Widom, J. Structure, dynamics, and function of chromatin in vitro. Annu. Rev. Biophys. Biomol. Struct. 27, 285–327 (1998).
Lee, T.I. & Young, R.A. Transcription of eukaryotic protein-coding genes. Annu. Rev. Genet. 34, 77–137 (2000).
Workman, J.L. & Kingston, R.E. Alteration of nucleosome structure as a mechanism of transcriptional regulation. Annu. Rev. Biochem. 67, 545–579 (1998).
Becker, P.B. & Horz, W. ATP-dependent nucleosome remodeling. Annu. Rev. Biochem. 71, 247–273 (2002).
Mellor, J. The dynamics of chromatin remodeling at promoters. Mol. Cell 19, 147–157 (2005).
Yuan, G.C. et al. Genome-scale identification of nucleosome positions in S. cerevisiae. Science 309, 626–630 (2005).
Segal, E. et al. A genomic code for nucleosome positioning. Nature 442, 772–778 (2006).
Ioshikhes, I.P., Albert, I., Zanton, S.J. & Pugh, B.F. Nucleosome positions predicted through comparative genomics. Nat. Genet. 38, 1210–1215 (2006).
Carroll, J.S. et al. Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein FoxA1. Cell 122, 33–43 (2005).
Kim, T.H. et al. A high-resolution map of active promoters in the human genome. Nature 436, 876–880 (2005).
Mucha, M., Lisowska, K., Goc, A. & Filipski, J. Nuclease-hypersensitive chromatin formed by a CpG island in human DNA cloned as an artificial chromosome in yeast. J. Biol. Chem. 275, 1275–1278 (2000).
Reid, D.G., Salisbury, S.A., Brown, T. & Williams, D.H. Conformations of two duplex forms of d(TCGA) in slow-exchange equilibrium characterized by NMR. Biochemistry 24, 4325–4332 (1985).
Wang, Y.H., Amirhaeri, S., Kang, S., Wells, R.D. & Griffith, J.D. Preferential nucleosome assembly at DNA triplet repeats from the myotonic dystrophy gene. Science 265, 669–671 (1994).
Horz, W. & Altenburger, W. Sequence specific cleavage of DNA by micrococcal nuclease. Nucleic Acids Res. 9, 2643–2658 (1981).
Anderson, J.D. & Widom, J. Sequence and position-dependence of the equilibrium accessibility of nucleosomal DNA target sites. J. Mol. Biol. 296, 979–987 (2000).
Lee, C.K., Shibata, Y., Rao, B., Strahl, B.D. & Lieb, J.D. Evidence for nucleosome depletion at active regulatory regions genome-wide. Nat. Genet. 36, 900–905 (2004).
Pokholok, D.K. et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122, 517–527 (2005).
Mito, Y., Henikoff, J.G. & Henikoff, S. Genome-scale profiling of histone H3.3 replacement patterns. Nat. Genet. 37, 1090–1097 (2005).
Hoffmann, A., Oelgeschlager, T. & Roeder, R.G. Considerations of transcriptional control mechanisms: do TFIID-core promoter complexes recapitulate nucleosome-like functions? Proc. Natl. Acad. Sci. USA 94, 8928–8935 (1997).
Garraway, L.A. et al. Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature 436, 117–122 (2005).
Bernstein, B.E., Liu, C.L., Humphrey, E.L., Perlstein, E.O. & Schreiber, S.L. Global nucleosome occupancy in yeast. Genome Biol. 5, R62 (2004).
Du, J. et al. Critical role of CDK2 for melanoma growth linked to its melanocyte-specific transcriptional regulation by MITF. Cancer Cell 6, 565–576 (2004).
McGill, G.G. et al. Bcl2 regulation by the melanocyte master regulator Mitf modulates lineage survival and melanoma cell viability. Cell 109, 707–718 (2002).
Chen, C. & Yang, T.P. Nucleosomes are translationally positioned on the active allele and rotationally positioned on the inactive allele of the HPRT promoter. Mol. Cell. Biol. 21, 7682–7695 (2001).
Zhang, Y. et al. Reproducible and inexpensive probe preparation for oligonucleotide arrays. Nucleic Acids Res. 29, E66 (2001).
Kouskouti, A. & Talianidis, I. Histone modifications defining active genes persist after transcriptional and mitotic inactivation. EMBO J. 24, 347–357 (2005).
Dai, M. et al. Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res. 33, e175 (2005).
Johnson, W.E. et al. Model-based analysis of tiling-arrays for ChIP-chip. Proc. Natl. Acad. Sci. USA 103, 12457–12462 (2006).
Fivaz, J., Bassi, M.C., Price, M., Pinaud, S. & Mirkovitch, J. Precisely positioned nucleosomes are not essential for c-fos gene regulation in vivo. Gene 255, 169–184 (2000).
Acknowledgements
We thank H. Widlund, E. Feige, V. Igras, I. Davis, W. Li and C. Meyer for helpful discussions and support. This work was supported by a grant from the US National Institutes of Health to D.E.F. and the Claudia Adams Barr Award for Innovative Basic Cancer Research to X.S.L. D.E.F. is Distinguished Clinical Scholar of the Doris Duke Charitable Foundation, and Charles and Jan Nirenberg Fellow in Pediatric Oncology at Dana-Farber Cancer Institute. J.S.S. was supported by the Claudia Adams Barr Award from Dana Farber Cancer Institute.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Genes with at least 2 additional positioned nucleosomes present in MCF7 and T47D but not in MEC. (XLS 60 kb)
Rights and permissions
About this article
Cite this article
Ozsolak, F., Song, J., Liu, X. et al. High-throughput mapping of the chromatin structure of human promoters. Nat Biotechnol 25, 244–248 (2007). https://doi.org/10.1038/nbt1279
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1279
This article is cited by
-
Construction of a transposase accessible chromatin landscape reveals chromatin state of repeat elements and potential causal variant for complex traits in pigs
Journal of Animal Science and Biotechnology (2022)
-
Comparative analysis and prediction of nucleosome positioning using integrative feature representation and machine learning algorithms
BMC Bioinformatics (2021)
-
Role of cell-type specific nucleosome positioning in inducible activation of mammalian promoters
Nature Communications (2020)
-
Born to run: control of transcription elongation by RNA polymerase II
Nature Reviews Molecular Cell Biology (2018)
-
Cell Type-Specific Survey of Epigenetic Modifications by Tandem Chromatin Immunoprecipitation Sequencing
Scientific Reports (2018)