Abstract
The B cell antigen receptor (BCR) is composed of a membrane-bound class M, D, G, E or A immunoglobulin for antigen recognition1,2,3 and a disulfide-linked Igα (also known as CD79A) and Igβ (also known as CD79B) heterodimer (Igα/β) that functions as the signalling entity through intracellular immunoreceptor tyrosine-based activation motifs (ITAMs)4,5. The organizing principle of the BCR remains unknown. Here we report cryo-electron microscopy structures of mouse full-length IgM BCR and its Fab-deleted form. At the ectodomain (ECD), the Igα/β heterodimer mainly uses Igα to associate with Cµ3 and Cµ4 domains of one heavy chain (µHC) while leaving the other heavy chain (µHC′) unbound. The transmembrane domain (TMD) helices of µHC and µHC′ interact with those of the Igα/β heterodimer to form a tight four-helix bundle. The asymmetry at the TMD prevents the recruitment of two Igα/β heterodimers. Notably, the connecting peptide between the ECD and TMD of µHC intervenes in between those of Igα and Igβ to guide TMD assembly through charge complementarity. Weaker but distinct density for the Igβ ITAM nestles next to the TMD, suggesting potential autoinhibition of ITAM phosphorylation. Interfacial analyses suggest that all BCR classes utilize a general organizational architecture. Our studies provide a structural platform for understanding B cell signalling and designing rational therapies against BCR-mediated diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data and materials reported in the main and supplementary data are available upon reasonable request. The electron density maps of 8.2 Å full-length IgM BCR, 3.3 Å IgM BCRΔfab and 3.9 Å IgM BCRΔfab have been deposited in the Electron Microscopy Data Bank (EMDB) with accession codes EMD-27848, EMD-27888 and EMD-28030, respectively. The atomic coordinates for IgM BCRΔfab and full-length IgM BCR have been deposited in the Protein Data Bank with the accession code of 8E4C and 8EMA.
References
Venkitaraman, A. R., Williams, G. T., Dariavach, P. & Neuberger, M. S. The B-cell antigen receptor of the five immunoglobulin classes. Nature 352, 777–781 (1991).
Alber, G., Flaswinkel, H., Kim, K.-M., Weiser, P. & Reth, M. in Progress in Immunology Vol. VIII (eds Gergely, J. et al.) 27–33 (Springer, 1993).
Hombach, J., Tsubata, T., Leclercq, L., Stappert, H. & Reth, M. Molecular components of the B-cell antigen receptor complex of the lgM class. Nature 343, 760–762 (1990).
Campbell, K. S. & Cambier, J. C. B lymphocyte antigen receptors (mIg) are non-covalently associated with a disulfide linked, inducibly phosphorylated glycoprotein complex. EMBO J. 9, 441–448 (1990).
Schamel, W. W. A. & Reth, M. Monomeric and oligomeric complexes of the B cell antigen receptor. Immunity 13, 5–14 (2000).
McHeyzer-Williams, L. J. & McHeyzer-Williams, M. G. Antigen-specific memory B cell development. Annu. Rev. Immunol. 23, 487–513 (2005).
Reth, M. & Wienands, J. Initiation and processing of signals from the B cell antigen receptor. Annu. Rev. Immunol. 15, 453–479 (1997).
Reth, M. Antigen receptors on B lymphocytes. Annu. Rev. Immunol. 10, 97–121 (1992).
Alt, F. W. & Bothwell, L. M. Synthesis of secreted and membrane-bound immunoglobulin mu heavy chains is directed by mRNAs that differ at their 3′ ends. Cell 20, 293–301 (1992).
Early, P. et al. Two mRNAs can be produced from a single immunoglobulin p gene by alternative RNA processing pathways. Cell 20, 313–3197 (1980).
Stavnezer, J. & Schrader, C. E. Ig heavy chain class switch recombination: mechanism and regulation. J. Immunol. 193, 5370–5378 (2014).
Stavnezer, J., Guikema, J. E. J. & Schrader, C. E. Mechanism and regulation of class switch recombination. Annu. Rev. Immunol. 26, 261–292 (2008).
Esser, C. Immunoglobulin class switching: molecular and cellular analysis. Annu. Rev. Immunol. 8, 717–735 (1990).
Reth, M. Antigen receptor tail clue. Nature 338, 383–384 (1989).
Samelson, L. E. & Klausner, R. D. Tyrosine kinases and tyrosine-based activation motifs. Current research on activation via the T cell antigen receptor. J. Biol. Chem. 267, 24913–24916 (1992).
Flaswinkel, H. & Reth, M. Dual role of the tyrosine activation motif of the Ig-alpha protein during signal transduction via the B cell antigen receptor. EMBO J. 13, 83–89 (1994).
Roux, K. H., Strelets, L., Brekke, O. H., Sandlie, I. & Michaelsen, T. E. Comparisons of the ability of human IgG3 hinge mutants, IgM, IgE, and IgA2, to form small immune complexes: a role for flexibility and geometry. J. Immunol. 161, 4083–4090 (1998).
Adlersberg, J. B. The immunoglobulin hinge (interdomain) region. Ric. Clin. Lab. 6, 191 (1976).
Sandin, S., Öfverstedt, L.-G., Wikström, A.-C., Wrange, Ö. & Skoglund, U. Structure and flexibility of individual immunoglobulin G molecules in solution. Structure 12, 409–415 (2004).
Wurzburg, B. A., Garman, S. C. & Jardetzky, T. S. Structure of the human IgE–Fc Cε3–Cε4 reveals conformational flexibility in the antibody effector domains. Immunity 13, 375–385 (2000).
Mirdita, M. et al. ColabFold—Making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Müller, R. et al. High-resolution structures of the IgM Fc domains reveal principles of its hexamer formation. Proc. Natl Acad. Sci. USA 110, 10183–10188 (2013).
Ban, N. et al. Structure of an anti‐idiotypic Fab against feline peritonitis virus‐neutralizing antibody and a comparison with the complexed Fab. FASEB J. 9, 107–114 (1995).
Dong, D. et al. Structural basis of assembly of the human T cell receptor–CD3 complex. Nature 573, 546–552 (2019).
Kato, K. et al. High-resolution cryo-EM structure of photosystem II reveals damage from high-dose electron beams. Commun. Biol. 4, 382 (2021).
Radaev, S. et al. Structural and functional studies of Igαβ and its assembly with the B Cell antigen receptor. Structure 18, 934–943 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Li, Y. et al. Structural insights into immunoglobulin M. Science 367, 1014–1017 (2020).
Hiramoto, E. et al. The IgM pentamer is an asymmetric pentagon with an open groove that binds the AIM protein. Sci. Adv. 4, eaau1199 (2018).
Sušac, L. et al. Structure of a fully assembled tumor-specific T cell receptor ligated by pMHC. Cell 185, 3201–3213.e19 (2022).
Schamel, W. W. A. & Reth, M. Stability of the B cell antigen receptor complex. Mol. Immunol. 37, 253–259 (2000).
Gottwick, C. et al. A symmetric geometry of transmembrane domains inside the B cell antigen receptor complex. Proc. Natl Acad. Sci. USA 116, 13468–13473 (2019).
Schwans, J. P. et al. Use of anion–aromatic interactions to position the general base in the ketosteroid isomerase active site. Proc. Natl Acad. Sci. USA 110, 11308–11313 (2013).
Kwong, P. D. et al. Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 393, 648–659 (1998).
Parikh, V. S. et al. Differential structure–function requirements of the transmembrane domain of the B cell antigen receptor. J. Exp. Med. 176, 1025–1031 (1992).
Shaw, C., Mitchell, N., Weaver, Y. K. & Abbas, A. K. Mutations of immunoglobulin transmembrane and cytoplasmic domains: effects on intracellular signaling and antigen presentation. Cell 63, 381–392 (1990).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430, 2237–2243 (2018).
Tolar, P., Sohn, H. W. & Pierce, S. K. The initiation of antigen-induced B cell antigen receptor signaling viewed in living cells by fluorescence resonance energy transfer. Nat. Immunol. 6, 1168–1176 (2005).
Su, Q. et al. Cryo-EM structure of the human IgM B cell receptor. Science 337, 875–880 (2022).
Ma, X. et al. Cryo-EM structures of two human B cell receptor isotypes. Science 377, 880–885 (2022).
Yang, J. & Reth, M. Oligomeric organization of the B-cell antigen receptor on resting cells. Nature 467, 465–469 (2010).
Gold, M. R. & Reth, M. G. Antigen receptor function in the context of the nanoscale organization of the B cell membrane. Annu. Rev. Immunol. 37, 97–123 (2019).
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013).
Zheng, S. Q. et al. MotionCor2 - anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Acknowledgements
We thank members of the Wu laboratory, especially Y. Zheng, for helpful discussions; R. Walsh, S. Sterling. M. Mayer and S. Rawson at the Harvard Cryo-EM Center for Structural Biology for cryo-EM training and data collection; K. Song at the University of Massachusetts Cryo-EM Core for screening and preliminary dataset collection; and J. Myers, V. Rayaprolu and the Pacific Northwest Center for Cryo-EM at Oregon Health and Science University for preliminary dataset collection, under the NIH grant U24GM129547 and accessed through EMSL (grid.436923.9), a DOE Office of Science User Facility sponsored by the Office of Biological and Environmental Research. We thank SBGrid for software and computing support. Research reported in this publication was supported by the National Institutes of Health through RO1 grant AI145656 (to M.R.), the DFG through TRR130-P02 (to M.R.), Germany’s Excellence Strategy (CIBSS-EXC-2189, Project ID390939984) and the Charles A. King Trust postdoctoral research fellowship (to Y.D.).
Author information
Authors and Affiliations
Contributions
M.R. and H.W. conceptualized the project. F.B.-B., D.S. and J.Y. made the stable cell lines for BCR expression. Z.L. validated BCR expression by flow cytometry under the supervision of F.W.A. Y.D. resorted and grew the cells, and purified the BCR samples for negative staining and cryo-EM. Y.D. and X.P. performed cryo-EM data acquisition. X.P. and Y.D. performed data processing. Y.D. and X.P. built and refined the model. Y.D. made the figures. H.W., Y.D. and M.R. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Robert Tampé and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Protein purification and cryo-EM data processing of full-length IgM BCR.
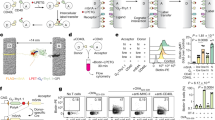
a, Selection of J558L B cells that expressed IgM BCR. Domain organization of IgM BCR and surface expression of IgM (APC) and Igα (GFP) by flow cytometry are shown. b-c, Purification of IgM BCR shown by SDS-PAGE, Blue-native (BN) PAGE and western blotting using antibodies against the individual subunits. H: heavy chain; L: light chain. d, Gel filtration profiles of IgM BCR. The peak of IgM BCR is shaded in grey. e-f, Representative 2D classes of IgM BCR (e) and specifically at its Fab region (f). g-h, Data processing flow chart (g) and local resolution distribution of IgM BCR (h).
Extended Data Fig. 2 Protein purification, raw images and 2D classifications of IgM BCRΔFab.
a, Selection of J558L B cells that expressed IgM BCRΔFab. Domain organization of IgM BCRΔFab and surface expression of IgM (APC) and Igα (GFP) by flow cytometry are shown. b-c, Purification of IgM BCRΔFab shown by SDS-PAGE, Blue-native (BN) PAGE and western blotting using antibodies against the individual subunits. H: heavy chain. d, Gel filtration profiles of IgM BCRΔFab. The peak of IgM BCRΔFab is shaded in grey e-f, Representative negative staining EM image (e) and cryo-EM image of IgM BCRΔFab (f). g, Representative 2D classes of IgM BCRΔFab.
Extended Data Fig. 3 Cryo-EM data processing of IgM BCRΔFab.
a, Cryo-EM data processing flow chart of the 3.9 Å intermediate cryo-EM map and the 3.3 Å final cryo-EM map of IgM BCRΔFab. b, Angular distribution of the particles used for the final reconstruction shown as a heat map. c, Fourier shell correlation (FSC) plots. d, 3D FSC plot.
Extended Data Fig. 4 AlphaFold predicted models of IgM BCR.
a, The top ranked model of IgM BCR (without Fab and Cµ2) predicted by AlphaFold, coloured by per-residue pLDDT score. b, Five predicted models of IgM BCR. c, Alignment of the five models of the Igα/β heterodimer at ECD (left) and TMD (right), showing consistent prediction of the ECD interaction (left) and the TMD interaction (right) separately. The CPs of the Igα/β heterodimer were not predicted correctly. d, Alignment of the five models of mIgM at ECD (left) and TMD (right) superimposed with IgM BCRΔFab model, showing consistent prediction of the ECD interaction (left) but incorrect prediction of the TMD interaction (right). The CPs of the mIgM dimer were not predicted correctly. In (c) and (d), the experimentally determined subunits of the IgM BCRΔFab model are shown in their model colours and AlphaFold predicted models are shown in grey.
Extended Data Fig. 5 Model fitting of IgM BCRΔFab in the 3.3 Å cryo-EM map (contour level: 4.0 σ).
a, Ig regions of Igα and Igβ for the Igα/β interaction superimposed with the cryo-EM map. b, Interface between µHC Cµ4 and Igα superimposed with the cryo-EM map. c, Four connecting peptide regions from µHC, µHC’, Igα and Igβ superimposed with the cryo-EM map. Acidic and basic residues are shown and labelled. d-e, TMD helices of µHC, µHC’, Igα and Igβ superimposed with the cryo-EM map (d). Key residues mediating the TMD interaction are shown (e). f, The fitting of five glycosylation sites on the Igα/Igβ heterodimer.
Extended Data Fig. 6 Structural and sequence alignment of Igα and Igβ.
a, Structural alignment between the Ig domains of Igα and Igβ (3.2 Å RMSD). b-c, Sequence alignment of Igα (b) and Igβ (c) among different species. Residues at the interface of Igα/β with µHC and µHC’ are indicated by triangle symbols. Igα C113 and Igβ C135 are marked by red triangle symbols.
Extended Data Fig. 7 Structural and sequence alignment of mIg.
a, Structural alignment of the Cµ3-Cµ4 homodimer of IgM BCR with that from the crystal structure of secreted IgM or sIgM (3.1 Å RMSD, PDB: 6KXS). b, Sequence alignment of mIg among different isotypes. Residues at the interface of µHC and µHC’ with Igα/β are indicated by solid and hollow triangle symbols respectively.
Extended Data Fig. 8 Interaction at TMD and sequence alignment of mIgM.
a—e, The interactions between pairs of helices of TMD. f, Sequence alignment of mIgM among different species.
Extended Data Fig. 9 AlphaFold prediction and secondary structure prediction of Igβ, and sequence conservation mapping on the IgM-BCR surface.
a, AlphaFold prediction of Igβ. The TMD and ICD of Igβ were predicted, showing the pLDDT scores. b, Secondary structure prediction of Igβ (residue 171–228). The ITAM consensus motif is highlighted by red rectangle. c-d, Conserved residue distribution on the Cµ3-Cµ4 homodimer surface (c) and the Igα/Igβ heterodimer surface (d) according to sequence alignment among species. The most conserved residues are shown in dark blue, less conserved residues in light blue, and the remaining residues in their chain colours are defined in Fig. 1d.
Supplementary information
Supplementary Information
This file contains Supplementary Figs. 1 and 2 and Table 1.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dong, Y., Pi, X., Bartels-Burgahn, F. et al. Structural principles of B cell antigen receptor assembly. Nature 612, 156–161 (2022). https://doi.org/10.1038/s41586-022-05412-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05412-7
This article is cited by
-
Raman spectroscopy of lymphocytes from patients with the Epstein–Barr virus infection
Scientific Reports (2024)
-
Structural basis for Fc receptor recognition of immunoglobulin M
Nature Structural & Molecular Biology (2023)
-
Immunoglobulin M perception by FcμR
Nature (2023)
-
Multi-faceted immunoglobulin M meets its elusive receptor
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.