Abstract
Characterizing how variation in the tempo and mode of evolution has structured the phenotypic diversity of extant species is a central goal of macroevolution1,2,3. However, studies are typically limited to a handful of traits4,5,6, providing incomplete information. We analyse morphological diversification in living birds, an ecologically diverse group7, documenting structural scales from ‘pan-skeletal’ proportions down to the localized three-dimensional shape changes of individual bones. We find substantial variation in evolutionary modes among avian subgroups and among skeletal parts, indicating widespread mosaicism and possible differences in the structure of the macroevolutionary landscape across Earth’s main environments. Water-linked groups, especially Aequorlitornithes (waterbirds), have repeatedly explored a large portion of their total morphospace, emphasizing variation in body proportions and in the shape of bones close to the body core, which are functionally related to the mechanics of locomotion8. By contrast, landbirds (Inopinaves) evolved distinct, group-specific body forms early in the aftermath of the K-Pg and subsequently emphasized local shape variation, especially in the head and distal limb bones, which interact more directly with the environment. Passerines, which comprise more than half of all bird species, show a conservative evolutionary dynamic that resulted in low disparity across all skeletal parts. Evidence for early establishment of the morphospace of living birds is clear for some skeletal parts, including beaks and the combined skeletal morphology. However, we find little evidence for early partitioning of that morphospace, contrary to more specific predictions of ‘niche-filling’ models1,9. Nevertheless, early divergence among broad environmental types may have caused an early divergence of evolutionary modes, suggesting an important role for environmental divergence in structuring the radiation of crown-group birds.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw landmark coordinates can be accessed as Supplementary Files linked to this article and accessing our project in OSF Public Repository following this link: https://osf.io/wjk3m/. All three-dimensional meshes can be freely downloaded following the links to Morphosource in Supplementary Materials Table 1.
Code availability
Custom code can be accessed in our project in OSF Public Repository following this link: https://osf.io/wjk3m/.
References
Simpson, G. G. The Major Features of Evolution (Columbia Univ. Press, 1953).
Gould, S. J. Ontogeny and Phylogeny (Harvard Univ. Press, 1985).
Jablonski, D. Approaches to macroevolution: 2. Sorting of variation, some overarching issues, and general conclusions. Evol. Biol. 44, 451–475 (2017).
Cooney, C. R. et al. Mega-evolutionary dynamics of the adaptive radiation of birds. Nature 542, 344–347 (2017).
Felice, R. N. & Goswami, A. Developmental origins of mosaic evolution in the avian cranium. Proc. Natl Acad. Sci. USA 115, 201716437 (2018).
Navalón, G., Marugán-Lobón, J., Bright, J. A., Cooney, C. R. & Rayfield, E. J. The consequences of craniofacial integration for the adaptive radiations of Darwin’s finches and Hawaiian honeycreepers. Nat. Ecol. Evol. 4, 270–278 (2020).
Gill, F. B. Ornithology (Macmillan, 1995).
Kardong, K. V. Vertebrates: Comparative Anatomy, Function, Evolution (Heinle and Heinle Publishers, 1997).
Schluter, D. The Ecology of Adaptive Radiation (Oxford Univ. Press, 2000).
Losos, J. B. Adaptive radiation, ecological opportunity, and evolutionary determinism: American Society of Naturalists EO Wilson Award address. The American Naturalist 175, 623–639 (2010).
Harmon, L. J., Schulte, J. A., Larson, A. & Losos, J. B. Tempo and mode of evolutionary radiation in iguanian lizards. Science 301, 961–964 (2003).
Hughes, M., Gerber, S. & Wills, M. A. Clades reach highest morphological disparity early in their evolution. Proc. Natl Acad. Sci. USA 110, 13875–13879 (2013).
Ackerly, D., Schwilk, D. & Webb, C. Niche evolution and adaptive radiation: testing the order of trait divergence. Ecology 87, S50–S61 (2006).
Gatesy, S. M. & Dial, K. P. Locomotor modules and the evolution of avian flight. Evolution 50, 331–340 (1996).
Dececchi, T. A. & Larsson, H. C. Body and limb size dissociation at the origin of birds: uncoupling allometric constraints across a macroevolutionary transition. Evolution 67, 2741–2752 (2013).
Clarke, J. A. & Middleton, K. M. Mosaicism, modules, and the evolution of birds: results from a Bayesian approach to the study of morphological evolution using discrete character data. Syst. Biol. 57, 185–201 (2008).
Prum, R. O. et al. A comprehensive phylogeny of birds (Aves) using targeted next-generation DNA sequencing. Nature 526, 569–573 (2015).
Oliveros, C. H. et al. Earth history and the passerine superradiation. Proc. Natl Acad. Sci. USA 116, 7916–7925 (2019).
Field, D. J. et al. Early evolution of modern birds structured by global forest collapse at the end-Cretaceous mass extinction. Curr. Biol. 28, 1825–1831. e1822 (2018).
Benito, J. et al. 40 new specimens of Ichthyornis provide unprecedented insight into the postcranial morphology of crownward stem group birds. PeerJ 10, e13919 (2022).
Field, D. J. et al. Timing the extant avian radiation: the rise of modern birds, and the importance of modeling molecular rate variation. Bull. Am. Mus. Nat. Hist. 440, 159–181 (2020).
Feduccia, A. ‘Big bang’ for tertiary birds? Trends Ecol. Evol. 18, 172–176 (2003).
Chiappe, L. M. & Qingjin, M. Birds of Stone: Chinese Avian Fossils from the Age of Dinosaurs (JHU Press, 2016).
Saupe, E. E. et al. Climatic shifts drove major contractions in avian latitudinal distributions throughout the Cenozoic. Proc. Natl Acad. Sci. USA 116, 12895–12900 (2019).
Nudds, R., Dyke, G. & Rayner, J. Forelimb proportions and the evolutionary radiation of Neornithes. Proc. R. Soc. London Ser. B Biol. Sci. 271, S324–S327 (2004).
Wang, X. & Clarke, J. A. Phylogeny and forelimb disparity in waterbirds. Evolution 68, 2847–2860 (2014).
Crouch, N. M. & Ricklefs, R. E. Speciation rate is independent of the rate of evolution of morphological size, shape, and absolute morphological specialization in a large clade of birds. Am. Naturalist 193, E78–E91 (2019).
Brocklehurst, N., Panciroli, E., Benevento, G. L. & Benson, R. B. Mammaliaform extinctions as a driver of the morphological radiation of Cenozoic mammals. Curr. Biol. 31, 2955–2963.e4 (2021).
Osborn, H. F. The law of adaptive radiation. Am. Naturalist 36, 353–363 (1902).
Felice, R. N. et al. Decelerated dinosaur skull evolution with the origin of birds. PLoS Biol. 18, e3000801 (2020).
Felice, R. N., Tobias, J. A., Pigot, A. L. & Goswami, A. Dietary niche and the evolution of cranial morphology in birds. Proc. R. Soc. B 286, 20182677 (2019).
Orkney, A., Bjarnason, A., Tronrud, B. C. & Benson, R. B. Patterns of skeletal integration in birds reveal that adaptation of element shapes enables coordinated evolution between anatomical modules. N. Ecol. Evol. 5, 1250–1258 (2021).
Chira, A. M. & Thomas, G. H. The impact of rate heterogeneity on inference of phylogenetic models of trait evolution. J. Evol. Biol. 29, 2502–2518 (2016).
Rombaut, L. M. et al. Allometric conservatism in the evolution of bird beaks. Evolution Lett. 6, 83–91 (2022).
Navalón, G., Bright, J. A., Marugán‐Lobón, J. & Rayfield, E. J. The evolutionary relationship among beak shape, mechanical advantage, and feeding ecology in modern birds. Evolution 73, 422–435 (2019).
Mayr, G. Avian Evolution: The Fossil Record of Birds and its Paleobiological Significance (John Wiley & Sons, 2016).
Foote, M. The evolution of morphological diversity. Ann. Rev. Ecol. Systematics 28, 129–152 (1997).
Bribiesca-Contreras, F., Parslew, B. & Sellers, W. I. Functional morphology of the forelimb musculature reflects flight and foraging styles in aquatic birds. J. Ornithol. 162, 779–793 (2021).
Phillimore, A. B. et al. Sympatric speciation in birds is rare: insights from range data and simulations. Am. Naturalist 171, 646–657 (2008).
MacArthur, R. H. Population ecology of some warblers of northeastern coniferous forests. Ecology 39, 599–619 (1958).
Cooney, C. R. et al. Sexual selection predicts the rate and direction of colour divergence in a large avian radiation. Nat. Commun. 10, 1773 (2019).
Tobias, J. A. et al. Species coexistence and the dynamics of phenotypic evolution in adaptive radiation. Nature 506, 359 (2014).
Del Hoyo, J. et al. Handbook of the Birds of the World Alive (Lynx Editions, 2017).
Botelho, J. F., Smith-Paredes, D. & Vargas, A. O. Altriciality and the evolution of toe orientation in birds. Evolution. Biol. 42, 502–510 (2015).
Natale, R. & Slater, G. J. The effects of foraging ecology and allometry on avian skull shape vary across levels of phylogeny. https://doi.org/10.1086/720745 (2022).
Pigot, A. L. et al. Macroevolutionary convergence connects morphological form to ecological function in birds. Nat. Ecol. Evol. 4, 230–239 (2020).
Slater, G. J. Iterative adaptive radiations of fossil canids show no evidence for diversity-dependent trait evolution. Proc. Natl Acad. Sci. USA 112, 4897–4902 (2015).
Rabosky, D. L. & Lovette, I. J. Density-dependent diversification in North American wood warblers. Proc. R. Soc. B. Biol. Sci. 275, 2363–2371 (2008).
Friedman, S., Collyer, M., Price, S. & Wainwright, P. Divergent processes drive parallel evolution in marine and freshwater fishes. Syst. Biol. 4, syab080 (2021).
Jetz, W., Thomas, G., Joy, J., Hartmann, K. & Mooers, A. The global diversity of birds in space and time. Nature 491, 444–448 (2012).
Bjarnason, A. & Benson, R. A 3D geometric morphometric dataset quantifying skeletal variation in birds. MorphoMuseuM 7, e125 (2021).
Abourachid, A., Fabre, A. C., Cornette, R. & Höfling, E. Foot shape in arboreal birds: two morphological patterns for the same pincer‐like tool. J. Anatomy 231, 1–11 (2017).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2019); https://www.R-project.org/
Adams, D., Collyer, M., Kaliontzopoulou, A. & Baken, E. Geomorph: Software for geometric morphometric analyses. R package version 4.0.4. https://cran.r-project.org/package=geomorph (2022).
Collyer, M., Adams, D. & Collyer, M. M. Package ‘RRPP’. RRPP: Linear Model Evaluation with Randomized Residuals in a Permutation Procedure, R package version 1.1.2. https://cran.r-project.org/package=RRPP (2021).
Harmon, L. et al. Package ‘geiger’. R package version 2 (R Foundation for Statistical Computing, 2020).
Bookstein, F. L. in Advances in Morphometrics (eds Marcus, L. F. et al.) 131–151 (Springer, 1996).
Gunz, P. & Mitteroecker, P. Semilandmarks: a method for quantifying curves and surfaces. Hystrix 24, 103–109 (2013).
Collyer, M. L., Davis, M. A. & Adams, D. C. Making heads or tails of combined landmark configurations in geometric morphometric data. Evolution. Biol. 47, 193–205 (2020).
Ciampaglio, C. N., Kemp, M. & McShea, D. W. Detecting changes in morphospace occupation patterns in the fossil record: characterization and analysis of measures of disparity. Paleobiology 27, 695–715 (2001).
Wills, M. A., Briggs, D. E. & Fortey, R. A. Disparity as an evolutionary index: a comparison of Cambrian and Recent arthropods. Paleobiology 20, 93–130 (1994).
Clavel, J., Escarguel, G. & Merceron, G. mvMORPH: an R package for fitting multivariate evolutionary models to morphometric data. Meth. Ecol. Evol. 6, 1311–1319 (2015).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Revell, L. J. phytools: An R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2011).
Acknowledgements
For access to specimens, we thank J. White and J. Cooper (NHMUK), J. Hinshaw (UMMZ), M. Lowe and M. Brooke (UMZC), M. Carnall and E. Westwig (OUMNH), K. Zyskowski (YPM), and B. Marks and J. Bates (FMNH). For access to CT scanning facilities, we thank K. Smithson (Cambridge Biotomography Centre); T. Davies, B. Moon and L. Martin-Silverstone (University of Bristol); V. Fernandez (Natural History Museum); A. Neander and Z.-X. Luo (University of Chicago PaleoCT) and M. Friedman (University of Michigan). We thank E. Griffiths, S. Wright, S. Poindexter, A. Wolniewicz and S. Evers for segmenting digital bone models from the CT scan data. We are grateful to A. Martin-Serra, J. L. Cantalapiedra, F. Blanco, M. Fabbri, I. Menéndez, S.M. Nebreda, J. Clavel, E.M. Steell, J. Marugán-Lobón and C. Navalón for enlightening discussion on contents, narrative and analytical approaches. We thank L. Balsa Pascual and Ó. Sanisidro for discussion on design choices. This work was supported by the European Union’s Horizon 2020 research and innovation programme 2014–2018 under grant agreement no. 677774 (European Research Council Starting grant no. TEMPO). Grant no. 677774 applies to the work of G.N., A.B. and R.B.J.B. G.N. acknowledges support from UKRI Future Leaders Fellowship no. MR/S032177/1.
Author information
Authors and Affiliations
Contributions
G.N. and R.B.J.B. conceptualized the study and designed the analytical approach. G.N., A.B., E.G. and R.B.J.B. collected and curated the data. G.N. and R.B.J.B. undertook the formal analyses. G.N. and R.B.J.B. wrote the manuscript. G.N. made the figures. G.N., A.B., E.G. and R.B.J.B. edited the text. R.B.J.B. acquired the funding used for this research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Gavin Thomas and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
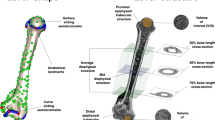
Extended Data Fig. 1 Infographic displaying the data and the methods’ pipeline used in this study.
A detailed account of the Material and Methods used in our study can be found in the main text.
Extended Data Fig. 2 Diagram showing how subclade disparity through time is calculated as test of early partitioning of morphospace, showing idealized contrasting patterns of morphological evolution.
a) Example time-calibrated phylogeny and points in time (cladogenesis events) for which b) subclade disparities values are calculated for a Brownian model of diffusive evolution and two alternative modes of morphological evolution. c) Corresponding patterns in morphospace exploration over time for two idealised patterns of morphological evolution: early partitioning of trait space (morphospace) in which morphological evolution is constrained at the level of subclades, and a non-partitioned pattern in which constraints are similar in the total group and within subclades.
Extended Data Fig. 3 Tests of early-partitioning of avian allometry-free shape and local shape variation of the individual elements in Aequorlithornithes, Inopinaves and Passeriformes.
See Extended Data Fig. 2 for a detailed account of the idealised patterns associated with each mode of evolution of disparity through time.
Extended Data Fig. 4 Deviations of group-specific disparities from the disparity expected if all birds were evolving under a uniform Brownian motion model of evolution for additional groups of water-based (Gruiformes, Anseriformes) and land-based (non-landbird terrestrial Neoaves, Galliformes) birds.
Boxplots summarize the distribution of delta disparities, calculated as empirical disparities minus disparities for the 200 simulated values for each of the three target clades. All partitions and data types are displayed, namely, the whole skeleton, three main skeletal regions and individual bones and the four different aspects of morphological variation in the three target lineages of birds. Arrows highlight the lineage/partition/data type which shows particularly high (up pointing arrows) or low (down pointing arrows) values. Values are normalised by interquartile range.
Extended Data Fig. 5 Subclade disparity–age plots for the whole skeleton and body regions for Aequorlitornithes (waterbirds).
X axes represent subclade disparities while y axes represent time in millions of years from the present time. Solid coloured lines represent mean empirical disparities through time, dashed lines represent mean BM-simulated disparities through time, shaded polygons display the space between 95% percentile and 5% percentiles of disparities through time for BM-simulated data. Colour code of individual subclade disparities (dots) as in Fig. 2.
Extended Data Fig. 6 Subclade disparity–age plots for the whole skeleton and body regions for Inopinaves (landbirds).
X axes represent subclade disparities while y axes represent time in millions of years from the present time. Solid coloured lines represent mean empirical disparities through time, dashed lines represent mean BM-simulated disparities through time, shaded polygons display the space between 95% percentile and 5% percentiles of disparities through time for BM-simulated data. Colour code of individual subclade disparities (dots) as in Fig. 2.
Extended Data Fig. 7 Subclade disparity–age plots for the whole skeleton and body regions for Passeriformes (passerines).
X axes represent subclade disparities while y axes represent time in millions of years from the present time. Solid coloured lines represent mean empirical disparities through time, dashed lines represent mean BM-simulated disparities through time, shaded polygons display the space between 95% percentile and 5% percentiles of disparities through time for BM-simulated data. Colour code of individual subclade disparities (dots) as in Fig. 2.
Extended Data Fig. 8 Subclade disparity–age plots for the individual elements for Aequorlitornithes (waterbirds).
X axes represent subclade disparities while y axes represent time in millions of years from the present time. Solid coloured lines represent mean empirical disparities through time, dashed lines represent mean BM-simulated disparities through time, shaded polygons display the space between 95% percentile and 5% percentiles of disparities through time for BM-simulated data. Colour code of individual subclade disparities (dots) as in Fig. 2.
Extended Data Fig. 9 Subclade disparity–age plots for the individual elements for Inopinaves (landbirds).
X axes represent subclade disparities while y axes represent time in millions of years from the present time. Solid coloured lines represent mean empirical disparities through time, dashed lines represent mean BM-simulated disparities through time, shaded polygons display the space between 95% percentile and 5% percentiles of disparities through time for BM-simulated data. Colour code of individual subclade disparities (dots) as in Fig. 2.
Extended Data Fig. 10 Subclade disparity–age plots for the individual elements for Passeriformes (passerines).
X axes represent subclade disparities while y axes represent time in millions of years from the present time. Solid coloured lines represent mean empirical disparities through time, dashed lines represent mean BM-simulated disparities through time, shaded polygons display the space between 95% percentile and 5% percentiles of disparities through time for BM-simulated data. Colour code of individual subclade disparities (dots) as in Fig. 2.
Supplementary information
Supplementary Information
This file contains Supplementary Figs. 1–37 and References.
Supplementary Table 1
List of specimens used in this study.
Supplementary Table 2
ANOVA tables used for proportions-normalizing individual elements.
Supplementary Table 3
Pairs of landmarks used to calculate maximum distances in each of the spatial axes.
Supplementary File 1
Bird_landmarks_Navalon_2022. Raw landmark coordinates for all specimens used in this study.
Supplementary File 2
Read_bird_landmarks_Navalon_2022. R code to read the raw landmarks coordinates in Supplementary File 1.
Supplementary File 3
Bird_processed_landmarks_Navalon_2022. Allometry-free and proportions free landmark coordinates for all specimens used in this study.
Supplementary File 4
Read_bird_processed_landmarks_Navalon_2022. R code to read the raw landmarks coordinates in Supplementary File 3.
Supplementary File 5
Combined phylogeny used in this study.
Supplementary File 6
Custom R functions used in this study.
Supplementary File 7
Custom R code used in this study.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Navalón, G., Bjarnason, A., Griffiths, E. et al. Environmental signal in the evolutionary diversification of bird skeletons. Nature 611, 306–311 (2022). https://doi.org/10.1038/s41586-022-05372-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05372-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.