Abstract
Severe COVID-19 is characterized by persistent lung inflammation, inflammatory cytokine production, viral RNA and a sustained interferon (IFN) response, all of which are recapitulated and required for pathology in the SARS-CoV-2-infected MISTRG6-hACE2 humanized mouse model of COVID-19, which has a human immune system1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20. Blocking either viral replication with remdesivir21,22,23 or the downstream IFN-stimulated cascade with anti-IFNAR2 antibodies in vivo in the chronic stages of disease attenuates the overactive immune inflammatory response, especially inflammatory macrophages. Here we show that SARS-CoV-2 infection and replication in lung-resident human macrophages is a critical driver of disease. In response to infection mediated by CD16 and ACE2 receptors, human macrophages activate inflammasomes, release interleukin 1 (IL-1) and IL-18, and undergo pyroptosis, thereby contributing to the hyperinflammatory state of the lungs. Inflammasome activation and the accompanying inflammatory response are necessary for lung inflammation, as inhibition of the NLRP3 inflammasome pathway reverses chronic lung pathology. Notably, this blockade of inflammasome activation leads to the release of infectious virus by the infected macrophages. Thus, inflammasomes oppose host infection by SARS-CoV-2 through the production of inflammatory cytokines and suicide by pyroptosis to prevent a productive viral cycle.
Similar content being viewed by others
Main
Acute SARS-CoV-2 infection resolves in most patients but becomes chronic and sometimes deadly in about 10–20% of patients1,2,3,4,5,6,7,14,15,16,20,24,25,26,27. Two hallmarks of severe COVID-19 are a sustained IFN response and viral RNA persisting for months1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,20,24,25,26,27,28. This chronicity is recapitulated in SARS-CoV-2-infected MISTRG6-hACE2 humanized mice19. High levels of IL-1β, IL-18 and lactate dehydrogenase (LDH) are correlated with COVID-19 severity in patients, suggesting a role for inflammasome activation and pyroptosis in pathology5,6,7,14,15,16,17,18,29. Here we show that human lung macrophages are infected by SARS-CoV-2. Replicating SARS-CoV-2 in these human macrophages activates inflammasomes and initiates an inflammatory cascade with a unique transcriptome, results in pyroptosis, and contributes to the downstream type-I IFN response. Blocking viral replication, the downstream IFN response or inflammasome activation in vivo during the chronic phase of the disease attenuates many aspects of the overactive immune inflammatory response (especially the inflammatory macrophage response) and disease.
Viral replication and the IFN response
Chronic interferon is associated with disease severity and impaired recovery in influenza infection30. To test whether a viral-RNA-dependent type-I IFN response was a driver of chronic disease, we treated SARS-CoV-2-infected MISTRG6-hACE2 mice with remdesivir21,22,23 and/or anti-IFNAR2 antibodies (Fig. 1a) to inhibit viral replication and/or the IFN response downstream of chronic infection, respectively. As a control, we used dexamethasone, which reverses many aspects of immunopathology in infected MISTRG6-hACE2 mice19 and in humans31. Although remdesivir and anti-IFNAR2 alone were partially therapeutic, combined therapy achieved more rapid weight recovery and suppression of the immune inflammatory response, especially macrophages, as effectively as dexamethasone (Fig. 1b–c and Extended Data Fig. 1a–f), suggesting that remdesivir and anti-IFNAR2 antibodies have a combinatorial effect in chronic infection.
a, Schematic of therapy: SARS-CoV-2-infected MISTRG6-hACE2 mice were treated with dexamethasone (Dex) and remdesivir (RDV) at 7, 8 and 9 d.p.i., and with anti-IFNAR2 antibodies at 7 and 11 d.p.i., and analysed at 14 or 28 d.p.i. i.p., intraperitoneal; i.v., intravenous; s.c., subcutaneous. b, Weight changes after infection. 28 d.p.i.: n = 5 (control infected), n = 4 (Dex; anti-IFNAR2 + RDV) and n = 3 (RDV; anti-IFNAR2) mice were examined over at least two experiments. Data are mean ± s.d. c, Human macrophages in the lungs. n = 10 (uninfected); 14 d.p.i.: n = 7 (control infected), n = 4 (Dex; RDV; anti-IFNAR; anti-IFNAR + RDV); 28 d.p.i.: n = 4 (control infected; Dex), n = 3 (RDV; anti-IFNAR) and n = 6 (anti-IFNAR + RDV) mice were examined over three experiments. Mean values are shown with data points. NS, not significant. d, Heat map of human genes suppressed by therapy in the lungs (log2[fold change] > 1; adjusted P with Bonferroni correction < 0.05). Differential expression was determined using DESeq2. Statistical analysis was performed using Wald tests. Transformed (minimum–maximum) normalized counts of duplicates. Hierarchical clustering (1 − Pearson). e, t-Distributed stochastic neighbour embedding (t-SNE) plot of human immune cells from uninfected or infected lungs (28 d.p.i.). Pooled duplicates. Cluster marker genes were identified using Wilcoxon rank-sum tests (Extended Data Fig. 2e). n = 3,655 (uninfected) and n = 3,776 (28 d.p.i.) cells analysed. NK, natural killer; Treg, T regulatory. f, t-SNE plots highlighting differentially abundant (DA) human immune cell populations identified by DA-seq61. Top, distribution/enrichment of DA populations. Bottom, DA clusters. g, t-SNE plots of human monocyte/macrophage clusters from 4 d.p.i., 14 d.p.i. and 28 d.p.i., and uninfected lungs. Left, d.p.i. Right, clusters. Different conditions were integrated as described in the Methods (ref. 62). Marker genes were identified using Wilcoxon rank-sum tests (Extended Data Fig. 3a,b). P values were adjusted with Bonferroni correction. n = 438 (uninfected), n = 336 (4 d.p.i.), n = 793 (14 d.p.i.) and n = 1,368 (28 d.p.i.) cells analysed. h, Heat map visualizing the response to the combined therapy based on differentially expressed genes associated with human monocytes and macrophages. Transformed (minimum–maximum) normalized expression of duplicates. Hierarchical clustering (1 − Pearson). i, Representative haematoxylin and eosin (H&E) staining and box plot of histopathological scores. n = 6 (uninfected), n = 7 (control infected) and n = 4 (anti-IFNAR2 + RDV) mice were examined over three experiments. The centre line shows the median value, the box limits show the 25th–75th percentiles, and the whiskers show the lowest (minimum) to the highest (maximum) value. Scale bars, 1 mm (first and third), 0.1 mm (second and fourth). For b, c and i, statistical analysis was performed using unpaired two-tailed t-tests. The data associated with dexamethasone used here as a control were reported previously19.
We assessed the effect of therapeutics on the lung transcriptome. Both dexamethasone and the combined therapy reversed overactive immune transcripts to the levels in uninfected animals (Fig. 1d, Extended Data Fig. 2a,b and Supplementary Table 1). The reduced transcripts were enriched for chemokine and cytokine networks (CXCL10, CXCL8, CCL2), inflammatory (TLR7, NLRP3, CASP1) and anti-viral (MPO, OAS1, OAS2) response, and interferon-stimulated genes (ISGs) (IFITM3, IFITM2, IRF7) (Supplementary Table 1 and Extended Data Fig. 2c,d), emphasizing the central role of IFN signalling and inflammatory cytokine-chemokines in chronic COVID-19. Comparison of single-cell transcriptomes of human immune cells from infected mice with their uninfected counterparts (Fig. 1e–g and Extended Data Fig. 2e) showed tissue-resident macrophages, such as alveolar macrophages (AMs), activated at the peak of infection, followed by an inflammatory response with infiltrating monocytes and monocyte-derived macrophages (MDM)(Fig. 1g, Extended Data Fig. 3a–c and Supplementary Table 2). As macrophages differentiated, they maintained their inflammatory signature and activated status throughout infection (Extended Data Fig. 3a–c and Supplementary Table 2). All macrophage subsets were enriched for ISGs at all timepoints (Extended Data Fig. 3c). These ISGs were suppressed after combination therapy with anti-IFNAR2 antibodies and remdesivir (Fig. 1h, Extended Data Fig. 4 and Supplementary Table 3). However, key anti-viral responses such as IFNG primarily produced by cytotoxic T cells were spared (Extended Data Fig. 5a), highlighting the selective effects of combined anti-IFNAR2–remdesivir therapy on the pathology of chronic COVID-19. Consistent with the fibrosis seen both in patients32,33,34,35,36 and humanized mice19, alveolar self-renewal and differentiation programs were inhibited, resulting in the accumulation of the pre-alveolar type 1 transitional cell state program in pneumocytes7,37,38,39. This state was reversed in infected MISTRG6-hACE2 mice by anti-IFNAR2–remdesivir combination therapy, restoring self-renewal and differentiation programs (Extended Data Fig. 5b). Overall, reducing chronic inflammation enhanced lung tissue recovery and prevented transition to fibrosis seen in humanized mice19 and humans32,33,34,35,36 (Fig. 1i and Extended Data Fig. 5c).
SARS-CoV-2 replicates in human macrophages
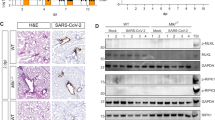
To determine the cellular source of persistent viral RNA and replication, we measured genomic RNA (gRNA) and subgenomic viral RNA (sgRNA)40 in lung tissue or in sorted lung epithelial cells or human immune cells from infected MISTRG6-hACE2 mice (Extended Data Fig. 6a–d). Notably, epithelial cells and human immune cells had similar levels of viral RNA (Extended Data Fig. 6d). Although gRNA was abundant, we could not discern sgRNA in either cell type. We tracked infected cells in MISTRG6-hACE2 mice using a reporter strain of virus—SARS-CoV-2-mNG41, which encodes the fluorescent protein mNG in infected cells. On the basis of this assay, most epithelial cells in the bronchioalveolar lavage (BAL) but only a few total lung epithelial cells were infected with SARS-CoV-2 (Extended Data Fig. 6e). Notably, human macrophages were strongly mNG positive throughout disease (Fig. 2a and Extended Data Fig. 6f,g). No mouse immune cells expressed mNG (Fig. 2a and Extended Data Fig. 6f). To address whether the SARS-CoV-2 viral RNA replicates in these cells or is acquired by phagocytosis, we measured the mNG signal in human macrophages from infected MISTRG6 mice untransduced with hACE2. In these mice, epithelial cells were not infected or infected poorly with SARS-CoV-219,42 (Extended Data Fig. 6h). However, these mice had similar levels of mNG+ human macrophages to AAV-hACE2 mice, suggesting that viral uptake by macrophages is independent of infected epithelial cells (Extended Data Fig. 6i). To determine whether SARS-CoV-2 replicates in human macrophages, we quantified gRNA and sgRNA40 in mNG+ versus mNG− epithelial or human immune cells at 4 days post-infection (d.p.i.) or 14 d.p.i. (Extended Data Fig. 6j). Only mNG+, and not mNG−, epithelial and immune cells had sgRNA (Fig. 2b). Second, we stained for double-stranded RNA (dsRNA), a diagnostic indicator of viral replication (Fig. 2c). As expected, mNG and dsRNA were detected/colocalized in human macrophages (Fig. 2c and Extended Data Fig. 7a). Third, we detected viral RNA-dependent RNA polymerase (RdRp) in human macrophages, which colocalized with a viral spike protein supporting specificity (Fig. 2d, Extended Data Fig. 7b and 8a–c). Viral RdRp and spike were also present in the human macrophages of autopsies of human lungs with SARS-CoV-2 pneumonia (Extended Data Fig. 9). Thus, the mouse model observations reflected the human disease. Remdesivir reduced the mNG signal and viral titres by the same amount in infected MISTRG6-hACE2 mice (Fig. 2e and Extended Data Fig. 10a). Thus, SARS CoV-2 appeared to replicate in human immune cells.
a, Representative flow cytometry plots and frequencies of mNG+ human (CD68+) or mouse (F4/80+) lung macrophages in SARS-CoV-2-mNG infected MISTRG6-hACE2 mice. n = 7 (human) and n = 6 (mouse) mice over at least three experiments. b, Quantification of gRNA and sgRNA (E gene)40,63 in sorted mNG+ or mNG− epithelial cells or human immune cells. n = 3 mice over 2 experiments. Mean values are shown with data points. ND, not detected. c, Representative fluorescence microscopy images of dsRNA (rJ2), CD68 and DAPI staining in fixed lung tissues from SARS-CoV-2-infected MISTRG6-hACE2 mice. Representative of n = 5 mice examined over 3 experiments. The yellow rectangle indicates the region magnified on the right. The yellow arrow indicates colocalization of CD68 with dsRNA. Pseudocolours were assigned. d, Representative fluorescence microscopy images of RdRp, CD68 and DAPI staining in fixed lung tissues from SARS-CoV-2-infected MISTRG6-hACE2 mice. Representative of n = 5 mice examined over 3 experiments. The yellow arrows indicate colocalization of human CD68 with dsRNA. The yellow rectangle indicates the region magnified above. Pseudocolours were assigned. For c and d, scale bars, 50 μm. e, The frequencies of mNG+ human immune cells in remdesivir-treated (1–3 d.p.i.) or control MISTRG6-hACE2 mice infected with SARS-CoV-2-mNG. n = 6 mice were examined over 3 experiments. Mean values are shown with data points. f, The frequencies of mNG+ human immune cells after ACE2 blockade (1–3 d.p.i.) in MISTRG6 (no AAV) mice infected with SARS-CoV-2-mNG. n = 5 (control infected) and n = 6 (anti-ACE2 treated) mice were examined over 2 experiments. Mean values are shown with data points. g, Representative flow cytometry plots and frequencies of mNG+ macrophages in infected MISTRG6-hACE2 mice treated with monoclonal antibodies (mAbs)19,45,64 at 35 h.p.i. n = 7 (control infected) and n = 4 (treated) mice were examined over 2 experiments. Mean values are shown with data points. h, The frequencies of mNG+ human immune cells in MISTRG6-hACE2 mice after CD16 blockade (2 d.p.i.). n = 6 mice were examined over 3 experiments. Mean values are shown with data points. Statistical analysis was performed using unpaired (a and g) and paired (e, f and h) two-tailed t-tests.
SARS-CoV-2 infects through ACE2 and CD16
The ACE2 receptor used by SARS-CoV-2 to infect the lung epithelium can be expressed in macrophages43. We measured ACE2 expression using flow cytometry and immunofluorescence staining in mouse epithelial cells and human lung macrophages (Extended Data Fig. 10b–f). Human lung macrophages of both MISTRG6 and MISTRG6-hACE2 mice, as well as the epithelial cells of only MISTRG6-hACE2 mice, expressed human ACE2 (Extended Data Fig. 10b–e). Interestingly, ACE2 expression was higher in both infected (mNG+) human macrophages and epithelial cells (Extended Data Fig. 10b–e). We treated SARS-CoV-2-infected MISTRG6 mice with a blocking antibody against human ACE2. In these mice, SARS-CoV-2 infects epithelial cells poorly19,42, as the mice did not receive AAV-ACE2 and only human macrophages express human ACE2 (Extended Data Fig. 6h). ACE2 blockade significantly diminished infected human macrophages (Fig. 2f), suggesting that ACE2 can mediate viral entry in human lung macrophages.
Antibodies can also mediate viral uptake by macrophages (for example, Dengue virus44). To test the role of antibody-mediated viral entry to macrophages, we treated infected mice with monoclonal antibodies45 against SARS-CoV-2 spike protein early (35 h post-infection (h.p.i.)) when the effects of endogenous antibodies are minimal or late (7 d.p.i.) (Extended Data Fig. 10g). Indeed, monoclonal antibody treatments increased infected lung macrophages (Fig. 2g and Extended Data Fig. 10h). Immune cells express a wide range of surface Fcγ receptors (FcγRs) that interact with the Fc moiety of antibodies. These interactions lead to multiple protective or pathological effector functions44,46. The severity of COVID-19 correlates with high serum IgG levels and specific IgG-Fc structures and interactions47,48,49. One such Fc interaction is mediated by CD16, which is expressed at high levels in mNG+ macrophages. We treated mice early (2 d.p.i., low antibody levels) as a proof of concept, or late (7 d.p.i. and 11 d.p.i., high antibody levels) as a possible therapeutic with anti-CD16 antibodies. Anti-viral antibody levels in the lung tissue were sufficient to mediate viral uptake and were positively correlated with mNG levels at 4 d.p.i. (Extended Data Fig. 10g,i). When dosing was optimized, CD16 blockade did not alter the distribution of macrophages, although it resulted in significantly fewer infected human macrophages at both timepoints (Fig. 2h and Extended Data Fig. 10j).
To elucidate whether viral replication products are the result of bona fide infection, we cultured human bone-marrow-derived macrophages (BMDMs, differentiated from bone-marrow cells of immune-reconstituted MISTRG6 mice) with SARS-CoV-2 in vitro. Indeed, SARS-CoV-2 was taken up by BMDMs and replicated in these cells, as measured by mNG signal (Extended Data Fig. 10k) and high levels of sgRNA (Extended Data Fig. 10l). This was true for multiple types of macrophage (Extended Data Fig. 10m). As in vivo, in vitro macrophage infection enhanced by antibodies (convalescent plasma or monoclonal antibodies) was reduced by CD16, ACE2 or RdRp blockade (Extended Data Fig. 10n,o). sgRNA levels in these macrophages were also reduced by these treatments (Extended Data Fig. 10l), further supporting a role for both ACE2 and CD16 in viral uptake and RdRp in viral replication. SARS-CoV-2 infection in human macrophages was not productive or produced very little as indicated by undetectable infectious virus, titred in culture, from sorted immune cells from infected mice at 4 d.p.i. and in vitro infected macrophages at 48 h.p.i. (Extended Data Fig. 10p–r).
The transcriptome of infected macrophages
We next determined the consequences of infection of human macrophages by SARS-CoV-2. Infected macrophages preferentially produced CXCL10, a chemokine that recruits many types of immune cells (Fig. 3a), but not TNF. Similar to mNG positivity, CXCL10 production by human macrophages was also enhanced by antibodies and inhibited by remdesivir, also reflected in serum levels and in vitro (Fig. 3b,c and Extended Data Fig. 11a–c). Thus, we used CXCL10 as a proxy for SARS-CoV-2-infected macrophages and identified a unique transcriptional signature enriched for genes encoded by tissue-resident macrophages, in particular AMs50 (APOC1, MRC1, ALOX5AP, FABP5, INHBA), chemokines of interstitial macrophages (CCL18, CCL3, CCL7, CCL8, CCL20, CXCL8), inflammatory cytokines (IL1A, IL18, IL27), complement genes (C1QA, C1QB) and ISGs (ISG20, IFI27) (Fig. 3d, Extended Data Figs. 11d,e and 12a,b, and Supplementary Table 4). Further flow cytometry characterization of mNG+ cells also confirmed enrichment for CD16+ AMs, which produced more CXCL10 (Fig. 3e and Extended Data Fig. 12c). Consistent with our findings, CD14highCD16high cells and AMs enriched with viral RNA in autopsies of the lungs of patients with COVID-197,20 also had distinct transcriptomes that were largely recapitulated in what we construe as CXCL10-associated genes (CXCL11, CCL18, CCL8, ISG15, CD83). Interestingly, this strong network of CXCL10-specific gene signature was no longer restricted to AMs later in infection as different macrophage subsets continuously differentiate, as evidenced by the high IL7R expression of developing lung macrophages50 (CXCL10+ and AMs) at all timepoints (Figs. 1g and 3d and Extended Data Fig. 12d).
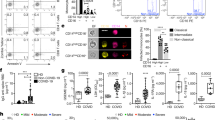
a, CXCL10+ or TNF+ human macrophages (MΦ). Representative of n = 6 mice over 3 experiments. b, CXCL10+ lung macrophage frequencies after monoclonal antibody or remdesivir therapy. n = 9 (control infected), n = 4 (monoclonal antibodies) and n = 6 (RDV) mice over 2 experiments. Mean values are shown with data points. c, Serum CXCL10 levels after monoclonal antibody or remdesivir therapy. Mean values are shown with data points. n = 4 (uninfected; monoclonal antibodies); n = 7 (control infected) and n = 3 (RDV) mice examined over 2 experiments. d, Correlation (Pearson and Spearman) of each gene with CXCL10, TNF or TLR7 in human lung monocytes and macrophages. k-means clustering. P values were calculated based on a t-distribution with length(x) − 2 d.f. or algorithm AS 89 with exact = TRUE; two-tailed. e, Representative plots and AM frequencies within mNG+ or mNG− macrophages. n = 8 mice examined over 4 experiments. f, ASC speck visualization/quantification and colocalization with active caspase-1 (FLICA) in mNG+ or mNG− human immune cells from MISTRG6-hACE2 mouse lungs. Cells were sorted as shown in Extended Data Fig. 6j. n = 1,000 cells were analysed per condition. ASC+ specks: n = 3 (A, 4 d.p.i.), n = 5 (B–D, 4 d.p.i.) and n = 3 mice (14 d.p.i.); FLICA: n = 3 mice examined over at least 2 experiments. Data are mean ± s.d. with data points. BF, bright field. NS, not significant. g, ASC speck visualization/quantification and colocalization with NLRP3 oligomerization in sorted mNG+ or mNG− human lung immune cells. n = 1,000 cells were analysed per condition. n = 3 mice over 2 experiments. Data are mean ± s.d. with data points. For f and g, scale bars 7 μm. BF, bright field. h, Serum IL-18, IL-1RA and GSDMD levels. IL-18: n = 4 mice were examined over 2 experiments; IL-1RA: n = 5 (uninfected), n = 7 (4 d.p.i.) and n = 4 (14 d.p.i.) mice were examined over 3 experiments; GSDMD: n = 4 (uninfected; 4 d.p.i.) and n = 5 (14 d.p.i.) mice over 3 experiments. Mean values are shown with data points. P < 0.0001 represents P = 3.32 × 10−7. i, Serum IL-18 and IL-1RA levels in mice treated with CD16-blocking or CD16-depleting antibodies or remdesivir. IL-18: n = 4 (uninfected; CD16 blocking; CD16 depleting), n = 7 (control infected) and n = 3 (RDV); IL-1RA: n = 5 (uninfected), n = 7 (control infected), n = 6 (RDV), n = 4 (CD16 blocking; CD16 depleting) mice were examined over at least 2 experiments. Mean values are shown with data points. P < 0.0001; uninfected, P = 3.28 × 10−5; CD16 depletion, P = 7.92 × 10−7. For b, c and f–i, statistical analysis was performed using unpaired two-tailed t-tests.
SARS-CoV-2 activates inflammasomes
Morphological analysis of sorted mNG+ cells revealed the appearance of membrane bubbles—a characteristic of pyroptosis—prompting us to investigate inflammasome activation as part of the inflammatory cascade initiated by infection. Inflammasomes are dynamic multiprotein complexes in which specific NOD-like receptors and adaptor molecules are assembled to activate caspases—the central effector proteins. We sorted mNG+ and mNG− human immune cells, mNG+ epithelial cells and mouse immune cells (Extended Data Fig. 6j), and assayed for sensors, adaptors and effectors of the inflammasome pathway. First, focusing on adaptor molecule apoptosis-associated speck-like protein containing a CARD (ASC) as the common adaptor molecule with an essential role in inflammasome assembly and activation, we found that infected (mNG+) human cells exclusively showed substantial inflammasome activation, quantified by ASC speck formation (Fig.3f and Extended Data Fig. 13a,b). ASC specks co-localized with both NLRP3 and active caspase-1 (visualized by fluorochrome-labelled inhibitor of caspases assay (FLICA)) (Fig. 3f,g and Extended Data Fig. 13a–d). Inflammasome activation in infected human macrophages was sustained during disease (4–14 d.p.i.; Fig. 3f,g and Extended Data Fig. 13c).
Once inflammasome complexes are formed, active caspase-1 cleaves and proteolytically activates the pro-inflammatory IL-1-family cytokines IL-1β and IL-18, which are typically elevated and characteristic of severe COVID-19 in patients. IL-18 levels in the blood and lungs were significantly elevated in SARS-CoV-2-infected mice and correlated well with the proportions of infected (mNG+) macrophages (Fig. 3h and Extended Data Fig. 13e). Although IL-1β levels in serum in vivo were not detectable, we measured IL-1RA. This specific receptor antagonist, induced by IL-1β, served as a proxy of IL-1β and it paralleled enhanced IL-18 levels and correlated with mNG+ cells (Fig. 3h and Extended Data Fig. 13f).
Finally, we assayed for pyroptosis by detecting LDH and gasdermin D (GSDMD) in the serum. GSDMD, which is a substrate of active caspase-1 and a pore-forming executer of pyroptosis, and LDH, released by pyroptosis, were particularly enriched in the serum of infected mice at the late timepoints (14 d.p.i.; Fig.3h and Extended Data Fig. 13g), further supporting continuous inflammasome activation during infection. Moreover, infected lung macrophages showed higher incorporation of a small fixable dye (Zombie Aqua) that enters dying cells with a compromised cell membrane, consistent with the pore-forming function of GSDMD and pyroptosis (Extended Data Fig. 13h).
All aspects of inflammasome activation were also recapitulated in vitro when BMDMs were infected in vitro with SARS-CoV-2. Active caspase-1 in infected BMDMs, which was dependent on viral replication, was inhibited by remdesivir (Extended Data Fig. 13i). High levels of the inflammasome products IL-18, IL-1β and IL-1RA (in response to IL-1β) and two measures of pyroptosis, GSDMD and LDH, that were detected at high levels in the supernatants of infected BMDMs were also inhibited by remdesivir (Extended Data Fig. 13j–n). In vitro infected cells also had higher incorporation of Zombie Aqua, consistent with pyroptosis (Extended Data Fig. 13o).
Infection affects the macrophage response
To determine the role of viral infection on the inflammatory macrophage response, we first blocked viral entry and replication in vivo and measured inflammatory cytokines and chemokines. Blocking viral entry (CD16 or ACE2 blockade) or inhibiting viral replication (remdesivir) all reduced IL-18, IL-1RA and CXCL10 levels, paralleling mNG levels (Fig. 3i and Extended Data Fig. 14a–e). Depletion of CD16+ cells in vivo (Extended Data Fig. 14f,g) resulted in the complete loss of IL-18 and IL-1RA in the serum, consistent with the concept that viral replication and inflammasome activation occurred mainly in myeloid cells (Fig. 3i). By contrast, monoclonal antibodies promoting viral infection in human macrophages (Fig. 2g and Extended Data Fig. 10h) enhanced systemic IL-18, IL-1RA and CXCL10, particularly during early disease (Extended Data Fig. 14h–k). Nonetheless, despite changes in the levels of the inflammatory cytokines and chemokines, neither monoclonal antibodies nor CD16 blockade affected lung pathology, potentially owing to the conflicting role of these antibodies on viral titres versus inflammation (Extended Data Fig. 14l,m). Consistent with the in vivo studies, IL-18, IL-1β, IL-1RA and CXCL10 levels were also reduced in the supernatants of in vitro infected BMDMs after CD16, ACE2 or RdRp inhibition, again paralleling the reduced viral replication inferred from mNG levels (Extended Data Figs. 10o and 14n–q).
Inflammasome inhibition in COVID-19
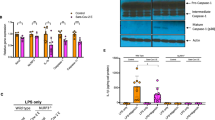
Finally, to assess the causal role of NLRP3 and caspase-1 activation in inflammasome-mediated inflammation and disease, we treated mice with caspase-1 and NLRP3 inhibitors (Fig. 4a). As expected, the proportion of infected cells did not diminish, but the inflammatory profile of these cells and other lung macrophages was considerably attenuated (Fig. 4b,c). In inhibitor-treated mice, mNG+ cells produced less CXCL10, which was also reflected in reduced serum levels (Fig. 4c and Extended Data Fig. 15a–c). Lung macrophages (mNG−) also produced less TNF (Fig. 4c and Extended Data Fig. 15a). Overall, inhibitor-treated mice had lower levels of caspase-1 activation and lower levels of IL-18, IL-1RA and GSDMD (Fig. 4d–g). The cumulative decrease in proinflammatory cytokines and chemokines after inflammasome inhibition reversed the immune-pathological state of the lungs, measured by scoring lung histopathology (Fig. 4h). Inflammasome inhibition reduced immune cell infiltration and enhanced tissue recovery to homeostasis in the lungs, despite persistently high levels of mNG+ human immune cells in the lungs.
a, Schematic of inflammasome inhibition in vivo. SARS-CoV-2-infected MISTRG6-hACE2 mice were treated with caspase-1 or NLRP3 inhibitors at 6–12 d.p.i. b, The frequencies of mNG+ human immune cells after inflammasome inhibition. n = 5 (control infected), n = 6 (caspase-1 inhibitor) and n = 4 (NLRP3 inhibitor) mice were examined over at least 2 experiments. Mean values are shown with data points. c, The frequencies of CXCL10+ or TNF+ human lung macrophages after inflammasome inhibition. n = 5 (control infected), n = 5 (caspase-1 inhibitor), n = 4 (NLRP3 inhibitor) mice were examined over at least 2 experiments. Mean values are shown with data points. d, Quantification of active caspase-1 in mNG+ human macrophages after inflammasome inhibition. n = 5 (control infected), n = 5 (caspase-1 inhibitor), n = 4 (NLRP3 inhibitor) mice examined over at least 2 experiments. Mean values are shown with data points. e, Serum human IL-18 levels after inflammasome inhibition. n = 4 (control; NLRP3), n = 5 (caspase-1) mice were examined over 2 experiments. Mean values are shown with data points. P < 0.0001 represents P = 1.00114 × 10−5. f, Human IL-1RA levels in lung homogenates after inflammasome inhibition. n = 4 mice were examined over 2 experiments. Mean values are shown with data points. g, Serum GSDMD levels after inflammasome inhibition. n = 5 (control infected) and n = 4 (caspase-1 inhibitor; NLRP3 inhibitor) mice were examined over 2 experiments. Mean values are shown with data points. h, Representative H&E staining (left) and histopathological scores (right) after inflammasome inhibition. n = 4 (control infected), n = 6 (caspase-1 inhibitor), n = 4 (NLRP3 inhibitor) independent mice over at least 2 experiments. For the box plots, the centre line shows the median value, the box limits show the 25th–75th percentiles, and the whiskers show the lowest (minimum) to the highest (maximum) value. Scale bars, 0.2 mm. i, Viral titres from the supernatants of BMDMs infected with SARS-CoV-2-mNG in vitro and treated with caspase-1 or NLRP3 inhibitors. n = 13 (control infected), n = 8 (caspase-1 inhibitor) and n = 5 (NLRP3 inhibitor) independent data points collected over 3 experiments. Mean values are shown with data points. Statistical analysis was performed using paired (b) and unpaired (c–i) two-tailed t-tests.
Caspase-1 and NLRP3 inhibitors blocked inflammasomes in vitro but did not affect macrophage infection, measured as mNG+ macrophage frequency, and reduced the inflammatory response to infection (Extended Data Fig. 15d). All parameters of inflammasome activation, including active caspase-1, IL-1β, IL-18, GSDMD and LDH, were significantly reduced after caspase-1 and NLRP3 inhibition in vitro (Extended Data Fig. 15e–i). Consistent with decreased pyroptosis, inflammasome blockade significantly reduced Zombie Aqua+ cells (Extended Data Fig. 15j). As seen in vivo, in vitro infected bone marrow macrophages produced less CXCL10 and IL-1RA (Extended Data Fig. 15k,l).
Finally, we tested whether inflammasome activation translated to any changes in the levels of infectious virus. We therefore first measured viral titres in the lungs of mice treated with caspase-1 inhibitor. Indeed, mice treated with caspase-1 inhibitor had a higher viral load at 14 d.p.i. in vivo (Extended Data Fig. 15m). Given that a reduced inflammatory response could result in deficient viral clearance, we infected macrophages in vitro and treated them with caspase-1 or NLRP3 inhibitors to test the direct effect of inflammasome activation on infectious virus. An analysis of the supernatants of these cultures showed that inhibitor-treated cells produce substantially higher amounts of virus compared with the uninhibited controls (Fig. 4i and Extended Data Fig. 15n). Thus, the activation of inflammasomes in infected macrophages has two protective functions—it attenuates virus production and signals infection to the immune system by releasing inflammatory cues to recruit and activate more immune cells at the site of infection.
Overall, these findings suggest that infection of macrophages by SARS-CoV-2 activates inflammasomes and drives pyroptosis. Pyroptosis interrupts the viral replication cycle and prevents viral amplification; in parallel it releases immune cell activators and recruiters. Viral RNA/pathogen-associated molecular patterns and proinflammatory cytokines released from these cells probably shape the hyperinflammatory macrophage response sustained by infiltrating monocytes and MDMs and drive immunopathology.
Discussion
The MISTRG6 COVID-19 model faithfully reflects many of the chronic immunoinflammatory features of the human disease, such as chronic viral RNA, IFN response and the inflammatory state in macrophages19. Overall, our mechanistic study of this model defines a cascade of events that initiates with the infection of lung macrophages generating replicative intermediates and products, including RdRp, dsRNA and sgRNA. SARS-CoV-2 replication activates an inflammatory program with activation of inflammasomes, production and release of inflammatory cytokines and chemokines, and pyroptosis. We established all steps of inflammasome activation by visualizing ASC oligomerization, colocalization with active caspase-1 and NLRP3, maturation of inflammasome-mediated cytokines IL-1β and IL-18, and pyroptosis assayed by GSDMD and LDH release. Inhibitors of both caspase-1 and NLRP3 blocked the downstream aspects of inflammasome activation and the inflammatory cascade both in vivo and in vitro. Notably, targeting inflammasome-mediated hyperinflammation, or the combined targeting of viral replication and the downstream interferon response in the chronic phase of the disease, prevented immunopathology associated with chronic SARS-CoV-2 infection in vivo.
In contrast to epithelial cells, infected macrophages produce little virus. However, inhibition of the inflammasome pathway led to a substantial increase in infectious virus produced by infected macrophages, although the degree to which these macrophages contribute, if at all, to high titres of virus production is unclear. Notably, inflammasome activation denies the virus the opportunity to replicate productively in these sentinel immune cells and, instead, broadcasts inflammatory signals that inform the immune system of the infection. Although this is potentially beneficial, excessive inflammation occurring through this mechanism coupled with the dysregulated interferon response may be the key factor leading to the excessive inflammation that typifies chronic COVID-192,5,51,52,53,54. Indeed, attenuation of the inflammasome in vivo blocks the inflammatory infiltrates in the lungs of infected mice in vivo. We speculate that, by contrast, an early interferon response, as may occur in the majority of patients who rapidly clear infection, as well as in the acute mouse models of infection in which human immune cells that can be infected are not present, leads to viral elimination before this inflammatory chain reaction can occur.
Viral RNA and particles can be detected by a variety of innate immune sensors. The inflammasome sensor NLRP3 is both upregulated and activated by replicating SARS-CoV-2. The NLRP3 inflammasome can directly sense viral replication/RNA or can rely on other viral RNA sensors such as MDA5 or RIG-I55,56,57. The loss of IL-18/IL-1β production after remdesivir treatment in our studies strongly suggests that viral replication is involved. Recent reports have also identified a possible role for NLRP3-driven inflammasome activation in SARS-CoV-2 infected myeloid cells in post-mortem tissue samples and PBMCs58. Although many candidates have been proposed (lytic cell death after infection, N protein59, ORF3A60), the exact mechanism of NLRP3 activation is still poorly understood29. Activation of other NOD-like receptors may also contribute to the process, as inhibition of caspase-1 was stronger than NLRP3 alone. Finally, there may be other mechanisms that enhance SARS-CoV-2 infection or the downstream inflammatory response in human macrophages that are unexplored here.
A role for inflammasome-driven hyperinflammation in COVID-19 pathophysiology in patients is now recognized5,6,7,14,15,16,17,18. Targeting inflammasome pathways in patients may provide alternative therapeutic options for resolving chronicity in COVID-19. However, the increased virus production seen after inflammasome blockade could pose a significant risk to the benefit of wholesale inhibition of the pathway. The findings from our study and its implications provide alternative therapeutic options to be investigated in the clinic and may guide new therapeutic developments and prompt clinical trials to investigate combinatorial therapies that target viral RNA, inflammasome activation or its products and sustained IFN response.
Methods
Mice
MISTRG6 mice were generated by the R. Flavell laboratory by combining mice generated by this laboratory, the laboratory of M. Manz and Regeneron Pharmaceuticals based on the Rag2−/−IL2rg−/− 129xBalb/c background supplemented with genes encoding human M-CSF, IL-3, SIRPα, thrombopoietin, GM-CSF and IL-6 knocked into their respective mouse loci65,66. MISTRG6 mice are deposited at Jackson Laboratories and have been made available to academic, non-profit and governmental institutions under a Yale-Regeneron material transfer agreement (already approved and agreed to by all parties). Instructions on obtaining the material transfer agreement for this mouse strain will be available along with strain information and on request. The CD1 strain of mice acquired from Charles River Laboratories was used for cross-fostering of MISTRG6 pups after birth to stabilize healthy microbiota. All mice were maintained under specific-pathogen free conditions in our animal facilities (biosafety level 1, 2 or 3) under our Animal Studies Committee-approved protocol. Unconstituted MISTRG6 mice were maintained with cycling treatment with enrofloxacin in the drinking water (Baytril, 0.27 mg ml−1). All animal experimentation was performed in compliance with Yale Institutional Animal Care and Use Committee protocols. For SARS-CoV-2-infected mice, all procedures were performed in a biosafety level 3 (BSL-3) facility with approval from the Yale Institutional Animal Care and Use Committee and Yale Environmental Health and Safety.
Transplantation of human CD34+ haematopoietic progenitor cells into mice
Fetal liver samples were cut into small fragments, treated for 45 min at 37 °C with collagenase D (Roche, 200 μg ml−1), and prepared into a cell suspension. Human CD34+ cells were purified by performing density-gradient centrifugation (Lymphocyte Separation Medium, MP Biomedicals), followed by positive immunomagnetic selection using the EasySep Human CD34 Positive Selection Kit (StemCell). For intrahepatic engraftment, newborn 1–3-day-old pups were injected with 20,000 fetal liver CD34+ cells in 20 μl of PBS into the liver with a 22-gauge needle (Hamilton Company). All use of human materials was approved by the Yale University Human Investigation Committee. At least three fetal liver samples (purchased) were used to account for variability and at least 8–10 gender-mixed mice were transplanted and analysed to account for reproducibility. Humanized mice that had lower than 30% humanization (percentage of human immune cells out of total (mouse and human combined) immune cells) in blood were excluded from the study or data collection.
AAV-hACE2 administration
AAV9 encoding hACE219,67 was purchased from Vector Biolabs (AAV9-CMV-hACE2). Animals were anaesthetized using isoflurane. The rostral neck was shaved and disinfected. A 5 mm incision was made, and the trachea was visualized. Using a 32-gauge insulin syringe, a 50 µl injection dose of 1011 genomic copies per ml of AAV-CMV-hACE2 was injected into the trachea. The incision was closed with 4–0 Vicryl suture and/or 3M Vetbond tissue adhesive. After administration of analgesic, the animals were placed into a heated cage until full recovery. The mice were then moved to BSL-3 facilities for acclimatization.
In vivo SARS-CoV-2 infection
The SARS-CoV-2 isolate USA-WA1/2020 was obtained from BEI reagent repository. SARS-CoV-2 mNG was obtained from P. Y. Shi41. All infection experiments were performed in a BSL-3 facility, licensed by the State of Connecticut and Yale University. Mice were anaesthetized using 20% (v/v) isoflurane diluted in propylene glycol. Using a pipette, 50 µl of SARS-CoV-2-WA1 or SARS-CoV-2-mNG (1–3 × 106 plaque-forming units (PFU)) was delivered intranasally.
Therapeutics
SARS-CoV-2-infected MISTRG6-hACE2 were treated intraperitoneally daily with dexamethasone at 10 mg kg−1 for 3 days starting at 7 d.p.i. Mice were treated subcutaneously with remdesivir at 25 mg kg−1 dosing as has been previously described22 for 3 consecutive days starting at 7 d.p.i. (Fig. 1) or 1 d.p.i. (Fig. 2; for human macrophage infection studies mice were treated twice, daily). Mice were treated with anti-IFNAR2 antibodies at 1.5 mg kg−1 dosing at 7 and 11 d.p.i. Weight changes post-infection were plotted as the percentage change compared with preinfection weight.
Infected MISTRG6-hACE2 mice were treated with two different clones of anti-human CD16 antibodies. For CD16 blockade experiments, mice were treated with anti-CD16 (Abcam, SP175) antibodies early and late. For early CD16 blockade studies, the mice were treated with anti-CD16 antibodies at 2 d.p.i. with a single dose (20 µg per mouse) and euthanized at 4 d.p.i. For late CD16 blockade studies, the mice were treated with anti-CD16 antibodies at 7 d.p.i. and 11 d.p.i. and euthanized at 14 d.p.i. For depletion experiments, the mice were treated with anti-CD16 (Thermo Fisher Scientific, 3G8) antibodies with a daily dose of 20 µg for 3 days starting 1 d.p.i. Rabbit IgG, monoclonal (EPR25A) isotype control (ab172730) and mouse IgG1 kappa isotype control (P3.6.2.8.1) were used.
Infected MISTRG6 (without AAV-hACE2) mice were treated with monoclonal antibodies against human ACE2 (MM0073-11A31, Abcam, ab89111) for 3 days i.p. with a daily dose of 20 µg starting at 1 d.p.i. In these mice only, epithelial cells were not infected or infected poorly with SARS-CoV-2 with undetectable titres using standard plaque assays19, presumably due to differences between mouse and human ACE2 that limit viral entry and replication42. Mouse IgG2 isotype was used as a control.
Infected MISTRG6-hACE2 mice received a mixed cocktail of monoclonal antibody clone 135 (m135) and clone 144 (m144) at 20 mg kg−1 at 35 h.p.i. or 7 d.p.i. Monoclonal recombinant antibodies used in this study were cloned from the convalescent patients (whose plasma was used for in vitro studies infecting BMDMs) and had high neutralizing activity against SARS-CoV-2 in vitro and in vivo in mouse adapted SARS-CoV-2 infection and ancestral stain of SARS-CoV-2/WA119,45,64.
For NLRP3-inhibitor experiments, infected MISTRG6-hACE2 mice were treated with MCC950 (R&D Systems) at a dose of 8 mg kg−1 i.p. on days 6, 8, 10 and 12 after infection and euthanized on day 1468,69,70. For caspase-1-inhibitor experiments, infected MISTRG6-hACE2 mice were treated with VX-765 (Invivogen) at a dose of 8 mg kg−1 on days 6, 8, 10 and 12 after infection and euthanized on day 1470. Control infected mice were treated with PBS.
Viral titres
Mice were euthanized in 100% isoflurane. Approximately half of the right lung lobe was placed into a bead homogenizer tube with 1 ml of DMEM + 2% FBS. After homogenization, 300 μl of this mixture was placed in 1 ml Trizol (Invitrogen) for RNA extraction and analysis. The remaining volume of lung homogenates was cleared of debris by centrifugation (3,900g for 10 min). Infectious titres of SARS-CoV-2 were determined by plaque assay in Vero E6 (standard) or Vero ACE2+TMPRSS2+ (sensitive) cells in DMEM 4% FBS and 0.6% Avicel RC-58171. Plaques were resolved at 48 h after infection by fixing in 10% formaldehyde for 1 h followed by staining for 1 h in 0.5% crystal violet in 20% ethanol. Plates were rinsed in water to visualize plaques. Multiple dilutions of lung homogenates were used to quantify infectious titres (the minimum number of plaques that can be quantified is 10 per ml of lung homogenate or ml of supernatant). Viral titres from supernatants of BMDM cultures were determined by plaque assay in Vero ACE2+TMPRSS2+ (sensitive) cells according to the same protocols described for lung homogenates. VERO C1008 (Vero 76, clone E6, Vero E6) were obtained from ATCC. Vero ACE2+TMPRSS2+ cells were obtained from B. Graham. None of the cell lines were authenticated or tested for mycoplasma contamination.
Viral RNA analysis
RNA was extracted using the RNeasy mini kit (Qiagen) according to the manufacturer’s protocol. SARS-CoV-2 RNA levels were quantified using the Luna Universal Probe Onestep RT-qPCR kit (New England Biolabs) and US CDC real-time RT-PCR primer/probe sets for 2019-nCoV_N1. For each sample, 1 µg of RNA was used. Subgenomic viral RNA was quantified using primer and probe sets targeting the E gene as previously described40,63. The primer/probe sequences were as follows: E_Sarbeco_F primer, ACAGGTACGTTAATAGTTAATAGCGT (400 nM per reaction); E_Sarbeco probe_P1, FAM-ACACTAGCCATCCTTACTGCGCTTCG-BBQ (200 nM per reaction); E_Sarbeco_R primer ATATTGCAGCAGTACGCACACA (400 nM per reaction); E leader specific primer sgLead-F, CGATCTCTTGTAGATCTGTTCTC (400 nM per reaction).
Histology and immunofluorescence
Yale pathology provided assistance with embedding and sectioning of lung tissue. A pulmonary pathologist reviewed the slides in a blinded manner and identified immune cell infiltration and other related pathologies. Paraffin-embedded lung tissue (fixed in 4% paraformaldehyde (PFA) for no more than 24 h) sections were deparaffinized in xylene and rehydrated. After antigen retrieval with 10 mM sodium citrate (pH 6) and permeabilization with 0.1% Triton-X for 10 min, the slides were blocked with 5% BSA in PBS with 0.05% Tween-20 for 1 h. The samples were next stained with primary antibodies against SARS-CoV-2-dsRNA, SARS-CoV2-RNA-dependent RNA polymerase, SARS-CoV-2 spike, human CD68, human ACE2 and their isotype controls diluted in 1% BSA overnight at 2–8 °C. The next day, the samples were washed and incubated with fluorescent secondary antibodies. After washes, the samples were treated with TrueBlack lipofuscin autofluorescence quencher for 30 s and mounted onto DAPI mounting medium (Sigma-Aldrich). Images were acquired using the Keyence BZ-X800 Fluorescence Microscope or Nikon ECLIPSE Ti Series Confocal Microscope. Pseudocolours were assigned for visualization. All histopathological assessment and scoring of H&E, trichrome and immunoflourescence staining were performed on blinded samples by a board-certified pathologist.
Isolation of cells and flow cytometry
All mice were analysed at approximately 9–14 weeks of age. Single-cell suspensions were prepared from blood, spleen, bronchioalveolar lavage and lungs. Mice were euthanized with 100% isoflurane. Bronchioalveolar lavage was performed using standard methods with a 22G Catheter (BD). Blood was collected either retro-orbitally or by cardiac puncture after euthanasia. Bronchioalveolar lavage was performed using standard methods with a 22G Catheter (BD)72. Lungs were collected, minced and incubated in a digestion cocktail containing 1 mg ml−1 collagenase D (Sigma-Aldrich) and 30 µg ml−1 DNase I (Sigma-Aldrich) in RPMI at 37 °C for 20 min with gentle shaking. Tissue was then filtered through a 70 or 100 µm filter. Cells were treated with ammonium-chloride-potassium buffer and resuspended in PBS with 1% FBS. Mononuclear cells were incubated at 4 °C with human (BD) and mouse (BioxCell, BE0307) Fc block for 10 min. After washing, primary antibody staining was performed at 4 °C for 20 min. After washing with PBS, cells were fixed using 4% PFA. For intracellular staining, cells were washed with BD permeabilization buffer and stained in the same buffer for 45 min at room temperature. Samples were analysed on an LSRII flow cytometer (BD Biosciences). Data were analysed using FlowJo software. Experimenters were not blinded to group allocation during sample collection of flow cytometric studies but were blinded during data analysis.
For cell sorting experiments, single-cell suspensions from digested lungs were stained with antibodies against human CD45, mouse CD45, mouse EPCAM and sorted using the BD FACS Aria II contained within a Baker BioProtect IV Biological Safety Cabinet. Cell viability was assessed with DAPI when applicable.
For imaging flow cytometry, cells from SARS-CoV-2-infected humanized mice were sorted on the basis of human immune cells (hCD45+); mouse immune cells (mCD45+) or epithelial mouse cells (EPCAM+). A−mNG+ epithelial cells (SARS-CoV-2-mNG+mCD45(PE)−EPCAM(APC)+hCD45(PB)−; B-total mouse immune cells (mCD45(PE)+EPCAM(APC)−hCD45(PB)−; C−mNG+ human immune cells (SARS-CoV-2-mNG+mCD45(PE)−EPCAM(APC)−hCD45(PB)+); D−mNG− human immune cells (SARS-CoV-2-mNG−mCD45(PE)−EPCAM(APC)−hCD45(PB)+). These sorted cells (epithelial or immune cells) were fixed in 4% PFA for at least 30 min. Fixed sorted cells (epithelial or immune cells) were permeabilized, stained with unconjugated primary antibodies against ASC (1:200, rabbit), NLRP3 (1:200, goat), then stained with secondary antibodies (donkey anti-rabbit or goat conjugated with AlexaFluor 546 or 647, at 1:1,000). Cells data were acquired using the ImageStream X MKII (Amnis) system with ×63 magnification and analysed using Ideas software (Amnis). ASC, NLRP3 specks were gated and quantified on the basis of fluorophore intensity/maximum pixels. For FLICA–caspase-1 colocalization, macrophages were pretreated with FLICA before sorting.
In vitro infection of human macrophages with SARS-CoV-2
Using aseptic techniques under sterile conditions, bone marrow cells were isolated from femurs of immune-reconstituted MISTRG6 mice. For differentiation into human macrophages in vitro, bone marrow cells were incubated in medium supplemented with 10% FBS, 1% penicillin–streptomycin and recombinant human M-CSF (50 ng ml−1), GM-CSF (50 ng ml−1) and IL-4 (20 ng ml−1) at 1 × 106 per ml concentration for 6 days in an incubator under 5% CO2 and at 37 °C. Medium supplemented with 10% FBS was replenished with new medium every 3–4 days. Before infection with SARS-CoV-2, these bone-marrow-derived macrophages were monitored for granularity, elongated morphology and stronger adherence to the plate. Purity of cultures was confirmed by flow cytometry. Human macrophages were then cultured with SARS-CoV-2-mNG in the presence or absence of the plasma of patients with COVID-19 or healthy individuals, monoclonal antibodies (mix of clones 135 and 144, described as therapeutics), remdesivir, anti-CD16 antibodies, anti-ACE2 antibodies, control isotype antibodies, caspase-1 inhibitor (VX-76573) or NLRP3 inhibitor (MCC950).
Human BMDMs in vitro or human lung macrophages ex vivo were cultured with a viral inoculum at 104 PFU of SARS-CoV-2-mNG (multiplicity of infection of around 0.1). These macrophage cultures were then incubated at 37 °C under 5% CO2 for 24, 48 and 72 h at which time cells were collected. Cells were dissociated from the culture plate with 10 mM EDTA or Accutase (Thermo Fisher Scientific) cell dissociation reagent (10–20 min). For studies pertaining to the mechanism of viral entry, viral replication and inflammasome activation, infected macrophages were treated with remdesivir (10 μM), anti-CD16 (Abcam clone, 10 µg ml−1) and anti-ACE2 (10 µg ml−1), caspase-1 inhibitor (VX765, 20 µM) and NLRP3 inhibitor (MCC950, 20 µg ml−1) in culture. Cells were stained when applicable and fixed for 30 min with 4% PFA. Convalescent plasma samples from the top 30 neutralizers in a cohort of 148 individuals were pooled to create a mixture with an NT50 titre of 1,597 against HIV-1 pseudotyped with SARS-CoV-2 S protein45. We used this pooled serum at a concentration of 5 µl plasma per ml for in vitro experiments and refer to it as plasma from patients with COVID-19. Healthy plasma was collected from healthy volunteers and pooled before the COVID-19 pandemic and was used at a concentration of 5 µl plasma per ml. Monoclonal antibodies (a mix of clones 135 and 144) were used at 4 µg per ml concentration.
Zombie Aqua and annexin V staining
Single-cell suspensions from in vitro cultures or enzymatically dissociated lungs were washed and stained for viability with Zombie Aqua (BioLegend, 423101) in PBS (1:400) for 15 min at 4 °C. Without washing the cells, cell surface antibody cocktail was added, and cells were incubated for another 15 min. Cells were then washed with PBS twice and resuspended in annexin V binding buffer. Cells were stained with annexin V PE (1:400) in binding buffer for 15 min at 4 °C. Cells were then washed with annexin V buffer and fixed in 4% PFA.
FLICA assay
Single-cell suspensions from in vitro cultures or enzymatically dissociated lungs were resuspended in RPMI 10% FBS with FLICA substrate (BioRad-FLICA 660 caspase-1 kit- ICT9122) and cultured for 1 h (for microscopy) or 30 min (for flow cytometry) at 37 °C. Cells were then washed twice with PBS and stained with Zombie Aqua and annexin V as described. Cells were then fixed with 1× Fixative (provided in the BioRad-FLICA caspase-1 kit) for at least 1 h, but not exceeding 16 h. Cells were kept at 4 °C until further staining and analysis. The FLICA 660 caspase-1 kit uses a target sequence (YVAD) sandwiched between a far-red fluorescent 660 dye (excitation max 660 nm, emission max 685 nm).
LDH measurement
LDH levels were measured from freshly collected supernatant of infected cells (BMDMs) or freshly collected serum using the CyQUANT LDH Cytotoxicity Assay (Thermo Fisher Scientific, C20300) according to the manufacturer’s instructions under BSL3 conditions.
Human samples
For this study we acquired six control uninfected, and two SARS-CoV-2-infected deidentified lung (4 different cuts) samples as paraffin-embedded tissues from autopsies of individuals admitted to Yale New Haven Hospital. Lungs were fixed in 10% formalin (details of patient samples are provided in Supplementary Table 5).
Cytokine, chemokine and IgG quantification
Human IL-18 (Sigma-Aldrich or RND), human CXCL10 (RND), human IL-1RA (Abcam), human gasdermin D (MyBioSource) were quantified from supernatants of BMDMs infected (or not) with SARS-CoV-2-mNG or from serum or lung homogenates of SARS-CoV-2-mNG-infected (or not) MISTRG6 or MISTRG6-hACE2 mice according to the manufacturer’s instructions. Human IL-1β was quantified from supernatants of BMDMs infected with SARS-CoV-2-mNG using the cytometric bead array for human IL-1B (BD) according to the manufacturer’s instructions. Human anti-spike-RBD IgG (BioLegend) was quantified from sera and lung homogenates of infected or uninfected MISTRG6-hACE2 mice.
Antibodies
Flow cytometry
All antibodies used in flow cytometry were obtained from BioLegend, unless otherwise specified.
Antibodies against the following antigens were used for characterization or isolation of cells by flow cytometry. Mouse antigens: CD45 (30-F11, 103130), CD45 (30-F11, 103108), CD45 (30-F11, 103147), CD326 (G8.8, 118218), F4/80 (BM8, 123117). Human antigens: CD45 (Hl30, 304044), CD45 (Hl30, 304029), CD3 (UCHT1, 300408), CD14 (HCD14, 325620), CD16 (3G8, 302030), CD16 (3G8, 302006), CD19 (HIB19, 302218), CD19 (HIB19, 302226), CD33 (WM53, 983902), CD20 (2H7, 302313), CD20 (2H7, 302322), CD206 (Clone15-2, 321106), CD206 (15-2, 321109), CD86 (BU63, 374210), CD123 (6H6, 306006), CD11B (M1/70, 101242), CD11C (3.9, 301608), HLA-DR (LN3, 327014), HLA-DR (LN3, 327020), HLA-DR (LN3, 327005), CD183 (G025H7, 353720), CD335-NKp46 (9E2, 331916), CD4 (OKT4, 317440), CD8 (SK1, 344718), CD8 (SK1, 344748), CD68 (Yl/82A, 333828).
Immunofluorescence
Anti-dsRNA antibodies (rJ2) were purchased from Sigma-Aldrich (MABE1134) or Antibodies online (Ab01299-23.0). Polyclonal SARS-CoV-2 RNA-dependent RNA polymerase antibodies were purchased from Cell Signaling (67988S). Monoclonal SARS-CoV-2 RNA-dependent RNA polymerase antibodies were purchased from Kerafest (ESG004). Anti-spike (spike 1) antibodies (1A9, GTX632604) were obtained from GeneTex. Anti-spike (spike 2) antibodies (T01Khu, 703958) were obtained from Thermo Fisher Scientific.
Imaging flow cytometry
Mouse anti-human PE-Cy7 CD16 (3G8) was purchased from BioLegend (302016). Rabbit anti-human ASC (polyclonal) was purchased from Santa Cruz (sc-22514-R). Goat anti-human NLRP3 (polyclonal) was purchased from Abcam (ab4207). Donkey anti-rabbit IgG (H+L) highly cross-adsorbed secondary antibody (Polyclonal) was purchased from Thermo Fisher Scientific (A-31573). Donkey anti-rabbit IgG (H+L) cross-adsorbed secondary antibody (polyclonal) was purchased from Thermo Fisher Scientific (A-10040). Donkey anti-goat IgG (H+L) cross-adsorbed secondary antibody (polyclonal) was purchased from Thermo Fisher Scientific (A-21447).
Therapeutic antibodies
Monoclonal antibodies against human CD16 used in blocking experiments were purchased from Abcam (SP175). Monoclonal antibodies against human ACE2 were purchased from Abcam. Anti-CD16 antibodies used in depletion experiments was purchased from Thermo Fisher Scientific (3G8). Monoclonal antibodies (clones 135 and 144) were acquired from M. Nussenzweig as previously described45. Anti-IFNAR2 antibodies were purchased from PBL Assay science (21385-1).
Gene expression
RNA was extracted using the RNeasy mini kit (Qiagen) according to the manufacturer’s protocol. The High-Capacity cDNA Reverse Transcription Kit was used to make cDNA. Quantitative PCR with reverse transcription was performed using the SYBR FAST universal qPCR kit (KAPA Biosystems). Predesigned KiCqStart primers for DDX58, IL6, IFITM3, IRF7, IFIH1, IFNA6, IFNG and HPRT1 were purchased from Sigma-Aldrich.
Bulk whole-tissue RNA-seq analysis
RNA isolated from homogenized lung tissue, also used for viral RNA analysis, was prepared for whole-tissue transcriptome analysis using low input (14 d.p.i.) or conventional (28 d.p.i.) bulk RNA sequencing (RNA-seq). Libraries were generated with the help of the Yale Center for Genomic Analysis. In brief, libraries were prepared using the Illumina rRNA depletion kit and sequenced on a NovaSeq system. Raw sequencing reads were aligned to the human–mouse combined genome with STAR74, annotated and counted with HTSeq75, normalized using DESeq276 and graphed using the Broad Institute Morpheus web tool. Heat maps visualize normalized counts of duplicates as minimum–maximum-transformed values, calculated by subtracting the row mean and diving by the s.d. for each gene. Rows (genes) were clustered by hierarchical clustering (1 − Pearson) or k-means clustering as indicated in the figure legends. Differential expression analysis was also performed with DESeq2. For IFN-stimulated gene identification, http://www.interferome.org was used with parameters -In Vivo, -Mus musculus or Homo sapiens -fold change up 2 and down 2.
10x Genomics scRNA-seq
Sorted human lung immune cells (hCD45+ in uninfected, 14 d.p.i. and 28 d.p.i.) were stained with TotalSeq (TotalSeq, B0251, anti-human hashtag 1 antibody, GTCAACTCTTTAGCG; TotalSeq, B0252, anti-human hashtag 2 antibody, TGATGGCCTATTGGG) antibodies (BioLegend) before processing for droplet-based single-cell RNA-seq (scRNA-seq). Single-cell transcriptomes and associated protocols of 4 d.p.i. lungs (total lung cells as opposed to sorted human immune cells analysed) were previously described19. Duplicates from each condition/time point were pooled in equal numbers to ensure 10,000 cells were encapsulated into droplets using 10x Chromium GEM technology. Libraries were prepared in house using the Chromium Next GEM Single Cell 3′ Reagent Kits v.3.1 (10x Genomics). scRNA-seq libraries were sequenced using the Nova-Seq system. Raw sequencing reads were processed with Cell Ranger v.3.1.0 using a human–mouse combined reference to generate a gene–cell count matrix. To distinguish between human and mouse cells, we counted the number of human genes (nHuman) and mouse genes (nMouse) with non-zero expression in each cell, and selected cells with nHuman > 20 × nMouse as human cells. The count matrix of human cells and human genes was used in the downstream analysis using Seurat (v.3.2)62. Specifically, this matrix was filtered to remove low-quality cells, retaining cells with >200 and <5,000 detected genes and <20% mitochondrial transcripts. We then log-normalized each entry of the matrix by computing log[CPM/100 + 1], where CPM is the counts per million. To visualize the cell subpopulations in two dimensions, we applied principal component analysis followed by t-SNE, a nonlinear dimensionality reduction method, to the log-transformed data. Graph-based clustering was then used to generate clusters that were overlaid on the t-SNE coordinates to investigate cell subpopulations. Marker genes for each cluster of cells were identified using the Wilcoxon test (two-tailed, rank-sum) with Seurat. For adjusted P values, Bonferroni correction was used. In this analysis, 438 cells (uninfected), 336 cells (4 d.p.i.), 793 cells (14 d.p.i.) and 1,368 (28 d.p.i.) cells were included.
To identify DA subpopulations that were not restricted to clusters, we applied DA-seq61, a targeted, multiscale approach that quantifies a local DA measure for each cell for comprehensive and accurate comparisons of transcriptomic distributions of cells. DA measure defined by DA-seq shows how much a cell’s neighbourhood is enriched by the cells from either uninfected or infected lungs. DA-seq analysis of our data revealed that T cells, monocytes and macrophages were responsible for most of the chronic infection driven changes. Red colouring marks enrichment at 28 d.p.i. lungs and blue colouring marks enrichment in uninfected lungs.
To combine cells from different d.p.i. (uninfected, 4 d.p.i., 14 d.p.i., 28 d.p.i.), we applied the integration method62 in Seurat to remove batch effects. We then performed principal component analysis and retained the top 30 PCs as the input to t-SNE, a nonlinear dimensionality reduction method, to embed the data onto two-dimensional space for visualization. Graph-based clustering with a resolution of 0.8 was then used to generate clusters that were overlaid on the t-SNE coordinates to investigate cell subpopulations. Marker genes for each cluster of cells were identified using the Wilcoxon test (two-tailed, rank-sum) with Seurat (for the adjusted P values, Bonferroni correction was used). After cell type identification, we separated out macrophage populations from all d.p.i., and applied the same procedures as described above to repreprocess and visualize the data. Clusters were redefined based on a resolution of 0.3.
Statistics and reproducibility
Unpaired or paired t-tests (always two-tailed) were used to determine statical significance for changes in immune cell frequencies and numbers while comparing infected mice with uninfected control mice or treated mice with untreated mice. To determine whether the viral RNA quantification was statistically significant across treatment groups or timepoints, two-tailed Mann–Whitney U-tests were used. Unpaired t-tests (two-tailed) or ratio paired t-tests (two-tailed) were used to determine whether the viral titre quantification of the untreated condition was significantly different from the treated groups. For the Pearson test, significance was deemed using t-test. The test statistic is based on Pearson's product-moment correlation coefficient cor(x, y) and follows a t-distribution with length(x) − 2 d.f. For Spearman tests, P values were computed using algorithm AS 89 with exact = TRUE. All micrographs presented in the study were representative of at least 3 animals or samples. Each experiment was repeated independently at least twice. All attempts yielded similar results. In the in vivo studies, each dot represents a biologically independent mouse.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
All data supporting the findings of this study are available within the Article and the Supplementary Information. All 10x Genomics single-cell RNA-seq and bulk RNA-seq data supporting the findings of this study have been deposited at the Gene Expression Omnibus (GEO) under accession codes GSE186794 and GSE199272. Source data are provided with this paper.
References
Gaebler, C. et al. Evolution of antibody immunity to SARS-CoV-2. Nature 591, 639–644 (2021).
Acharya, D., Liu, G. & Gack, M. U. Dysregulation of type I interferon responses in COVID-19. Nat. Rev. Immunol. 20, 397–398 (2020).
Pairo-Castineira, E. et al. Genetic mechanisms of critical illness in COVID-19. Nature 591, 92–98 (2020).
Nienhold, R. et al. Two distinct immunopathological profiles in autopsy lungs of COVID-19. Nat. Commun. 11, 5086 (2020).
Zhou, Z. et al. Heightened innate immune responses in the respiratory tract of COVID-19 patients. Cell Host Microbe 27, 883–890 (2020).
Dolan, M. E. et al. Investigation of COVID-19 comorbidities reveals genes and pathways coincident with the SARS-CoV-2 viral disease. Sci. Rep. 10, 20848 (2020).
Delorey, T. M. et al. COVID-19 tissue atlases reveal SARS-CoV-2 pathology and cellular targets. Nature 595, 107–113 (2021).
Singanayagam, A. et al. Duration of infectiousness and correlation with RT-PCR cycle threshold values in cases of COVID-19, England, January to May 2020. Euro Surveill. 25, 2001483 (2020).
Decker, A. et al. Prolonged SARS-CoV-2 shedding and mild course of COVID-19 in a patient after recent heart transplantation. Am. J. Transplant. 20, 3239–3245 (2020).
Jeong, H. W. et al. Viable SARS-CoV-2 in various specimens from COVID-19 patients. Clin. Microbiol. Infect. 26, 1520–1524 (2020).
Liu, W. D. et al. Prolonged virus shedding even after seroconversion in a patient with COVID-19. J. Infect. 81, 318–356 (2020).
van Kampen, J. J. A. et al. Duration and key determinants of infectious virus shedding in hospitalized patients with coronavirus disease-2019 (COVID-19). Nat. Commun. 12, 267 (2021).
Folgueira, M. D., Luczkowiak, J., Lasala, F., Perez-Rivilla, A. & Delgado, R. Prolonged SARS-CoV-2 cell culture replication in respiratory samples from patients with severe COVID-19. Clin. Microbiol. Infect. 27, 886–891 (2021).
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Del Valle, D. M. et al. An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat. Med. 26, 1636–1643 (2020).
Wu, C. et al. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 pneumonia in Wuhan, China. JAMA Intern. Med. 180, 934–943 (2020).
Rodrigues, T. S. et al. Inflammasomes are activated in response to SARS-CoV-2 infection and are associated with COVID-19 severity in patients. J. Exp. Med. 218, e20201707 (2021).
Henry, B. M. et al. Lactate dehydrogenase levels predict coronavirus disease 2019 (COVID-19) severity and mortality: a pooled analysis. Am. J. Emerg. Med. 38, 1722–1726 (2020).
Sefik, E. et al. A humanized mouse model of chronic COVID-19. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01155-4 (2021).
Grant, R. A. et al. Circuits between infected macrophages and T cells in SARS-CoV-2 pneumonia. Nature 590, 635–641 (2021).
Beigel, J. H. et al. Remdesivir for the treatment of COVID-19—final report. N. Engl. J. Med. 383, 1813–1826 (2020).
Pruijssers, A. J. et al. Remdesivir inhibits SARS-CoV-2 in human lung cells and chimeric SARS-CoV expressing the SARS-CoV-2 RNA polymerase in mice. Cell Rep. 32, 107940 (2020).
Yin, W. et al. Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 368, 1499–1504 (2020).
Havervall, S. et al. Symptoms and functional impairment assessed 8 months after mild COVID-19 among health care workers. JAMA 325, 2015–2016 (2021).
Nasserie, T., Hittle, M. & Goodman, S. N. Assessment of the frequency and variety of persistent symptoms among patients with COVID-19: a systematic review. JAMA Netw. Open 4, e2111417 (2021).
Ginestra, J. C., Mitchell, O. J. L., Anesi, G. L. & Christie, J. D. COVID-19 critical illness: a data-driven review. Annu. Rev. Med. 73, 95–111 (2022).
Phetsouphanh, C. et al. Immunological dysfunction persists for 8 months following initial mild-to-moderate SARS-CoV-2 infection. Nat. Immunol. 23, 210–216 (2022).
Basile, K. et al. Cell-based culture of SARS-CoV-2 informs infectivity and safe de-isolation assessments during COVID-19. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa1579 (2020).
Vora, S. M., Lieberman, J. & Wu, H. Inflammasome activation at the crux of severe COVID-19. Nat. Rev. Immunol. 21, 694–703 (2021).
Major, J. et al. Type I and III interferons disrupt lung epithelial repair during recovery from viral infection. Science 369, 712–717 (2020).
The RECOVERY Collaborative Group. Dexamethasone in hospitalized patients with Covid-19—preliminary report. N. Engl. J. Med. 384, 693–704 (2020).
Polak, S. B., Van Gool, I. C., Cohen, D., von der Thusen, J. H. & van Paassen, J. A systematic review of pathological findings in COVID-19: a pathophysiological timeline and possible mechanisms of disease progression. Mod. Pathol. 33, 2128–2138 (2020).
Tian, S. et al. Pathological study of the 2019 novel coronavirus disease (COVID-19) through postmortem core biopsies. Mod. Pathol. 33, 1007–1014 (2020).
Menter, T. et al. Postmortem examination of COVID‐19 patients reveals diffuse alveolar damage with severe capillary congestion and variegated findings in lungs and other organs suggesting vascular dysfunction. Histopathology 77, 198–209 (2020).
Barton, L. M., Duval, E. J., Stroberg, E., Ghosh, S. & Mukhopadhyay, S. Covid-19 autopsies, Oklahoma, USA. Am. J. Clin. Pathol. 153, 725–733 (2020).
Xu, Z. et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 8, 420–422 (2020).
Strunz, M. et al. Alveolar regeneration through a Krt8+ transitional stem cell state that persists in human lung fibrosis. Nat. Commun. 11, 3559 (2020).
Kobayashi, Y. et al. Persistence of a regeneration-associated, transitional alveolar epithelial cell state in pulmonary fibrosis. Nat. Cell Biol. 22, 934–946 (2020).
Choi, J. et al. Inflammatory signals induce AT2 cell-derived damage-associated transient progenitors that mediate alveolar regeneration. Cell Stem Cell 27, 366–382 e367 (2020).
Wolfel, R. et al. Virological assessment of hospitalized patients with COVID-2019. Nature 581, 465–469 (2020).
Xie, X. et al. An infectious cDNA clone of SARS-CoV-2. Cell Host Microbe 27, 841–848 (2020).
Hassan, A. et al. A SARS-CoV-2 infection model in mice demonstrates protection by neutralizing antibodies. Cell. 182, 744–753 (2020).
Muus, C. et al. Single-cell meta-analysis of SARS-CoV-2 entry genes across tissues and demographics. Nat. Med. 27, 546–559 (2021).
Bournazos, S., Gupta, A. & Ravetch, J. V. The role of IgG Fc receptors in antibody-dependent enhancement. Nat. Rev. Immunol. 20, 633–643 (2020).
Davide, F. R. et al. Convergent antibody responses to SARS-CoV-2 in convalescent individuals. Nature 584, 437–442 (2020).
Bournazos, S. & Ravetch, J. V. Fcγ receptor function and the design of vaccination strategies. Immunity 47, 224–233 (2017).
Chakraborty, S. et al. Proinflammatory IgG Fc structures in patients with severe COVID-19. Nat. Immunol. 22, 67–73 (2021).
Hoepel, W. et al. High titers and low fucosylation of early human anti-SARS-CoV-2 IgG promote inflammation by alveolar macrophages. Sci. Transl. Med. 13, eabf8654 (2021).
Larsen, M. D. et al. Afucosylated IgG characterizes enveloped viral responses and correlates with COVID-19 severity. Science 371, eabc8378 (2021).
Evren, E. et al. Distinct developmental pathways from blood monocytes generate human lung macrophage diversity. Immunity 54, 259–275 (2021).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Chua, R. L. et al. COVID-19 severity correlates with airway epithelium-immune cell interactions identified by single-cell analysis. Nat. Biotechnol. 38, 970–979 (2020).
Lei, X. et al. Activation and evasion of type I interferon responses by SARS-CoV-2. Nat. Commun. 11, 3810 (2020).
Zhang, J. Y. et al. Single-cell landscape of immunological responses in patients with COVID-19. Nat. Immunol. 21, 1107–1118 (2020).
Allen, I. C. et al. The NLRP3 inflammasome mediates in vivo innate immunity to influenza A virus through recognition of viral RNA. Immunity 30, 556–565 (2009).
Thomas, P. G. et al. The intracellular sensor NLRP3 mediates key innate and healing responses to influenza A virus via the regulation of caspase-1. Immunity 30, 566–575 (2009).
Pothlichet, J. et al. Type I IFN triggers RIG-I/TLR3/NLRP3-dependent inflammasome activation in influenza A virus infected cells. PLoS Pathog. 9, e1003256 (2013).
Ferreira, A. C. et al. SARS-CoV-2 engages inflammasome and pyroptosis in human primary monocytes. Cell Death Discov. 7, 43 (2021).
Pan, P. et al. SARS-CoV-2 N protein promotes NLRP3 inflammasome activation to induce hyperinflammation. Nat. Commun. 12, 4664 (2021).
Xu, H. et al. SARS-CoV-2 viroporin encoded by ORF3a triggers the NLRP3 inflammatory pathway. Virology 568, 13–22 (2022).
Zhao, J. et al. Detection of differentially abundant cell subpopulations in scRNA-seq data. Proc. Natl Acad. Sci. USA 118, e2100293118 (2021).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Corman, V. M. et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 25, 2000045 (2020).
Schäfer, A. et al. Antibody potency, effector function, and combinations in protection and therapy for SARS-CoV-2 infection in vivo. J. Exp. Med. 218, e20201993 (2020).
Yu, H. et al. A novel humanized mouse model with significant improvement of class-switched, antigen-specific antibody production. Blood 129, 959–969 (2017).
Rongvaux, A. et al. Development and function of human innate immune cells in a humanized mouse model. Nat. Biotechnol. 32, 364–372 (2014).
Israelow, B. et al. Mouse model of SARS-CoV-2 reveals inflammatory role of type I interferon signaling. J. Exp. Med. 217, e20201241 (2020).
Coll, R. C. et al. A small-molecule inhibitor of the NLRP3 inflammasome for the treatment of inflammatory diseases. Nat. Med. 21, 248–255 (2015).
Perera, A. P. et al. MCC950, a specific small molecule inhibitor of NLRP3 inflammasome attenuates colonic inflammation in spontaneous colitis mice. Sci. Rep. 8, 8618 (2018).
Dixon, K. O. et al. TIM-3 restrains anti-tumour immunity by regulating inflammasome activation. Nature 595, 101–106 (2021).
Wei, J. et al. Genome-wide CRISPR screens reveal host factors critical for SARS-CoV-2 infection. Cell 184, 76–91 (2020).
Sun, F., Xiao, G. & Qu, Z. Murine bronchoalveolar lavage. Bio Protoc. 7, e2287 (2017).
Stack, J. H. et al. IL-converting enzyme/caspase-1 inhibitor VX-765 blocks the hypersensitive response to an inflammatory stimulus in monocytes from familial cold autoinflammatory syndrome patients. J. Immunol. 175, 2630–2634 (2005).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
The generation of the original MISTRG6 model was supported by the Bill and Melinda Gates Foundation. We thank G. Yancopoulos, D. Valenzuela, A. Murphy, and W. Auerbach at Regeneron Pharmaceuticals who generated, in collaboration with our groups, the individual knockin alleles combined in MISTRG; M. Chiorazzi, I. Odell and W. Philbrick and all of the other members of the Flavell laboratory for discussions and comments; J. Alderman, B. Cadugan and E. Hughes-Picard for administrative assistance; B. Cadugan and E. Eynon for manuscript editing; P. Ranney, C. Hughes for mouse colony management; D. Urbanos for human CD34+ cell isolation; L. Devine and E. Manet for help with cell sorting; B. Fontes and M. Schniederberend for biosafety guidance and training; R. Filler for help with viral stocks and cell lines. E. Sefik is a HHMI Fellow of the Damon Runyon Cancer Research Foundation (DRG-2316–18). H.R.S was supported by an NSF Graduate Research Fellowship Program under DGE1752134. This work was funded by the Howard Hughes Medical Institute (to R.A.F., M.N. and A.I.). This study was also supported in part by awards from National Institute of Health grants R35 GM142687 (to Y.G.C.), R01AI157488 (to A.I.), 2T32AI007517 (to B.I.), R01 AI061093 (to E.M.), AI118855 (to E.M.), CA016359 (to E.M.), K08 AI128043 (to C.B.W.), U01 CA260507 (to S.H.), Animal Modeling Core of the Yale Cooperative Center of Excellence in Hematology at Yale University School of Medicine U54 DK106857 (S.H., R.A.F.), R01 GM131642 and GM135928 (to R.A.F. and Y.K.), the Burroughs Wellcome Fund (to C.B.W.), the Patterson Foundation (to C.B.W.), a Fast Grant from Emergent Ventures at the Mercatus Center (to A.I. and C.B.W.), the Mathers Foundation (to A.I., C.B.W. and E.M.), the Ludwig Family Foundation (to A.I. and C.B.W.), the Harvard Lemann Brazil Research Fund (to J.L.) and the Rita Allen Foundation (to Y.G.C.).
Author information
Authors and Affiliations
Contributions
E.S. conceived the project, performed experiments, analysed the data and wrote the manuscript. R.Q. and J.Z. performed bioinformatics analysis. C.J. performed imaging flow cytometry experiments for characterization of the NLRP3 inflammasome. E.K. prepared samples for histopathological assessment and performed all immunofluorescence staining. H.M. performed histopathological assessment of lung pathology, quantification of immunofluorescence staining and offered essential conceptual insights in interpreting lung pathology. B.I. helped to establish the model in biosafety level 3. M.N. provided monoclonal antibodies used in the study. H.N.B. helped with tissue preparation and immunofluorescence staining. S.V. provided help with IL-1 quantification protocols. Y.G.C. provided protocols and insights on dsRNA staining. J.R.B., A.H., H.R.S., S.H., A.I., E.M., M.N., J.L., C.B.W. and Y.K. offered conceptual insights, contributed to the overall interpretation of this work and helped with writing the manuscript. R.A.F. co-conceived and supervised the project, helped to interpret the work and supervised the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.A.F. is an advisor to Glaxo Smith Kline, Zai Labs and Ventus Therapeutics. J.L. is an advisor of Ventus Therapeutics. S.H. is a consultant for FORMA Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Stanley Perlman, Gestur Vidarsson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Targeting viral replication and the downstream interferon response attenuates the hyperactive immune/inflammatory response (matched to figure 1).
a. Representative gating strategy of human immune cells in the lungs of SARS-CoV-2 infected MISTRG6-hACE2 mice. Cells isolated from lungs or bronchioalveolar lavage (BAL) were stained with antibodies against human CD45, HLA-DR, CD68, CD16, CD14, CD206, CD86, CD11B, CD11C, CD123, CD3, and mouse CD45. Cell numbers were calculated using counting beads. b. Human immune cells (numbers) in BAL (14 d.p.i.) or lungs (14 and 28 d.p.i.) of SARS-CoV-2 infected MISTRG6-hACE2 mice treated with dexamethasone (Dex), Remdesivir (RDV), anti-IFNAR2 or a combined therapy of Remdesivir (RDV) and anti-IFNAR2. 14 d.p.i., BAL: CTRL-infected n = 5; Dex, RDV, anti-IFNAR2 n = 3; anti-IFNAR2+RDV n = 4 biologically independent mice examined over 2 independent experiments. 14 d.p.i., Lung: CTRL-infected n = 7; Dex, RDV, anti-IFNAR2, anti-IFNAR2+RDV n = 4 biologically independent mice examined over 3 independent experiments. 28 d.p.i., Lung: CTRL infected n = 5, Dex n = 4, RDV n = 3, anti-IFNAR2 n = 3, anti-IFNAR2+RDV n = 6 biologically independent mice examined over 3 independent experiments. Means with individual datapoints plotted. Unpaired, two-tailed t-test. P < 0.0001 = 8.19x10−5. c. Representative flow cytometry plots and frequencies of alveolar macrophages (AMs) (middle: hCD206hihCD86+hCD68+) or inflammatory macrophages (bottom: hCD206lo/-hCD86+hCD68+) in 14 d.p.i. or 28 d.p.i. lungs of treated or untreated MISTRG6-hACE2 mice. 14 d.p.i.: CTRL infected n = 5, Dex n = 4, RDV n = 4, anti-IFNAR2 n = 4, anti-IFNAR2+RDV n = 4 biologically independent mice examined over at least 2 independent experiments. 28 d.p.i.: CTRL infected n = 5, Dex n = 4, RDV n = 3, anti-IFNAR2 n = 3, anti-IFNAR2+RDV n = 6 biologically independent mice examined over 3 independent experiments. Means with individual datapoints. Unpaired, two-tailed t-test. P < 0.0001 = 4.67x10−5. d. Frequencies (left) and numbers (right) of lung pDCs at 14 d.p.i. CTRL-infected n = 6, Dex n = 4, RDV, anti-IFNAR2 n = 3, anti-IFNAR2+RDV n = 6 mice examined over at least 2 experiments. Means with datapoints. Unpaired, two-tailed t-test. P < 0.0001 = 7.29x10−5. e IFNA transcript levels measured by qPCR in treated or control untreated MISTRG6-hACE2 mice infected with SARS-CoV-2: Uninfected n = 5; CTRL infected: 4 d.p.i. n = 8, 14 d.p.i. n = 9, 28 d.p.i. n = 6; Dex: 14 d.p.i. n = 4, 28 d.p.i. n= 4; RDV 14 and 28 d.p.i. n = 3; anti-IFNAR2 14 and 28 d.p.i. n = 3; anti-IFNAR2+ Remdesivir 14 and 28 d.p.i. n = 4 biologically independent mice examined over at least 2 independent experiments. Normalized to HPRT1. Violin plots with individual datapoints. Unpaired, two-tailed t test. f. Representative histograms and frequencies of HLA-DR+ activated T cells in treated or control mice. 14 d.p.i.: CTRL-infected n = 5, Dex, RDV, anti-IFNAR2, anti-IFNAR2+RDV n = 4; 28 d.p.i.: CTRL infected, Dex, anti-IFNAR2+RDV n = 4, RDV, anti-IFNAR2 n = 3 biologically independent mice examined over 3 independent experiments. Means with datapoints. Unpaired, two-tailed t-test. Some of the data associated with dexamethasone therapy used here as a control have been reported19.
Extended Data Fig. 2 Anti-IFNAR2 and Remdesivir therapy reverses infection induced transcriptional changes (matched to figure 1).
a. Similarity comparison of uninfected, infected, and therapeutically manipulated lungs based on dexamethasone suppressed genes. Pearson correlation. Duplicates analysed for each condition. b. Genes suppressed by both dexamethasone and combined therapy of Remdesivir and anti-IFNAR2 (log2, foldchange < −1, P adj < 0.05). P adj: For the adjusted P values the Bonferroni correction was used. Duplicates analysed for each condition. Dexamethasone suppressed genes significantly overlapped with genes significantly suppressed by combined anti-IFNAR2 and Remdesivir therapy (64% overlap). See Supplementary Table 1 for a full list of genes and their normalized expression. c. Network analysis (STRING v11.0) of genes suppressed by both dexamethasone and combined therapy of Remdesivir and anti-IFNAR2 (as shown in Extended Data Fig. 2b). Duplicates analysed for each condition. K- means clustering (n = 4). d. Pathway (Ingenuity) analysis of genes suppressed by both dexamethasone and combined therapy of Remdesivir and anti-IFNAR2 (as shown in Extended Data Fig. 2b). Duplicates analysed for each condition. Fisher's Exact Test was used to determine statistical significance in the overlap between the dataset genes and the genes suppressed by therapy. e. Transcriptional landscape of human immune cells at single cell level in uninfected or infected (28 d.p.i.). MISTRG6-hACE2 mice. Cluster identifying genes comparing human immune cells from infected (28 d.p.i.) or uninfected lungs for 17 clusters shown in Fig. 1e. Marker genes for each cluster of cells were identified using the Wilcoxon rank-sum test with Seurat. Pooled duplicates analysed for each condition.
Extended Data Fig. 3 Deeper characterization of monocyte/macrophage clusters at early (4 d.p.i.) or late (14 and 28 d.p.i.) SARS-CoV-2 infection (matched to figure 1).
a. Heatmap visualizing cluster identifying genes comparing human monocytes and macrophages from infected (4, 14 or 28 d.p.i.) or uninfected lungs (as shown in Fig. 1g). Pooled duplicates. Uninfected: 438 cells, 4 d.p.i.: 336 cells, 14 d.p.i.: 793 cells, 28 d.p.i.: 1368 cells were analysed. This analysis allowed step by step characterization of the inflammatory macrophage response. Marker genes for each cluster of cells were identified using the Wilcoxon rank-sum test (two-tailed) with Seurat. b. Temporal distribution of transcriptional changes associated with monocytes and macrophages in infected (4, 14 or 28 d.p.i.) or uninfected lungs (as shown in Fig. 1g). Pooled duplicates analysed. Uninfected: 438 cells, 4 d.p.i.: 336 cells, 14 d.p.i.: 793 cells, 28 d.p.i.: 1368 cells included in analysis. c. Top: Heatmap of representative genes that are differentially regulated (DEGs) in human macrophages from 4, 14, 28 d.p.i. lungs compared with uninfected lungs. Uninfected: 438 cells, 4 d.p.i.: 336 cells, 14 d.p.i.: 793 cells, 28 d.p.i.: 1368 cells included in analysis. Bottom: Distribution of interferon stimulated genes within these DEGs. Pooled duplicates analysed.
Extended Data Fig. 4 Therapeutics reduced expression of a representative list of interferon stimulated genes- ISGs (DDX58, IFIH1, IFITM3, IRF7) or inflammatory markers (IL6) (matched to figure 1).
Relative expression of interferon inducible or inflammatory genes in treated or untreated MISTRG6- hACE2 mice infected with SARS-CoV-2 mice at 14 d.p.i. or 28 d.p.i. Uninfected baseline expression values are presented as reference. The distribution of cells that preferentially express these genes is overlayed on the tSNE plots showing 14 d.p.i. and 28 d.p.i. human immune cells. IFITM3 and IL6 were particularly enriched in human macrophage/monocyte clusters, while IRF7, DDX58 and IFIH1 were enriched in multiple human immune cells such as T cells, B cells, and myeloid cells. Normalized to HPRT1. DDX58: uninfected n = 3; CTRL-infected: 14 d.p.i. n = 8, 28 d.p.i. n=: 5; Dex 14 d.p.i. n = 3, 28 d.p.i. n = 6; RDV 14 and 28 d.p.i. n = 3; anti-IFNAR2 14 and 28 d.p.i. n = 3, anti-IFNAR2+ Remdesivir 14 and 28 d.p.i. n = 5 biologically independent mice examined over at least 2 independent experiments. IFIH1: uninfected n = 5; CTRL-infected: 14 d.p.i. n = 11, 28 d.p.i. n=: 3; Dex 14 and 28 d.p.i. n = 4; RDV 14 and 28 d.p.i. n = 3; anti-IFNAR2 14 and 28 d.p.i. n = 3, anti-IFNAR2+ Remdesivir 14 d.p.i. n = 4 and 28 d.p.i. n = 5 biologically independent mice examined over at least 2 independent experiments. IFITM3: uninfected n = 4; CTRL-infected: 14 d.p.i. n = 7, 28 d.p.i. n=: 4; Dex 14 and 28 d.p.i. n = 4; RDV 14 and 28 d.p.i. n = 3; anti-IFNAR2 14 and 28 d.p.i. n = 3, anti-IFNAR2+ Remdesivir 14 and 28 d.p.i. n = 4 biologically independent mice examined over at least 2 independent experiments. IRF7: uninfected n = 4; CTRL-infected: 14 d.p.i. n = 7, 28 d.p.i. n=: 5; Dex 14 and 28 d.p.i. n = 4; RDV 14 and 28 d.p.i. n = 3; anti-IFNAR2 14 and 28 d.p.i. n = 3, anti-IFNAR2+ Remdesivir 14 d.p.i. n = 4, 28 d.p.i. n = 5 biologically independent mice examined over at least 2 independent experiments. IL6: uninfected n = 5; CTRL-infected: 14 d.p.i. n = 9, 28 d.p.i. n=: 4; Dex 14 d.p.i. n = 3, 28 d.p.i. n = 4; RDV 14 and 28 d.p.i. n = 3; anti-IFNAR2 14 and 28 d.p.i. n = 3, anti- IFNAR2+ Remdesivir 14 d.p.i. n = 4, 28 d.p.i. n = 5 biologically independent mice examined over at least 2 independent experiments. Unpaired, two-tailed t-test.
Extended Data Fig. 5 Anti-IFNAR2 and Remdesivir combined therapy reverses fibrotic transcriptional signature and prevents the transition to fibrosis seen in the infected mice (matched to figure 1).
a. Relative expression of IFNG in treated or untreated MISTRG6-hACE2 mice infected with SARS-CoV-2 mice at 14 d.p.i. or 28 d.p.i. Uninfected baseline expression values are presented as reference. Normalized to HPRT1. Uninfected n = 7; CTRL infected: 14 d.p.i. n = 7, 28 d.p.i. n=: 6; Dex 14 d.p.i. = 3, 28 d.p.i. = 5; anti-IFNAR2+ Remdesivir 14 d.p.i. n = 4, 28 d.p.i. n = 6 biologically independent mice examined over at least 2 independent experiments. Unpaired, two-tailed t-test. b. Heatmap of AT2 cell self-renewal and AT1 differentiation and pre-alveolar type 1 transitional cell state (PATS) associated genes at in uninfected or infected (14 d.p.i.) lungs in response to therapeutics. AT2 cell self-renewal and AT1 differentiation gene signature was inhibited while PATS gene signature was enriched in autopsy lungs of patients with severe COVID-197. Top differentially expressed genes in epithelial cluster 7 of autopsy lungs7 were used in the analysis. Duplicates were analysed for each condition. Normalized counts of duplicates visualized as min-max transformed values, calculated by subtracting row mean and diving by SD for each gene. Rows (genes) clustered by hierarchical clustering (one-minus Pearson). c. Representative images of trichrome staining and box and whisker plot (min to max, with all datapoints) of the trichrome scoring of MISTRG6-hACE2 mice treated with a combined therapy of Remdesivir and anti-IFNAR2 or not (CTRL infected). The whiskers go down to the smallest value (minimum) and up to the largest value (maximum). The box extends from the 25th to 75th percentiles. The median is shown as a line in the center of the box. N = 4 biologically independent mice examined over 2 independent experiments. Unpaired, two-tailed t-test.
Extended Data Fig. 6 Cellular source of persistent SARS-CoV-2 viral RNA and sustained viral replication in lungs (matched to figure 2).
a. Quantification of genomic (gRNA) and subgenomic (sgRNA) viral RNA (E-gene) in whole homogenized lung tissue at 4, 14 and 28 d.p.i. 4 d.p.i.: n = 7, 14 d.p.i. n = 5, 28 d.p.i. n = 4 biologically independent mice examined over 3 independent experiments. Means with all datapoints and SD. b. Quantification of genomic (gRNA) and subgenomic (sgRNA) viral RNA (E-gene) in whole homogenized lung tissue at 14 d.p.i. in mice treated with combined therapy of Remdesivir and anti-IFNAR2. CTRL: n = 4, anti-IFNAR2+RDV: n = 4 biologically independent mice examined over 2 independent experiments. N.D.= not detected. c. Quantification of genomic (gRNA) and subgenomic (sgRNA) viral RNA (E-gene) in whole homogenized lung tissue at 28 d.p.i. in mice treated with Remdesivir, anti-IFNAR2 or combined therapy of Remdesivir and anti-IFNAR2. N = 3 biologically independent mice representative of 2 independent experiments. N.D.= not detected. d. Representative gating strategy for sorting human immune cells (human CD45+) or mouse epithelial cells (mouse EPCAM+) from lungs of mice infected with SARS-CoV-2 and quantification of viral RNA (E and N genes) in these sorted cells. N gene: 4 d.p.i. n = 3, 14 d.p.i. n = 6(epithelial), n = 5 (immune), 28 d.p.i. n = 4 (epithelial) n = 3 (immune) biologically independent mice analysed over 3 independent experiments. E gene: 4 d.p.i. n = 3, 14 d.p.i. n = 7 (epithelial), n = 6 (immune), 28 d.p.i. n = 4 (epithelial) n = 3 (immune) biologically independent mice analysed over 3 independent experiments. e. mNG signal in epithelial (EPCAM+) cells from lungs and BAL of mice infected with reporter SARS-CoV-2-mNG or control wild type SARS-CoV-2/WA1. mNG is expressed in infected cells following viral replication. Representative of n = 4 biologically independent mice examined over 2 independent experiments. f. Representative histograms of mNG expression in human or mouse lung macrophages isolated from BAL of infected MISTRG6-hACE2 mice at 4 d.p.i. Representative of n = 3 biologically independent mice examined over 2 independent experiments. g. Frequencies of mNG+ cells within human lung immune cells (hCD45+) of SARS-CoV-2-mNG infected MISTRG6-ACE2 mice at 4 d.p.i. and 14 d.p.i. 4 d.p.i. n = 4, 14 d.p.i. n = 6 biologically independent mice examined over at least 2 experiments. Unpaired, two-tailed t-test. P value = 0.066. h. Viral titres measured as PFU using Vero ACE2+TMPRSS2+ cells that over express human ACE2 from lung homogenates of MISTRG6 mice transduced with AAV-hACE2 (+AAV) or not (−AAV) and infected with SARS-CoV-2. MISTRG6-hACE2 (+AAV): 2 d.p.i. n = 2, 4 d.p.i. n = 5, 7 d.p.i. n = 2, 14 d.p.i. n = 6 MISTRG6(−AAV): 2 d.p.i. n = 4, 4 d.p.i. n = 10 and 7, 14 d.p.i. n = 2, biologically independent mice representative of at least 2 independent experiments. Viral titres using standard Vero E6 cells do not have any detectable titres (previously reported19) in MISTRG6 mice without AAV-hACE2. Some of the MISTRG6-hACE2 data presented here have been previously reported as part of the characterization of the model19. i. Frequencies of mNG+ cells within human macrophages (human CD68+) isolated from lungs of infected MISTRG6 mice transduced with AAV-hACE2 (AAV+) or not (AAV−). MISTRG6 mice with and without AAV-hACE2 were reconstituted with human progenitor cells from the same donor. AAV+ n = 6, AAV− n = 5 biologically independent mice examined over 3 independent experiments. j. Representative gating strategy for sorting mNG+ and mNG− human immune cells, mNG+ and mNG− mouse epithelial cells and mouse immune cells. Lung cells from SARS-CoV2-mNG infected MISTRG6-hACE2 mice were stained with antibodies against human CD45, mouse CD45, and mouse EPCAM. Sorted cells were used for viral RNA quantification (Fig. 2) and characterization of the inflammasome pathway (Fig. 3).
Extended Data Fig. 7 Viral replication products are detected in human lung macrophages of infected MISTRG6-hACE2 mice (matched to figure 2).
a. Representative fluorescent microscopy images showing colocalization of double stranded RNA (dsRNA, clone rJ2) staining, mNG signal and DAPI staining in fixed lung tissue at 4 d.p.i. Representative of n = 4 biologically independent mice examined over 2 independent experiments. b. Representative fluorescent microscopy images of RNA dependent RNA polymerase (RdRp), anti- human CD68 and DAPI staining in fixed lung tissue from SARS-CoV-2 infected or control MISTRG6-hACE2 mice (Non-SARS-CoV-2 pneumonia). Representative of n = 7 biologically independent SARS-CoV-2 infected mice examined over 3 independent experiments. Yellow arrows mark RdRp+ human macrophages. Blue arrows mark RdRp− human macrophages. Isotype controls (bottom panels) and non- COVID pneumonia lungs (bacterial infection, top panels) n = 3 biologically independent mice are presented as controls. Pseudo-colours were assigned for visualization.
Extended Data Fig. 8 Viral RNA dependent RNA polymerase (RdRp) and Spike in human lung macrophages of MISTRG6-hACE2 mice infected with SARS-CoV-2 (matched to figure 2).
a. Representative fluorescent microscopy images of Spike (S), human CD68, and DAPI staining in fixed lungs of SARS-CoV-2- infected MISTRG6-hACE2 mice. Yellow rectangle provides a higher magnification view of the selected area. Pseudo-colours were assigned for visualization. Representative of n = 5 biologically independent mice examined over 3 independent experiments. b. Quantification of viral replication products or machinery in human lung macrophages from SARS-CoV-2 infected MISTRG6-hACE2 mice measured by immunofluorescence staining. Quantification was performed based on representative high-power images (40x) in areas showing diffuse alveolar damage. Frequencies of dsRNA, mNG, RdRp, and Spike positive human macrophages out of hCD68+DAPI+ cells are plotted. dsRNA: 20, 81, 133, 135, 52 human macrophages were counted. mNG: 30, 103, 110 human macrophages were counted. RdRp monoclonal: 187, 59, 85, 106, 142, 63, 59 human macrophages were counted. RdRp polyclonal: 134, 21, 22, 218, 44 human macrophages were counted. Spike (antibody 1): 21, 22, 218, 44 total human macrophages were counted. Spike (antibody 2): 63, 83, 163, 101, 57 human macrophages were counted. N = 5 (dsRNA+), N = 3 (mNG), N = 7 (RdRp+, monoclonal), N = 5 (RdRp+, polyclonal), N = 4 (Spike+-1), N = 5 (Spike+-2) biologically independent mice representative of at least 2 independent experiment. Means with all datapoints are shown. See Methods for details of antibodies used. c.Representative fluorescent microscopy images and quantification of colocalization of Spike (S), RNA dependent RNA polymerase (RdRp), human CD68, and DAPI staining in fixed lungs of SARS-CoV-2- infected MISTRG6-hACE2 mice. Top panel: isotype control staining of SARS-CoV-2- infected lungs. Middle panel: control lungs with Non-SARS-CoV-2, bacterial pneumonia. Bottom panels: SARS-CoV-2- infected MISTRG6-hACE2 mice. Yellow rectangle provides a higher magnification view of the selected area. Pseudo-colours are assigned for visualization. Representative of n = 5 biologically independent mice over 3 independent experiments.
Extended Data Fig. 9 Viral RNA dependent RNA polymerase (RdRp) and Spike in human macrophages of human autopsy lungs with SARS-CoV-2 pneumonia (matched to figure 2).
Representative fluorescent microscopy images and quantification of colocalization of Spike (S), RNA dependent RNA polymerase (RdRp), human CD68 and DAPI staining in fixed human autopsy lungs with SARS-CoV-2 pneumonia or non-SARS-CoV-2 pneumonia. Quantification was performed based on representative high-power images (40x) in areas showing diffuse alveolar damage. Top panels: Representative of RdRp staining with human CD68; middle panels: Representative of Spike staining with CD68; bottom panels: RdRp and Spike staining in SARS-CoV-2- infected autopsy lungs. Yellow rectangle provides a higher magnification view of the selected area. Pseudo-colours are assigned for visualization. SARS-CoV-2 pneumonia n = 4, non-SARS-CoV-2 pneumonia n = 3 biologically independent specimens.
Extended Data Fig. 10 Human macrophage infection was enhanced by antibodies and reduced by CD16, ACE2, or RdRp blockade in vivo and in vitro (matched to figure 2).
a. Viral titres in lung homogenates of Remdesivir (RDV) treated or control untreated MISTRG6-hACE2 mice infected with SARS-CoV-2-mNG. CTRL infected n = 6, RDV treated n = 6 biologically independent mice examined over 3 independent experiments. b. Representative histograms and mean florescent intensity (MFI) for human ACE2 expression in mNG+ or mNG− epithelial cells from MISTRG6-hACE2 (AAV+) mice or total epithelial cells from MISTRG6 (AAV−) mice infected with SARS-CoV-2-mNG. AAV+ n = 10, AAV− n = 6 biologically independent mice examined over at least 3 independent experiments. Paired, two-tailed t-test. c. Representative histograms for human ACE2 expression in mNG+ or mNG− human macrophages, human B cells (CD19+) or mouse immune cells isolated from MISTRG6-hACE2 mice infected with SARS-CoV-2-mNG. Representative of N = 10 mice for epithelial cells, n = 7 mice for human macrophages examined over at least 3 independent experiments. d. MFI of human ACE2 expression in mNG+ or mNG− human macrophages or mouse epithelial cells isolated from SARS-CoV-2-mNG infected MISTRG6-hACE2 mice. Epithelial cells n = 10, human macrophages n = 7 biologically independent mice examined over at least 3 independent experiments. Paired, two-tailed t-test. e. MFI of human ACE2 expression in mNG+ or mNG− human macrophages isolated from MISTRG6 (AAV−) mice infected with SARS-CoV-2-mNG. Epithelial cells are virtually not infected with SARS-CoV-2-mNG in MISTRG6 mice without transduced human ACE2. N = 8 biologically independent mice examined over at least 3 independent experiments. Paired, two-tailed t-test. f. Representative fluorescent microscopy images showing colocalization of human ACE2 and human CD68 cells in SARS-CoV-2 infected MISTRG6-hACE2 mice. Representative of 3 independent mice over 2 independent experiments. g. Anti-Spike (RBD) IgG levels measured by ELISA in serum or lung homogenates of SARS-CoV-2 infected (4 and 14 d.p.i.) or uninfected MISTRG6-hACE2 mice treated therapeutically with monoclonal antibodies (mAbs) against SARS-CoV-2 spike protein (treated at 35 h.p.i. or 7 d.p.i.) or not. Lung homogenates: Uninfected n = 5, 4 d.p.i. n = 8, 14 d.p.i. n = 8, 4 d.p.i.+mAb n = 2, 14 d.p.i.+mAb n = 2 biologically independent mice representative of at least 2 experiments. Serum: Uninfected n = 3, 4 d.p.i. n = 3, 14 d.p.i. n = 3, 4 d.p.i.+mAb n = 3, 14 d.p.i.+mAb n = 2 biologically independent mice representative of at least 2 experiments. Unpaired, two-tailed t-test. h. Frequencies of mNG signal in human immune cells in infected mice (14 d.p.i.) treated therapeutically with monoclonal antibodies45,64 (mAb) at 7 d.p.i. CTRL infected n = 5, mAb treated n = 4 biologically independent mice examined over 2 independent experiments. Means with datapoints and SD. Paired, two-tailed t-test. i. Two-way plot showing anti-Spike (RBD) IgG levels and corresponding mNG+ human immune cell proportions in lungs of infected MISTRG6-hACE2 mice at 4 d.p.i. Pearson’s correlation value = 0.70. N = 8 biologically independent mice examined over 4 independent experiments. j. Frequencies of mNG+ human macrophages in human immune cells in SARS-CoV-2-mNG infected MISTRG6-hACE2 mice treated with anti-CD16 antibody (Abcam-clone SP175) at 7 d.p.i. and 11 d.p.i. and analysed at 14 d.p.i. n = 6 biologically independent mice examined over 3 independent experiments. Unpaired, two-tailed t-test. k. Representative histograms and frequencies of mNG+ cells in human BMDMs cultured (or not) with SARS-CoV-2-mNG for 48 h. Human BMDMs were differentiated in vitro from bone marrow cells of immune-reconstituted uninfected MISTRG6 mice. BMDMs were treated with pooled plasma from healthy controls (prior to COVID-19 pandemic) or convalescent COVID-1945 patients during the course of SARS-CoV-2 infection in vitro. Uninfected n = 3, infected+ healthy plasma n = 7, infected+ COVID plasma n = 10 independent samples cultured and analysed over at least 3 experiments. Means with datapoints. Unpaired t-test. P < 0.0001 = 1.57x10−5. l. Quantification of genomic (gRNA) and subgenomic (sgRNA) viral RNA (E gene) in infected BMDMs at 48 h.p.i. Cells were treated with plasma from healthy controls or convalescent COVID-19 patients. Healthy plasma: n = 4, COVID plasma n = 6, RDV: n = 6, anti-CD16+anti-ACE2 n = 4 independent samples analysed over at least 2 independent experiments. Means with datapoints. Mann-Whitney, two-tailed, t-test. m. Representative histograms and frequencies of mNG+ cells in BMDMs and lung macrophages cultured with SARS-CoV-2 in presence of plasma of convalescent COVID-19 patients. mNG+ macrophages were analysed at 48 h.p. BMDMs n= 6, Lung macrophages n = 4 independent samples analysed over 2 independent experiments. Unpaired, two-tailed t-test. n. Frequencies of mNG+ cells in BMDMs cultured with SARS-CoV-2 or not in presence of healthy patient plasma, COVID plasma, monoclonal antibodies (clones 135 and 144) or no antibodies. COVID plasma n = 5, mAb n = 4, healthy plasma n = 4, no Ab n = 5 independent samples analysed over 2 independent experiments. Means with datapoints and SD. The same monoclonal antibody cocktail used was used in vivo (Figs. 2 and 3). Unpaired, two-tailed t-test. o. Representative histograms and frequencies of mNG+ cells in BMDMs cultured with SARS-CoV-2-mNG (or not) in presence or absence of COVID plasma. Cultures were treated with Remdesivir, anti-human CD16 antibody and/or anti-human ACE2 antibody. Healthy plasma n = 5, COVID plasma n = 10, RDV n = 5, anti-CD16 n = 6, anti-ACE2 n = 4, anti-CD16+ACE2 n = 4 independent samples analysed over at least 2 independent experiments. Means with datapoints. Unpaired two-tailed t-test. P values < 0.0001: anti-CD16 vs. COVID plasma=1.98x10−5, RDV vs. COVID plasma = 5.24x10−6. p. Viral titres and representative plaque images from supernatants of human or mouse BMDMs infected with SARS-CoV-2 mNG in vitro (without COVID plasma). Infectious virus from supernatants of infected macrophage cultures collected at 24, 48 and 72 h.p.i. was plaqued using Vero ACE2+TMPRSS2+ cells. Supernatant collected from Vero E6 cell cultures were provided as reference. Human: 24 h.p.i. n = 9, 48 h.p.i. n = 13, 72 h.p.i. n = 4. Mouse: 24 h.p.i. n = 6 independent samples analysed over at least 2 independent experiments. q. Viral titres from supernatants of BMDMs infected with SARS-CoV-2 mNG in vitro and treated with Remdesivir (RDV) or a combination of anti-CD16 and anti-ACE2 antibodies. Cultures were not supplemented with COVID plasma. Infectious virus from supernatants of infected macrophage cultures collected at 24 h.p.i. was plaqued using Vero ACE2+TMPRSS2+ cells. CTRL n = 9, RDV n = 4, anti-CD16 and anti-ACE2 n = 4 independent samples representative of 2 independent experiments. Means with datapoints. r. Viral titres measured as PFUs using supernatants containing concentrations of Remdesivir (1 µm) or anti-ACE2 (1 µg/ml) and anti-CD16 antibodies diluted to (1:10) allow quantification of PFUs at 24 h.p.i. from macrophage cultures. Supernatants were applied on Vero ACE2+TMPRSS2+ cells which were then infected with a matched inoculum of SARS-CoV-2 mNG (103 PFU quantified in Vero-E6 cells) to test carry over effect in plaque quantification. Untreated N = 9, RDV N = 6, anti-ACE2+anti-CD16 n = 4 independent datapoints collected over 3 independent experiments. Means with datapoints. Unpaired, two-tailed, t-test.
Extended Data Fig. 11 SARS-CoV-2 infected human macrophages have a unique transcriptional signature (matched to figure 3).
a. Representative gating strategy of CXCL10 or TNF producing human macrophages in MISTRG6-hACE2 mice infected with SARS-CoV-2-mNG. b. Representative flow cytometry plots of CXCL10 and TNF staining in mice therapeutically treated with mAb or control untreated mice. Representative of n = 4 biologically independent mice examined over 2 independent experiments. c. CXCL10 production measured by ELISA in supernatants of BMDMs infected with SARS-CoV-2 in vitro. Infected BMDM cultures were supplemented with pooled plasma from COVID-19 patients and were treated with Remdesivir or not. Uninfected n = 5, CTRL infected n = 12, RDV n = 4 over 3 independent experiments. Means with individual values are plotted. Unpaired, two-tailed t-test. c. Spearman correlation values of each gene based on its correlation with CXCL10 or TNF or TLR7. d. Expression and distribution of CXCL10, TNF and TLR7 in human immune cells from infected (4, 14 and 28 d.p.i.) MISTRG6-hACE2 mice.
Extended Data Fig. 12 CXCL10-associated genes (matched to figure 3).
a. Network (STRING v11.0) analysis of top CXCL10-associated genes (top 200 genes). K-means clustering. Clusters and their corresponding pathway analysis are available as source files. Top genes that correlate with CXCL10 (4 d.p.i., Pearson and Spearmen correlation combined) are enriched for distinct inflammatory molecules. b. Network (STRING) analysis of genes that are preferentially associated with CXCL10 but not with TLR7 or TNF. Disconnected nodes in the network are not displayed. K-means clustering. Clusters and their corresponding pathway analysis are presented as source files. c. Proportions of TNF or CXCL10 producing macrophages among alveolar (CD206hiCD68+) macrophages. Unpaired, two-tailed t-test. N = 6 biologically independent mice examined over 3 independent experiments. MISTRG6-hACE2 mice were infected with SARS-CoV-2-mNG and lungs were analysed at 4 d.p.i. d. Distribution of CXCL10 or TNF associated genes at 4, 14, 28 d.p.i. in lungs infected with SARS-CoV-2 or not. Analysis performed on macrophages of 4 d.p.i. lungs in Fig. 3d was extended to more timepoints. Pearson (top) and Spearman (bottom) correlation values were calculated for each gene for its correlation with CXCL10 or TNF in human monocytes and macrophages isolated from uninfected and infected (4, 14 and 28 d.p.i.) lungs of MISTRG6-hACE2 mice. K-means clustering analysis.
Extended Data Fig. 13 SARS-CoV-2 infection of human macrophages activates inflammasomes and leads to death by pyroptosis in vivo and in vitro (matched to figure 3).
a. Representative images of single stained cells for ASC specks, NLRP3, human CD14, human CD45, mouse CD45 and mouse EPCAM. Cells from SARS-CoV-2 infected humanized mice were sorted based on (Extended Data Fig. 6j): human immune cells (hCD45+); mouse immune cells (mCD45+) or epithelial mouse cells (EPCAM+). Sorted cells were stained with single antibodies against ASC, CD14 or NLRP3. Left panel shows human immune cells and right panel shows mouse immune cells (mCD45+) and mouse epithelial cells (EPCAM+). Representative of n = 5 independent mice examined over 3 independent experiments. b. Visualization of ASC specks as a measure of inflammasome activation in mNG+ (SARS-CoV-2+) or mNG− (SARS-CoV-2−) human immune cells at 4 d.p.i. Human immune cells were sorted from SARS-CoV-2 infected humanized mice were sorted based on expression of human CD45 and mNG and lack of mouse CD45 and EPCAM expression (Extended Data Fig. 6j). Representative of n = 5 biologically independent mice examined over 3 independent experiments. c. Left: Representative flow cytometry plot displaying SARS-CoV-2-mNG and Casp1-FLICA staining of CD11b+ human immune cells. Right: quantification of FLICA+ cells (%) as a measure of active caspase-1 in infected (mNG+) and uninfected (mNG−) human lung macrophages (CD11b+hCD45+) at 4 d.p.i. and 14 d.p.i. 4 d.p.i.: n = 3 biologically independent mice examined over 2 independent experiments, 14 d.p.i. n = 5 biologically independent mice examined over 3 independent experiments. Lung cells were incubated with FLICA-Casp1 substrate for 30 min. Means with individual datapoints plotted. Paired, two-tailed t-test. P < 0.0001 = 4.29x10−9. d. Quantification of Casp1-FLICA staining as a measure of active caspase-1 in infected (mNG+) human or total mouse CD11b+ cells at 4 d.p.i. Mouse cells: mCD45+CD11b+hCD45−. Human cells: mCD45−CD11b+hCD45+mNG+. N = 6 biologically independent mice examined over 3 independent experiments. Means with individual datapoints plotted. Paired, two-tailed t-test. P < 0.0001 = 1.79x10−9. e. Human IL-18 (measured by ELISA) in lungs and serum and corresponding mNG levels (measured as percent within human immune cells by flow cytometry) in lungs of infected MISTRG6-hCE2 mice at 4 d.p.i. Lung: Pearson’s correlation value = 0.69. N = 8 biologically independent mice examined over 3 independent experiments. Serum: Pearson’s correlation value = 0.75. n = 7 biologically independent mice examined over 3 independent experiments. Unpaired, two-tailed t-test. f. Human IL-1RA (measured by ELISA) in lungs and serum and corresponding mNG levels (measured as percent within human immune cells by flow cytometry) in lungs of infected MISTRG6-hCE2 mice at 4 d.p.i. Lung: Pearson’s correlation value = 0.82 n = 8 biologically independent mice examined over 3 independent experiments. Serum: Pearson’s correlation value = 0.46 n = 8 biologically independent mice examined over 3 independent experiments. Unpaired t-test, two-tailed. g. LHD levels measured as absorbance at OD 490nm in serum of uninfected or infected MISTRG6-hACE2 mice at 4 d.p.i. and 14 d.p.i. Fresh serum was assayed for LDH. Uninfected n = 3, 4 d.p.i. n = 4, 14 d.p.i. n = 4 biologically independent mice examined over 2 independent experiments. Means with individual datapoints. Unpaired, two-tailed t-test. h. Zombie Aqua incorporation in infected (mNG+) or uninfected (mNG−) CD16+CD11b+ or CD16−CD11b+ human myeloid cells. Frequencies of Zombie+ cells were measured in Annexin V− cells. N = 5 biologically independent mice examined over 2 independent experiments. Means with individual datapoints. Paired, two-tailed t-test. i. Representative histograms and quantification of Casp1-FLICA staining as a measure of active caspase-1 in bone marrow derived macrophages (BMDMs) infected with SARS-CoV-2 in vitro or not for 48 h. BMDM cultures were either supplemented with healthy or COVID plasma or monoclonal antibodies for the duration of the infection. Cultures were treated with Remdesivir, anti-ACE2 and anti-CD16 to block viral replication or viral entry. Colouring on the histograms matches the bar graph legend. Uninfected n = 4; healthy plasma CTRL infected n = 9, anti-ACE2 n = 4, RDV n = 4; mAb n = 4; COVID plasma CTRL infected n = 12, anti-ACE2+anti-CD16 n = 6, RDV n = 5 independent datapoints collected over at least 2 independent experiments. Means with all datapoints. Unpaired, two-tailed t-test. P- values < 0.0001: COVID plasma vs. RDV: 5.85x10−6. j.Representative histograms and quantification of IL-1β in supernatants of BMDMs infected with (or not) SARS-CoV-2 in vitro for 48 or 72 h. Uninfected n = 7, 48 h.p.i. n = 10, 72 h.p.i. n = 5 independent datapoints collected over 3 independent experiments. Means with SD and individual datapoints. Unpaired, two-tailed t-test. P- values < 0.0001: uninfected vs. 72 h.p.i. = 4.96x10−7, 48 h.p.i. vs. 72 h.p.i. = 1.17 x10−8. k. Human IL-18 levels at 48 h.p.i. in supernatants of BMDMs infected or not with SARS-CoV-2 in vitro. BMDM cultures were supplemented with COVID plasma for the duration of the infection. Uninfected n = 4, 48 h.p.i. n = 7 independent datapoints collected over at least 2 independent experiments. Unpaired, two-tailed t-test. Means with individual datapoints. Unpaired, two-tailed t-test. l. Human IL-1RA levels at 48 h.p.i in supernatants of BMDMs infected with SARS-CoV-2 in vitro or not. Uninfected n = 3, infected n = 7 independent datapoints collected over at least 2 independent experiments. Means with individual datapoints. Unpaired, two-tailed t-test. m. Gasdermin D (GSDMD) levels in supernatants of BMDMs infected with (or not) SARS-CoV-2 in vitro for 48 h. BMDM cultures were supplemented with COVID plasma for the duration of the infection. Cultures were treated with Remdesivir to block viral replication. Uninfected N = 3, CTRL infected n = 10, RDV n = 3 independent datapoints collected over at least 2 independent experiments. Means with individual datapoints. Unpaired, two-tailed t-test. n. LHD levels measured by absorbance at OD 490 nm in supernatants of infected or uninfected BMDMs. Uninfected n = 6, Infected n = 11 independent datapoints collected over 3 independent experiments. Means with individual datapoints. Unpaired, two-tailed t-test. P-value = 7.38x10−6. o. Zombie Aqua incorporation in uninfected or SARS-CoV-2-mNG infected BMDMs. Frequencies of Zombie+ cells within Annexin V− population at 48 h.p.i. are presented. Uninfected n = 4, infected n = 7 independent datapoints collected over 3 independent experiments. Means with individual datapoints. Unpaired, two-tailed t-test.
Extended Data Fig. 14 Promoting or blocking viral entry or replication in human macrophages in vivo and in vitro impacts inflammatory profile of macrophages (matched to figure 3).
a. Human IL-18 levels measured in lung homogenates of infected (4 d.p.i.) MISTRG6-hACE2 mice treated (or not) with CD16 blocking antibody (Abcam). CTRL infected n = 7, anti-CD16 n = 5 biologically independent mice examined over 3 independent experiments. Means with datapoints. Unpaired, two-tailed t-test. b. Human IL-1RA levels measured in lung homogenates of infected (4 d.p.i.) MISTRG6-hACE2 mice treated (or not) with CD16 blocking antibody (Abcam). N = 4 biologically independent mice examined over 2 independent experiments. Means with all datapoints. Unpaired, two-tailed t-test. c. Human CXCL10 levels measured in serum of infected (4 d.p.i.) MISTRG6-hACE2 mice treated (or not) with CD16 blocking antibody (Abcam). CTRL infected n = 7, anti-CD16 n = 4 biologically independent mice examined over at least 2 experiments. Means with individual datapoints. Unpaired, two-tailed t-test. d. Human IL-18 levels measured in serum of infected (14 d.p.i.) MISTRG6-hACE2 mice treated (or not) with CD16 blocking antibody (Abcam). Mice were treated with anti-CD16 blocking antibody at 7 d.p.i. and 11 d.p.i. CTRL infected n = 4, anti-CD16 n = 4 biologically independent mice examined over 2 independent experiments. Means with individual datapoints. Unpaired, two-tailed t-test. e. Human IL-18 levels measured in serum of infected (4 d.p.i.) MISTRG6 mice treated (or not) with anti-ACE2 antibody (Abcam). Mice were treated with anti-ACE2 antibody at 1,2,3 d.p.i. Uninfected n = 4, CTRL infected n = 5, anti-ACE2 n = 4 biologically independent mice examined over 2 independent experiments. Means with individual datapoints. Unpaired, two-tailed t-test. P < 0.0001 = uninfected vs. CTRL-infected = 1.43x10−5, CTRL-infected vs. anti-ACE2 = 6.95x10−5. f. Representative flow cytometry plots of CD14 staining on total human immune cells (hCD45+) as a proxy for myeloid cells in infected MISTRG6-hACE2 mice (4 d.p.i.) treated (or not) with a depleting antibody against CD16 (ThermoFisher, clone 3G8). MISTRG6-hACE2 mice were infected with SARS-CoV-2-mNG. Representative of n = 4 biologically independent mice examined over 2 independent experiments. g. Frequencies of mNG+ cells in infected MISTRG6-hACE2 mice at 4 d.p.i. treated (or not) with a depleting antibody against CD16 (ThermoFisher, clone 3G8). N = 4 biologically independent mice examined over 2 independent experiments. Means with datapoints. Unpaired, two-tailed t-test. h. Human IL-18 levels in serum of infected or uninfected MISTRG6-hACE2 mice that were therapeutically treated with monoclonal antibodies (mAb) at 36 h.p.i. or not. Sera from infected mice were analysed at 4 d.p.i. N = 4 biologically independent mice examined over 2 independent experiments. Means with datapoints. Unpaired, two-tailed t-test. i. Human IL-1RA levels in serum of infected (4 d.p.i.) or uninfected MISTRG6-hACE2 mice that were therapeutically treated with mAb at 36 h.p.i. or not. Uninfected and mAb treated n = 4, CTRL infected n = 3 (matched to mAb treatment) biologically independent mice examined over 2 independent experiments. Means with datapoints. Paired, two-tailed t-test. j. Human IL-18 levels measured in serum of infected (14 d.p.i.) MISTRG6-hACE2 mice treated (or not) with mAb (clone 135+ clone 144) at 7 d.p.i. and analysed at 14 d.p.i. N = 3 biologically independent mice examined over 2 independent experiments. Mean with individual datapoints. Unpaired, two-tailed t-test. k. Frequencies of CXCL10+ macrophages within total human macrophages (hCD45+hCD68+) in lungs of infected MISTRG6-hACE2 mice treated (or not) with mAb (clone 135, clone 144) at 7 d.p.i. and analysed at 14 d.p.i. Mean with individual values. Unpaired, two-tailed t-test, not significant (ns). CTRL-infected n = 5, mAb n = 4 biologically independent mice examined over 2 independent experiments. l. Box and whisker plot (min to max, with all datapoints) of the histopathological scoring of the H&E staining of infected MISTRG6-hACE2 lungs at 4 d.p.i. Mice were either treated with monoclonal antibodies at 35 h.p.i. or anti-CD16 at 2 d.p.i. CTRL infected n = 9, mAb treated n = 6, anti-CD16 treated n = 3 biologically independent mice examined over at least 2 independent experiments. The whiskers go down to the smallest value (minimum) and up to the largest value (maximum). The box extends from the 25th to 75th percentiles. The median is shown as a line in the center of the box. Unpaired t-test not significant. m. Box and whisker plot (min to max, with all datapoints) of the histopathological scoring of the H&E staining of infected MISTRG6-hACE2 lungs at 14 d.p.i. Mice were either treated with monoclonal antibodies at 7 d.p.i. or anti-CD16 at 7 and 11 d.p.i. CTRL infected n = 4, mAb treated n = 4, anti-CD16 treated n = 4 biologically independent mice examined over 2 independent experiments. The whiskers go down to the smallest value (minimum) and up to the largest value (maximum). The box extends from the 25th to 75th percentiles. The median is shown as a line in the center of the box. Unpaired, two-tailed t-test, not significant. n. Human IL-18 levels in supernatants of SARS-CoV-2 infected BMDMs treated with anti-CD16 and anti-ACE2 antibodies to block viral entry or with Remdesivir to block viral replication. CTRL infected n = 8, anti-CD16+anti-ACE2 n = 5, RDV n = 5 independent datapoints over 3 independent experiments. Means with all datapoints. Unpaired, two-tailed t-test. P < 0.0001 = 5.0x10−5. o. Human IL-1β levels in supernatants of SARS-CoV-2 infected BMDMs treated with anti-CD16 and anti-ACE2 antibodies to block viral entry or with Remdesivir to block viral replication. N = 4 independent datapoints over 2 independent experiments. Means with all datapoints. Unpaired, two-tailed, t-test. p. Human IL-1RA in supernatants of SARS-CoV-2 infected BMDMs treated with anti-CD16 and anti-ACE2 antibodies to block viral entry or with Remdesivir to block viral replication. CTRL infected n = 7, anti-CD16+anti-ACE2 n = 3, RDV n = 3 independent datapoints over 2 independent experiments. Means with all datapoints. Unpaired, two-tailed, t-test. q. Human CXCL10 levels in supernatants of BMDMs infected in vitro in presence or absence of anti-CD16 and anti-ACE2 antibodies to block viral entry. CTRL infected N = 12, anti-ACE2+ anti-CD16 treated n = 6 independent datapoints over 2 independent experiments. Means with all datapoints. Unpaired, two-tailed, t-test.
Extended Data Fig. 15 Blockade of inflammasome activation leads to reduced cytokine production in vitro (matched to figure 4).
a. Representative flow cytometry plots of CXCL10 and TNF staining in total human macrophages and histograms of CXCL10 expression in infected (mNG+) and uninfected (mNG−) macrophages from lungs of SARS-CoV-2-mNG infected MISTRG6-hACE2 mice treated with caspase-1 (Casp1) or NLRP3 inhibitors in vivo. Mice were treated on days 6,8,10,12 post-infection and analysed at 14 d.p.i. Representative of n = 5 biologically independent mice. b. Mean fluorescent intensity (MFI) of CXCL10 expression in human macrophages isolated from infected MISTRG6-hACE2 mice treated with caspase-1 inhibitor or left untreated. N = 3 biologically independent mice. Representative of 3 independent experiments. Means with all datapoints and SD. Unpaired, two-tailed t-test. c. CXCL10 levels in serum of SARS-CoV-2-mNG infected MISTRG6-hACE2 mice (14 d.p.i.) treated with caspase-1 or NLRP3 inhibitors. N = 4 biologically independent mice examined over 2 independent experiments. Means with all datapoints and SD. Unpaired, two-tailed t-test. d. Frequencies of mNG+ bone marrow-derived macrophages (BMDMs) infected with SARS-CoV-2-mNG in vitro. BMDMs were treated with caspase-1 (Casp1) or NLRP3 inhibitors or left untreated and analysed at 48 h.p.i. CTRL infected n = 22, Casp1 inhibitor-treated n = 17, NLRP3 inhibitor-treated n = 6 independent datapoints collected over at least 3-experiments. Means with all datapoints and SD. Unpaired, two-tailed t-test. e. Frequencies of FLICA+ BMDMs infected with SARS-CoV-2 in vitro for 48 h. BMDMs were treated with caspase-1 or NLRP3 inhibitors or left untreated. CTRL infected n = 13, Casp1 inhibitor-treated n = 12, NLRP3 inhibitor-treated n = 6 independent datapoints collected over at least 3 independent experiments. Means with all datapoints. Unpaired, two-tailed t-test. P < 0.0001 = 3.33x10−5. f. Human IL-18 levels in supernatants of SARS-CoV-2-mNG infected BMDMs treated with caspase-1 (Casp1) inhibitor or left untreated. CTRL infected n = 8, Casp1 inhibitor-treated n = 6 independent datapoints collected over 2 independent experiments. Means with all datapoints and SD. Unpaired, two-tailed t-test. g. Representative histograms and quantification of IL-1β in supernatants of BMDMs infected with SARS-CoV-2 in vitro. Cultures were treated with caspase-1 (Casp1) inhibitor. Uninfected n = 7; 48 h.p.i. CTRL infected n = 10, Casp1 inhibitor-treated n = 9, NLRP3 inhibitor-treated n = 3; 72 h.p.i. CTRL infected n = 5 Casp1 inhibitor-treated n = 3, NLRP3 inhibitor-treated n = 3 independent datapoints collected over at least 2 experiments. Means with all datapoints and SD. Unpaired, two-tailed t-test. P < 0.0001 = 4.96x10−7. h. Human Gasdermin D (GSDMD) levels at 48 h.p.i. in supernatants of SARS-CoV-2-mNG infected BMDMs treated with caspase-1 inhibitor or left untreated. CTRL infected n = 10, Casp1 inhibitor-treated n = 6 independent datapoints collected over at least 3 independent experiments. Means with all datapoints. Unpaired, two-tailed t-test. i. LDH levels measured as absorbance at OD 490nm in supernatants of uninfected or SARS-CoV-2-mNG infected BMDMs treated with Casp1 or NLRP3 inhibitor or left untreated in vitro. Uninfected n = 6, CTRL infected (48 h.p.i.) n = 11, Casp1 inhibitor-treated (48 h.p.i.) n = 9, NLRP3 inhibitor-treated (48 h.p.i.) n = 5 independent datapoints collected over 2 independent experiments. Means with all datapoints and SD. Unpaired, two-tailed t-test. P < 0.0001 = 7.38x10−6. j. Zombie Aqua incorporation in SARS-CoV-2-mNG infected BMDM treated with caspase-1 or NLRP3 inhibitors or left untreated (CTRL infected). Frequencies of Zombie+ cells within Annexin V− population at 48 h.p.i. are reported. Uninfected n = 4, CTRL infected n = 7, Casp1 inhibitor n = 4, NLRP3 inhibitor n = 3 over 2 experiments. Means with all datapoints. Unpaired, two-tailed t-test. k. Human CXCL10 levels in supernatants of infected BMDMs treated with caspase-1 or NLRP3 inhibitors or left untreated. Supernatants were collected at 48 h.p.i. Uninfected n = 5, CTRL infected n = 12, Casp1 inhibitor n = 5, NLRP3 n = 4 independent datapoints over at least 2 independent experiments. Means with all datapoints and SD. Unpaired, two-tailed t-test. l. Human IL-1RA levels in supernatants of SARS-CoV-2-mNG infected BMDMs treated with caspase-1 inhibitor or not. Supernatants were collected at 48 h.p.i. CTRL infected n = 7, Casp1 inhibitor-treated n = 6, NLRP3 inhibitor-treated n = 4 independent datapoints collected over at least 2 independent experiments. Means with all datapoints. Unpaired, two-tailed t-test. m. Viral titres measured as PFU in lung homogenates of MISTRG6-hACE2 mice infected with SARS-CoV-2 and treated with caspase-1 inhibitor in vivo. Infected MISTRG6-hACE2 mice were treated with caspase-1 inhibitor on days 6,8,10,12 post-infection and analysed at 14 d.p.i. Lung homogenates were plaqued using Vero ACE2+TMPRSS2+ cells. of CTRL infected: n = 7, Casp1 inhibitor-treated: n = 6 biologically independent mice examined over 3 independent experiments. Box and whisker plot (min to max, with all datapoints) The whiskers go down to the smallest value (minimum) and up to the largest value (maximum). The box extends from the 25th to 75th percentiles. The median is shown as a line in the center of the box. Ratiopaired, two-tailed t-test. n. Representative images of plaque assays used to quantify infectious virus in supernatants of BMDMs infected with SARS-CoV-2-mNG and treated with caspase-1 or NLRP3 inhibitors. Supernatants of infected macrophage cultures were collected at 48 h.p.i. and plaqued using Vero ACE2+TMPRSS2+ cells. Plaques were resolved at 48 h.p.i. Representative of CTRL infected: n = 13, Casp1 inhibitor-treated: n = 8, NLRP3 inhibitor-treated: n = 5 independent datapoints collected over 3 independent experiments.
Supplementary information
Supplementary Information
Supplementary Discussion, a guide for Supplementary Tables 1–5 and Supplementary References.
Supplementary Table 1
Human genes that are differentially regulated in lungs of infected MISTRG6-hACE2 in response to therapeutics. Genes that are upregulated in response to infection and downregulated in response to therapeutics (dexamethasone, anti-IFNAR + remdesivir) in these infected mice at 14 d.p.i. were included in the analysis (matched to Fig. 1d). Normalized expression of duplicates. n = 2 biologically independent mice examined over two independent experiments. Differential expression analysis was performed using DESeq2 and statistical significance was determined using the Wald test.
Supplementary Table 2
Cluster identifying markers and markers that identify temporal transcriptional changes associated with monocytes and macrophages in infected (4, 14 or 28 d.p.i.) or uninfected lungs of MISTRG6-hACE2 mice (matched to Fig. 1g). n = 2 biologically independent mice for each condition were pooled. Marker genes for each cluster of cells were identified using the Wilcoxon rank-sum tests with Seurat. For the adjusted P values, Bonferroni correction was used.
Supplementary Table 3
Expression of human genes that are enriched in macrophages (clusters identified as part of Fig. 1g) during SARS-Cov-2 infection and their response to anti-IFNAR2 and remdesivir therapy (matched to Fig. 1h). Normalized expression of duplicates was analysed. n = 2 biologically independent mice examined over two independent experiments. Differential expression analysis was performed usingDESeq2 and statistical significance was determined using the Wald test.
Supplementary Table 4
Pearson and Spearman correlation values were calculated for each gene for its correlation with CXCL10, TNF or TLR7 in human monocytes and macrophages at 4 d.p.i. (based on Fig. 1g, matched to Fig. 3d) For the Pearson test, significance was based on the t-test with statistics based on Pearson's product–moment correlation coefficient cor(x, y) and following a t distribution with length(x)-2 d.f. For Spearman's test, P values werecomputed using algorithm AS 89 with exact = TRUE. Correlation values, P values (two-tailed) and FDR-adjusted P values are presented.
Supplementary Table 5
The patient samples used for immunofluorescence staining. Details of patient demographics for specimens use in immunofluorescence staining, including age, gender, medication, time of death post-symptom onset (dps), co-morbidities, cause of death and histopathological findings. This table is presented as part of the Supplementary Methods.
Source data
Rights and permissions
About this article
Cite this article
Sefik, E., Qu, R., Junqueira, C. et al. Inflammasome activation in infected macrophages drives COVID-19 pathology. Nature 606, 585–593 (2022). https://doi.org/10.1038/s41586-022-04802-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04802-1
This article is cited by
-
cGAS-STING, inflammasomes and pyroptosis: an overview of crosstalk mechanism of activation and regulation
Cell Communication and Signaling (2024)
-
ATF3 coordinates the survival and proliferation of cardiac macrophages and protects against ischemia–reperfusion injury
Nature Cardiovascular Research (2024)
-
Single-cell division tracing and transcriptomics reveal cell types and differentiation paths in the regenerating lung
Nature Communications (2024)
-
The role of inflammasomes in human diseases and their potential as therapeutic targets
Signal Transduction and Targeted Therapy (2024)
-
Characteristics of COVID-19 and Impact of Disease Activity in Patients with Adult-Onset Still’s Disease
Rheumatology and Therapy (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.