Abstract
The gut microbiome is associated with diverse diseases1,2,3, but a universal signature of a healthy or unhealthy microbiome has not been identified, and there is a need to understand how genetics, exposome, lifestyle and diet shape the microbiome in health and disease. Here we profiled bacterial composition, function, antibiotic resistance and virulence factors in the gut microbiomes of 8,208 Dutch individuals from a three-generational cohort comprising 2,756 families. We correlated these to 241 host and environmental factors, including physical and mental health, use of medication, diet, socioeconomic factors and childhood and current exposome. We identify that the microbiome is shaped primarily by the environment and cohabitation. Only around 6.6% of taxa are heritable, whereas the variance of around 48.6% of taxa is significantly explained by cohabitation. By identifying 2,856 associations between the microbiome and health, we find that seemingly unrelated diseases share a common microbiome signature that is independent of comorbidities. Furthermore, we identify 7,519 associations between microbiome features and diet, socioeconomics and early life and current exposome, with numerous early-life and current factors being significantly associated with microbiome function and composition. Overall, this study provides a comprehensive overview of gut microbiome and the underlying impact of heritability and exposures that will facilitate future development of microbiome-targeted therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The raw microbiome sequencing data, processed microbiome data (including taxonomy, pathway, virulence factor and antibiotic resistance gene profiles) and basic phenotypes (including age, sex and BMI) used in this study are available at the European Genome-Phenome Archive under accession EGAS00001005027. These datasets can be accessed from https://forms.gle/eHeBdXJMXbVvCJRc8 or by email from the corresponding author (R.K.W.) at the address listed at the EGA data access committee EGAC00001001996. The phenotype data can be requested, for a fee, by filling the application form at https://www.lifelines.nl/researcher/how-to-apply/apply-here. Lifelines will not charge an access fee for controlled access to the full dataset used in the manuscript (including phenotype and sequencing data), for the specific purpose of replication of the results presented in this Article or for further assessment by the reviewers, for a period of three months. Researchers interested in such a replication study or review assessment can contact Lifelines at research@lifelines.nl. Source data are provided with this paper.
Code availability
Open source codes and scripts used for the analyses or figures are available at the GitHub repository (https://github.com/GRONINGEN-MICROBIOME-CENTRE/DMP) and Zenodo (https://doi.org/10.5281/zenodo.5910709). To facilitate the re-use of the codes, the repository also includes example datasets that enable users to test the codes without the need to apply for access to phenotypes.
References
Lynch, S. V. & Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375, 2369–2379 (2016).
Liang, D., Leung, R. K., Guan, W. & Au, W. W. Involvement of gut microbiome in human health and disease: brief overview, knowledge gaps and research opportunities. Gut Pathog. 10, 3 (2018).
Zmora, N., Soffer, E. & Elinav, E. Transforming medicine with the microbiome. Sci. Transl. Med. 11, eaaw1815 (2019).
Garber, K. First microbiome-based drug clears phase III, in clinical trial turnaround. Nat. Rev. Drug Discov. 19, 655–656 (2020).
Zhernakova, A. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352, 565–569 (2016).
Gaulke, C. A. & Sharpton, T. J. The influence of ethnicity and geography on human gut microbiome composition. Nat. Med. 24, 1495–1496 (2018).
Vatanen, T. et al. Genomic variation and strain-specific functional adaptation in the human gut microbiome during early life. Nat. Microbiol. 4, 470–479 (2019).
Berry, D. & Widder, S. Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front. Microbiol. 5, 219 (2014).
Miquel, S. et al. Faecalibacterium prausnitzii and human intestinal health. Curr. Opin. Microbiol. 16, 255–261 (2013).
Huang, R. et al. Metagenome-wide association study of the alterations in the intestinal microbiome composition of ankylosing spondylitis patients and the effect of traditional and herbal treatment. J. Med. Microbiol. 69, 797–805 (2020).
Kandeel, W. A. et al. Impact of Clostridium bacteria in children with autism spectrum disorder and their anthropometric measurements. J. Mol. Neurosci. 70, 897–907 (2020).
Tap, J. et al. Identification of an intestinal microbiota signature associated with severity of irritable bowel syndrome. Gastroenterology 152, 111–123.e8 (2017).
Costea, P. I. et al. Enterotypes in the landscape of gut microbial community composition. Nat. Microbiol. 3, 8–16 (2018).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743 (2016).
GEM Project Research Consortium. Association of host genome with intestinal microbial composition in a large healthy cohort. Nat. Genet. 48, 1413–1417 (2016).
Wang, K. et al. Gut microbiota as prognosis markers for patients with HBV-related acute-on-chronic liver failure. Gut Microbes 13, 1–15 (2021).
Wang, P. et al. Cigarette smoking status alters dysbiotic gut microbes in hypertensive patients. J. Clin. Hypertens. 23, 1431–1446 (2021).
Gupta, V. K. et al. A predictive index for health status using species-level gut microbiome profiling. Nat. Commun. 11, 1–16 (2020).
Bowyer, R. et al. Socioeconomic status and the gut microbiome: a TwinsUK cohort study. Microorganisms 7, 17 (2019).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Vandeputte, D. et al. Stool consistency is strongly associated with gut microbiota richness and composition, enterotypes and bacterial growth rates. Gut 65, 57–62 (2016).
Davenport, E. R. et al. Seasonal variation in human gut microbiome composition. PLoS ONE 9, e90731 (2014).
The Microbiome Quality Control Project Consortium. Assessment of variation in microbial community amplicon sequencing by the Microbiome Quality Control (MBQC) project consortium. Nat. Biotechnol. 35, 1077–1086 (2017).
Toorn, J. E. et al. Seasonal variation of diet quality in a large middle-aged and elderly Dutch population-based cohort. Eur. J. Nutr. 59, 493 (2020).
Tucker, P. & Gilliland, J. The effect of season and weather on physical activity: a systematic review. Public Health 121, 909–922 (2007).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Song, S. J. et al. Cohabiting family members share microbiota with one another and with their dogs. eLife 2, e00458 (2013).
Xie, H. et al. Shotgun metagenomics of 250 adult twins reveals genetic and environmental impacts on the gut microbiome. Cell Syst. 3, 572–584.e3 (2016).
Oh, M. & Zhang, L. DeepMicro: deep representation learning for disease prediction based on microbiome data. Sci. Rep. 10, 6026 (2020).
Zimmermann, M., Zimmermann-Kogadeeva, M., Wegmann, R. & Goodman, A. L. Mapping human microbiome drug metabolism by gut bacteria and their genes. Nature 570, 462–467 (2019).
Imhann, F. et al. Proton pump inhibitors affect the gut microbiome. Gut 65, 740–748 (2016).
Wu, H. et al. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat. Med. 23, 850–858 (2017).
Bach, J. F. The hygiene hypothesis in autoimmunity: the role of pathogens and commensals. Nat. Rev. Immunol. 18, 105–120 (2018).
Marynowski, M. Role of environmental pollution in irritable bowel syndrome. World J. Gastroenterol. 21, 11371 (2015).
Dujardin, C. E. et al. Impact of air quality on the gastrointestinal microbiome: a review. Environ. Res. 186, 109485 (2020).
Fouladi, F. et al. Air pollution exposure is associated with the gut microbiome as revealed by shotgun metagenomic sequencing. Environ. Int. 138, 105604 (2020).
Manor, O. et al. Health and disease markers correlate with gut microbiome composition across thousands of people. Nat. Commun. 11, 5206 (2020).
Sogin, M. L., Morrison, H., McLellan, S., Welch, D. & Huse, S. The rare biosphere: sorting out fact from fiction. Genome Biol. 11, I19 (2010).
Sun, Z. et al. Challenges in benchmarking metagenomic profilers. Nat. Methods 18, 618–626 (2021).
Scholtens, S. et al. Cohort Profile: LifeLines, a three-generation cohort study and biobank. Int. J. Epidemiol. 44, 1172–1180 (2015).
Tigchelaar, E. F. et al. Cohort profile: LifeLines DEEP, a prospective, general population cohort study in the northern Netherlands: study design and baseline characteristics. BMJ Open 5, e006772 (2015).
Siebelink, E., Geelen, A. & de Vries, J. H. M. Self-reported energy intake by FFQ compared with actual energy intake to maintain body weight in 516 adults. Br. J. Nutr. 106, 274–281 (2011).
Brouwer-Brolsma, E. M. et al. A National Dietary Assessment Reference Database (NDARD) for the Dutch population: rationale behind the design. Nutrients 9, 1136 (2017).
Willett, W. C. et al. Reproducibility and validity of a semiquantitative food frequency questionnaire. Am. J. Epidemiol. 122, 51–65 (1985).
Vinke, P. C. et al. Development of the food-based Lifelines Diet Score (LLDS) and its application in 129,369 Lifelines participants. Eur. J. Clin. Nutr. 72, 1111–1119 (2018).
Møller, G. et al. A protein diet score, including plant and animal protein, investigating the association with HbA1c and eGFR—the PREVIEW project. Nutrients 9, 763 (2017).
Leeming, E. R., Johnson, A. J., Spector, T. D. & Le Roy, C. I. Effect of diet on the gut microbiota: rethinking intervention duration. Nutrients 11, 2862 (2019).
Eeftens, M. et al. Development of land use regression models for PM2.5, PM2.5 absorbance, PM10 and PMcoarse in 20 European study areas; results of the ESCAPE project. Environ. Sci. Technol. 46, 11195–11205 (2012).
Beelen, R. et al. Development of NO2 and NOx land use regression models for estimating air pollution exposure in 36 study areas in Europe—the ESCAPE project. Atmos. Environ. 72, 10–23 (2013).
Eeftens, M. et al. Stability of measured and modelled spatial contrasts in NO2 over time. Occup. Environ. Med. 68, 765–770 (2011).
Ford, A. C. et al. Validation of the Rome III criteria for the diagnosis of irritable bowel syndrome in secondary care. Gastroenterology 145, 1262–1270.e1 (2013).
Angulo, P. et al. The NAFLD fibrosis score: a noninvasive system that identifies liver fibrosis in patients with NAFLD. Hepatology 45, 846–854 (2007).
Bedogni, G. et al. The fatty liver index: a simple and accurate predictor of hepatic steatosis in the general population. BMC Gastroenterol. 6, 33 (2006).
Imhann, F. et al. The 1000IBD project: multi-omics data of 1000 inflammatory bowel disease patients; data release 1. BMC Gastroenterol. 19, 5 (2019).
McIver, L. J. et al. bioBakery: a meta’omic analysis environment. Bioinformatics 34, 1235–1237 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Truong, D. T. et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 12, 902–903 (2015).
Franzosa, E. A. et al. Species-level functional profiling of metagenomes and metatranscriptomes. Nat. Methods 15, 962–968 (2018).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45, D158–D169 (2017).
Swertz, M. A. et al. The MOLGENIS toolkit: rapid prototyping of biosoftware at the push of a button. BMC Bioinf. 11, S12 (2010).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Hsieh, T. C., Ma, K. H. & Chao, A. iNEXT: an R package for rarefaction and extrapolation of species diversity (Hill numbers). Methods Ecol. Evol. 7, 1451–1456 (2016).
Kaminski, J. et al. High-specificity targeted functional profiling in microbial communities with ShortBRED. PLoS Comput. Biol. 11, e1004557 (2015).
Chen, L. et al. VFDB: a reference database for bacterial virulence factors. Nucleic Acids Res. 33, D325–D328 (2005).
Jia, B. et al. CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 45, D566–D573 (2017).
Ziyatdinov, A. et al. lme4qtl: linear mixed models with flexible covariance structure for genetic studies of related individuals. BMC Bioinf. 19, 68 (2018).
Aitchison, J. The Statistical Analysis of Compositional Data. J. R. Stat. Soc. Series B Stat. Methodol. 44, 139–177 (1982).
Turnbaugh, P. J. et al. The Human Microbiome Project. Nature 449, 804–810 (2007).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Goodrich, J. K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8, e1002687 (2012).
Chen, L. et al. Gut microbial co-abundance networks show specificity in inflammatory bowel disease and obesity. Nat. Commun. 11, 4018 (2020).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Acknowledgements
We acknowledge and thank the late M. Hofker who initiated the Lifelines DAG3/Dutch Microbiome Project. We acknowledge the services of Lifelines Cohort Study, the contributing research centres delivering data to Lifelines and all the study participants. The Lifelines Biobank initiative has been made possible by subsidies from the Dutch Ministry of Health, Welfare and Sport, the Dutch Ministry of Economic Affairs, the University Medical Center Groningen (UMCG the Netherlands), the University of Groningen and the Northern Provinces of the Netherlands. We thank the Center for Information Technology of the University of Groningen (RUG) for their support and for providing access to the Peregrine high performance computing cluster, the Genomics Coordination Center (UMCG and RUG) for their support and for providing access to Calculon and Boxy high-performance computing clusters and the MOLGENIS team for data management and analysis support. Metagenomics library preparation and sequencing was done at Novogene. We also thank K. Mc Intyre for English and content editing and Tania Ballve Fernandez for illustration of Fig. 1a. Sequencing of the cohort was funded by a grant from CardioVasculair Onderzoek Nederland (CVON 2012-03) to M.H., J.F. and A.Z. R.G., H.J.M.H. and R.K.W. are supported by the collaborative TIMID project (LSHM18057-SGF) financed by the PPP allowance made available by Top Sector Life Sciences & Health to Samenwerkende Gezondheidsfondsen (SGF) to stimulate public–private partnerships and co-financed by health foundations that are part of the SGF. R.K.W. is supported by the Seerave Foundation and the Dutch Digestive Foundation (16-14). A.Z. is supported by European Research Council (ERC) Starting Grant 715772, Netherlands Organization for Scientific Research (NWO) VIDI grant 016.178.056, CVON grant 2018-27 and NWO Gravitation grant ExposomeNL 024.004.017. JF is supported by the Dutch Heart Foundation IN-CONTROL (CVON2018-27), the ERC Consolidator grant (grant agreement No. 101001678), NWO-VICI grant VI.C.202.022, and the Netherlands Organ-on-Chip Initiative, an NWO Gravitation project (024.003.001) funded by the Ministry of Education, Culture and Science of the government of The Netherlands. C.W. is further supported by an ERC advanced grant (ERC-671274) and an NWO Spinoza award (NWO SPI 92-266). L.C. is supported by a joint fellowship from the University Medical Center Groningen and China Scholarship Council (CSC201708320268) and a Foundation De Cock-Hadders grant (20:20-13). M.A.S. is supported by NWO VIDI grant 016 and EUCAN-connect, a project funded by European Commission H2020 grant 824989.
Author information
Authors and Affiliations
Contributions
R.G. designed and implemented the metagenomic data analysis pipelines, analysed metagenomic data, performed heritability analysis and drafted the manuscript. A.K. designed the prediction models and implemented statistical methods for association analyses and assisted in drafting of the manuscript. A.V.V., L.C., V.C., S.H., M.A.Y.K., S.A.-S., J.R.B., L.A.B., V.C.L., T.S., M.H., J.C.S. and S.S. assisted in other statistical analyses, interpretation of data and drafting of the manuscript. M.A.S. provided data stewardship and analysis infrastructure. B.H.J., J.A.M.D., S.J. and J.G.-A. collected data, assisted in study planning and critically reviewed the manuscript. S.S. supervised and coordinated heritability analysis. R.C.H.V. provided the air pollution data and supervised the air pollution analysis. H.J.M.H., A.Z., R.K.W., J.F. and C.W. conceived, coordinated and supported the study. All authors critically revised and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Estimation of total number of species and genera in the DMP population.
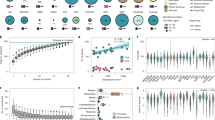
Figure shows rarefaction and extrapolation sampling curve for a, genera and b, species richness calculated using Hill numbers implemented in the iNEXT package for R. The extrapolated part of rarefaction curve is shown dotted. The SD of the estimate is shaded and the asymptotic richness estimate is shown.
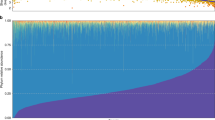
Extended Data Fig. 2 Overview of DMP microbiome composition and function.
a, First two principal coordinates of the Bray-Curtis distance matrix calculated on microbial species of the DMP cohort, coloured by the relative abundance of Prevotella copri bacterium. b, Average relative abundances of bacterial phyla present in > 0.1% of the DMP cohort. Red vertical line indicates rare phyla (abundance < 0.1%). c, Phylum-level composition of all samples in the cohort, sorted by abundance of phylum Bacteroidetes. Each vertical line indicates one sample. * phylum has significantly higher variance when compared to each of pathway classes (one-sided F test of variances FDRs < 0.05, Supplementary Table 1G) d, Relative abundances of the top 10 MetaCyc pathways of all samples (sorted to match panel c). Each vertical line indicates one sample. The means of standard deviations of taxa and pathways were found to be significantly different (mean(sd(tax1),...,sd(taxn)) - mean(sd(pwy1),...,sd(pwnm) > 0, two-sided permutation test (1,000 permutations) P < 1.0 x 10−3). All panels show results generated from n = 8,208 independent samples.
Extended Data Fig. 3 Clusters determined by bi-modally distributed Prevotella copri.
a, Density plots of log2-transformed relative abundances of the 10 most abundant bacterial species. b, Log2-transformed relative abundance of Prevotella copri per microbiome cluster (n (cluster1, red) = 6,346, n (cluster2, blue) = 1,862; boxplot: centre line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range; points, outliers; outer line, distribution of data). The clusters were determined using the partitioning around the medoid method on the relative abundances of microbial species. c. Association of P. copri with metadata (n = 8,208 independent samples; dot, mean; lines, 95% confidence intervals).
Extended Data Fig. 4 Bray-Curtis distances of microbiome features of cohabiting and non-cohabiting participants.
Pairwise microbiome Bray-Curtis dissimilarity comparisons of groups of random, non-cohabiting pairs (RND.PAIR, n = 2,000) compared to cohabitating partners (PARTNERS, n = 1,710); cohabiting parent–child pairs (PAR_CH, n = 285) and cohabiting siblings (SIBL, n = 144); and random pairs (n = 2,000) compared to non-cohabiting 1st-degree relatives (1stDEG.SEP, n = 600) and cohabiting 1st-degree relatives (1stDEG.COH, n = 429). a, MetaCyc pathways. b, Virulence factor gene families. c, antibiotic resistance gene families (centre line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range; points, outliers; outer line: distribution of data). Significantly different groups are marked with ** for FDR < 1.0e–5 or * for FDR < 0.05 (two-sided Wilcoxon test).
Extended Data Fig. 5 Overview of microbiome–phenotype associations.
Figure shows the number of study-wide significant associations (FDR < 0.05) per phenotype group, clustered by taxonomy. Bar heights represent the number of associations relative to the maximal number of associations for the phenotype group.
Extended Data Fig. 6 Gut Microbiome Health Index calculated for DMP cohort.
Box-plots of the Gut Microbiome Health Index (GMHI) for healthy participants of the DMP cohort samples (Y, n = 1,876 independent participants) vs participants who reported one or more diseases (N, n = 6,332 independent participants) (centre line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range; points, outliers; outer line, distribution of data). P-value is shown for two-sided Wilcoxon rank-sum test.
Extended Data Fig. 7 Microbiome associations with diseases and medication use.
Heatmap of microbiome–phenotype associations, with microbial species clustered by Z scores (multivariate linear regression of CLR-transformed relative abundance of taxa, correcting for age, Sex, BMI, Bristol stool scale of the faecal sample and technical factors (DNA concentration, sequencing read depth, sequencing batch and sampling season)) using hierarchical clustering and coloured by the direction of association. Study-wide significant associations (Benjamini-Hochberg corrected p-value < 0.05) are marked with +/−. Coloured associations without a label indicate nominally significant associations (Benjamini-Hochberg corrected p-value < 0.05).
Extended Data Fig. 8 Microbiome association with early-life exposures.
Heatmap of microbiome–phenotype associations, with microbial species clustered by Z scores (multivariate linear regression of CLR-transformed relative abundance of taxa, correcting for age, Sex, BMI, Bristol stool scale of the faecal sample and technical factors (DNA concentration, sequencing read depth, sequencing batch and sampling season)) using hierarchical clustering and coloured by the direction of association. Study-wide significant associations (Benjamini-Hochberg corrected p-value < 0.05) are marked with +/−. Coloured associations without a label indicate nominally significant associations (Benjamini-Hochberg corrected p-value < 0.05).
Extended Data Fig. 9 Microbiome association with smoking, pollutants and greenspace.
Heatmap of microbiome–phenotype associations, with microbial species clustered by Z scores (multivariate linear regression of CLR-transformed relative abundance of taxa, correcting for age, Sex, BMI, Bristol stool scale of the faecal sample and technical factors (DNA concentration, sequencing read depth, sequencing batch and sampling season)) using hierarchical clustering and coloured by the direction of association. Study-wide significant associations (Benjamini-Hochberg corrected p-value < 0.05) are marked with +/−. Coloured associations without a label indicate nominally significant associations (Benjamini-Hochberg corrected p-value < 0.05).
Extended Data Fig. 10 Microbiome association with diet.
Heatmap of microbiome–phenotype associations, with microbial species clustered by Z scores (multivariate linear regression of CLR-transformed relative abundance of taxa, correcting for age, Sex, BMI, Bristol stool scale of the faecal sample and technical factors (DNA concentration, sequencing read depth, sequencing batch and sampling season)) using hierarchical clustering and coloured by the direction of association. Study-wide significant associations (Benjamini-Hochberg corrected p-value < 0.05) are marked with +/−. Coloured associations without a label indicate nominally significant associations (Benjamini-Hochberg corrected p-value < 0.05).
Supplementary information
Supplementary Discussion
Elaborating on the results of microbiome models for prediction of general health and specific diseases in our study; elaborating on the associations of gut microbiome with diseases and medication use in our study; elaborating on diet stability over 5-year time period
Supplementary Table 1
Excel file containing Descriptions of Supplementary Tables (Data_description sheet) and Supplementary Tables 1–10 (in separate excel sheets).
Rights and permissions
About this article
Cite this article
Gacesa, R., Kurilshikov, A., Vich Vila, A. et al. Environmental factors shaping the gut microbiome in a Dutch population. Nature 604, 732–739 (2022). https://doi.org/10.1038/s41586-022-04567-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04567-7
This article is cited by
-
Case report: Aberrant fecal microbiota composition of an infant diagnosed with prolonged intestinal botulism
Gut Pathogens (2024)
-
Gut microbial network signatures of early colonizers in preterm neonates with extrauterine growth restriction
BMC Microbiology (2024)
-
Differences in gut microbiota between Dutch and South-Asian Surinamese: potential implications for type 2 diabetes mellitus
Scientific Reports (2024)
-
A robust microbiome signature for autism spectrum disorder across different studies using machine learning
Scientific Reports (2024)
-
Variant of the lactase LCT gene explains association between milk intake and incident type 2 diabetes
Nature Metabolism (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.