Abstract
The natural habitats of microorganisms in the human microbiome, ocean and soil ecosystems are full of colloids and macromolecules. Such environments exhibit non-Newtonian flow properties, drastically affecting the locomotion of microorganisms1,2,3,4,5. Although the low-Reynolds-number hydrodynamics of swimming flagellated bacteria in simple Newtonian fluids has been well developed6,7,8,9, our understanding of bacterial motility in complex non-Newtonian fluids is less mature10,11. Even after six decades of research, fundamental questions about the nature and origin of bacterial motility enhancement in polymer solutions are still under debate12,13,14,15,16,17,18,19,20,21,22,23. Here we show that flagellated bacteria in dilute colloidal suspensions display quantitatively similar motile behaviours to those in dilute polymer solutions, in particular a universal particle-size-dependent motility enhancement up to 80% accompanied by a strong suppression of bacterial wobbling18,24. By virtue of the hard-sphere nature of colloids, whose size and volume fraction we vary across experiments, our results shed light on the long-standing controversy over bacterial motility enhancement in complex fluids and suggest that polymer dynamics may not be essential for capturing the phenomenon12,13,14,15,16,17,18,19,20,21,22,23. A physical model that incorporates the colloidal nature of complex fluids quantitatively explains bacterial wobbling dynamics and mobility enhancement in both colloidal and polymeric fluids. Our findings contribute to the understanding of motile behaviours of bacteria in complex fluids, which are relevant for a wide range of microbiological processes25 and for engineering bacterial swimming in complex environments26,27.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions herein are available from the University of Minnesota Data Repository: https://doi.org/10.13020/nfr5-te36.
Code availability
The codes used for tracking bacterial swimming speeds and wobble angles are available from the University of Minnesota Data Repository: https://doi.org/10.13020/nfr5-te36.
References
Celli, J. P. et al. Helicobacter pylori moves through mucus by reducing mucin viscoelasticity. Proc. Natl Acad. Sci. USA 106, 14321–14326 (2009).
Suarez, S. S. & Pacey, A. A. Sperm transport in the female reproductive tract. Hum. Reprod. Update 12, 23–37 (2006).
Wells, M. L. & Goldberg, E. D. Occurrence of small colloids in sea water. Nature 353, 342–344 (1991).
Verdugo, P. et al. The oceanic gel phase: a bridge in the DOM–POM continuum. Mar. Chem. 92, 67–85 (2004).
Azam, F. & Malfatti, F. Microbial structuring of marine ecosystems. Nat. Rev. Microbiol. 5, 782–791 (2007).
Lauga, E. & Powers, T. R. The hydrodynamics of swimming microorganisms. Rep. Prog. Phys. 72, 096601 (2009).
Childress, S. Mechanics of Swimming and Flying (Cambridge Univ. Press, 1981).
Berg, H. C. E. coli in Motion (Springer, 2004).
Lauga, E. Bacterial hydrodynamics. Annu. Rev. Fluid Mech. 48, 105–130 (2016).
Elfring, G. J. & Lauga, E. in Complex Fluids in Biological Systems (ed. Spagnolie, S.) 283–317 (Springer, 2015).
Patteson, A. E., Gopinath, A. & Arratia, P. E. Active colloids in complex fluids. Curr. Opin. Colloid Interf. Sci. 21, 86–96 (2016).
Shoesmith, J. G. The measurement of bacterial motility. Microbiology 22, 528–535 (1960).
Schneider, W. R. & Doetsch, R. N. Effect of viscosity on bacterial motility. J. Bacteriol. 117, 696–701 (1974).
Berg, H. C. & Turner, L. Movement of microorganisms in viscous environments. Nature 278, 349–351 (1979).
Magariyama, Y. & Kudo, S. A mathematical explanation of an increase in bacterial swimming speed with viscosity in linear-polymer solutions. Biophys. J. 83, 733–739 (2002).
Martinez, V. A. et al. Flagellated bacterial motility in polymer solutions. Proc. Natl Acad. Sci. USA 111, 17771–17776 (2014).
Zhang, Y., Li, G. & Ardekani, A. M. Reduced viscosity for flagella moving in a solution of long polymer chains. Phys. Rev. Fluids 3, 023101 (2018).
Patteson, A. E., Gopinath, A., Goulian, M. & Arratia, P. E. Running and tumbling with E. coli in polymeric solutions. Sci. Rep. 5, 15761 (2015).
Qu, Z., Temel, F. Z., Henderikx, R. & Breuer, K. S. Changes in the flagellar bundling time account for variations in swimming behavior of flagellated bacteria in viscous media. Proc. Natl Acad. Sci. USA 115, 1707–1712 (2018).
Qu, Z. & Breuer, K. S. Effects of shear-thinning viscosity and viscoelastic stresses on flagellated bacteria motility. Phys. Rev. Fluids 5, 073103 (2020).
Zöttl, A. & Yeomans, J. M. Enhanced bacterial swimming speeds in macromolecular polymer solutions. Nat. Phys. 15, 554–558 (2019).
Binagia, J. P., Phoa, A., Housiadas, K. D. & Shaqfeh, E. S. G. Swimming with swirl in a viscoelastic fluid. J. Fluid Mech. 900, A4 (2020).
Man, Y. & Lauga, E. Phase-separation models for swimming enhancement in complex fluids. Phys. Rev. E 92, 023004 (2015).
Hyon, Y., Marcos, Powers, T. R., Stocker, R. & Fu, H. C. The wiggling trajectories of bacteria. J. Fluid Mech. 705, 58–76 (2012).
Hibbing, M. E., Fuqua, C., Parsek, M. R. & Peterson, S. B. Bacterial competition: surviving and thriving in the microbial jungle. Nat. Rev. Microbiol. 8, 15–25 (2010).
Nelson, B. J., Kaliakatsos, I. K. & Abbott, J. J. Microrobots for minimally invasive medicine. Annu. Rev. Biomed. Eng. 12, 55–85 (2010).
Bechinger, C. et al. Active particles in complex and crowded environments. Rev. Mod. Phys. 88, 045006 (2016).
Peng, Y., Liu, Z. & Cheng, X. Imaging the emergence of bacterial turbulence: phase diagram and transition kinetics. Sci. Adv. 7, eabd1240 (2021).
Liu, Z., Zeng, W., Ma, X. & Cheng, X. Density fluctuations and energy spectra of 3D bacterial suspensions. Soft Matter 17, 10806–10817 (2021).
Lauga, E., DiLuzio, W. R., Whitesides, G. M. & Stone, H. A. Swimming in circles: motion of bacteria near solid boundaries. Biophys. J. 90, 400–412 (2006).
Hiemenz, P. C. & Lodge, T. Polymer Chemistry 2nd edn (CRC Press, 2007).
Darnton, N. C., Turner, L., Rojevsky, S. & Berg, H. C. On torque and tumbling in swimming Escherichia coli. J. Bacteriol. 189, 1756–1764 (2007).
Macosko, C. W. Rheology: Principles, Measurements, and Applications (VCH, 1994).
Jeffrey, D. J. & Onishi, Y. Calculation of the resistance and mobility functions for two unequal rigid spheres in low-Reynolds-number flow. J. Fluid Mech. 139, 261–290 (1984).
Zhang, B. K., Leishangthem, P. K., Ding, Y. & Xu, X. L. An effective and efficient model of the near-field hydrodynamic interactions for active suspensions of bacteria. Proc. Natl Acad. Sci. USA 118, e2100145118 (2021).
Li, G., Tam, L.-K. & Tang, J. X. Amplified effect of Brownian motion in bacterial near-surface swimming. Proc. Natl Acad. Sci. USA 105, 18355–18359 (2008).
Block, S. M., Blair, D. F. & Berg, H. C. Compliance of bacterial flagella measured with optical tweezers. Nature 338, 514–518 (1989).
Berg, H. C. & Brown, D. A. Chemotaxis in Escherichia coli analysed by three-dimensional tracking. Nature 239, 500–504 (1972).
Crenshaw, H. C. A new look at locomotion in microorganisms: rotating and translating. Am. Zool. 36, 608–618 (1996).
Rossi, M., Cicconofri, G., Beran, A., Noselli, G. & DeSimone, A. Kinematics of flagellar swimming in Euglena gracilis: helical trajectories and flagellar shapes. Proc. Natl Acad. Sci. USA 114, 13085–13090 (2017).
Cortese, D. & Wan, K. Y. Control of helical navigation by three-dimensional flagellar beating. Phys. Rev. Lett. 126, 088003 (2021).
Shimogonya, Y. et al. Torque-induced precession of bacterial flagella. Sci. Rep. 5, 18488 (2015).
Poon, W. C. K., Weeks, E. R. & Royall, C. P. On measuring colloidal volume fractions. Soft Matter 8, 21–30 (2012).
Crocker, J. C. & Grier, D. G. Methods of digital video microscopy for colloidal studies. J. Colloid Interf. Sci. 179, 298–310 (1996).
Acknowledgements
We thank P. Agrawal, S. Ghosh, Y. Kim, Z. Liu, R. Patel and G. Pradipta for help with experiments and data analysis and S. Guo, P. Kolliopoulos, E. Lauga, Y. Man and Z. Qu for fruitful discussions. We would also like to acknowledge the late Professor Howard Berg, who answered our questions on E. coli mutant strains. This research is supported by the IPRIME program of University of Minnesota, and by the US National Science Foundation CBET-1702352 and 2028652. X.X. acknowledges the financial support from National Natural Science Foundation of China no. 11974038 and no. U1930402. Portions of this work were conducted in the Minnesota Nano Center, which is supported by the US National Science Foundation through the National Nano Coordinated Infrastructure Network (NNCI) under award number ECCS-2025124.
Author information
Authors and Affiliations
Contributions
S.K., L.F.F. and X.C. designed the research. S.K. performed the experiments. S.K., S.S., L.F.F. and X.C. discussed and analysed the experimental data. X.X. proposed the model and conducted the analytical calculations with input from X.C. P.L. and X.X. performed the simulation of Extended Data Fig. 7. X.C. conceived the project. L.F.F., X.X. and X.C. supervised the project. S.K. and X.C. co-wrote the manuscript. All authors discussed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Paulo Arratia and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Experimental methods.
a, The average swimming speed of bacteria V as a function of time after a bacterial suspension is injected into a PDMS microchannel. V is normalized by the average speed at time t = 0. The vertical dashed line indicates the maximum measurement time of our experiments of 15 min. b, Comparison of the average swimming speed of bacteria V in suspensions of Ficoll 400 of increasing concentrations c from our experiments with that from a previous study. V is normalized by the average swimming speed in the pure buffer V0. c is the weight percentage concentration. Blue circles are from our experiments, whereas orange diamonds are from ref. 16. c. Determination of the intrinsic viscosity of dilute solutions of carboxymethyl cellulose (CMC) of average molecular weight 700,000. The viscosities of the solutions η as a function of polymer concentrations c are extracted from ref. 18. ηs is the solvent viscosity. c is in unit of g/ml. The slope of the linear fit gives [η] = 56,344.1 ml/g.
Extended Data Fig. 3 Anticorrelation between bacterial wobbling and motility enhancement.
a, The swimming speed versus the wobble angle of individual bacteria in pure buffer (black squares) and in a suspension of colloids of radius R = 500 nm and volume fraction ϕ = 4% (blue triangles). The data are obtained by averaging the swimming speed of many bacteria binned over a small range of wobble angles. The raw data for each individual bacteria in the 4% colloidal suspension are also shown as the background (grey triangles). For clarify, we do not show the data of individual bacteria in buffer, which show a similar degree of scattering. V0 = 13 μm/s and θ0 = 45° are the average swimming speed and the average wobble angle of bacteria in buffer, respectively. As a comparison, the average swimming speed versus the average wobble of bacteria in polymer solutions of different concentrations from ref. 18 is also shown (red discs). b, A violin plot showing the probability distribution of the wobble angle of bacteria in buffer and in the 4% colloidal suspension. The interquartile range (IQR) gives the difference between the 75th and 25th percentiles of the data. a shows that at a given wobble angle, the swimming speed of a bacterium is nearly constant, independent whether it swims in buffer or in colloidal suspensions. b shows that the average swimming speed of bacteria increases in colloidal suspensions, because there are more weakly-wobbling bacteria in the suspensions.
Extended Data Fig. 4 Model description and prediction.
a, A schematic showing the 3D helical trajectory of a bacterium. The 2D projection of the 3D trajectory manifests as bacterial wobbling under optical microscopy. The pitch P and the radius Rw of the trajectory are indicated. The velocity of the bacterium tangential to the helical trajectory Vb and the swimming velocity of the bacterium measured in experiments V are also shown. The schematic is not to scale. Rw is comparable to the size of bacteria and much smaller than P for real trajectories (see Extended Data Fig. 5a). The detailed 3D configuration of a bacterium w.r.t. its trajectory is shown in Extended Data Fig. 6. b, A schematic showing the motion of a bacterium in our model. The angular velocity of the bacterial body ωb and the flagellar bundle ωt, as well as the angular velocity of the entire bacterium as a rigid body, ωcm, are shown. ωb, ωt and ωcm are in the same plane, which we define as the ω plane. The misaligned angle between ωb and ωt, α, and the angle of ωcm with respect to the negative z direction, β, are also shown. The wobble angle θ = β – α. The coordinate system used in our model is defined at the upper right corner. c, Wobble angle θ as a function of the misaligned angle α. A maximum wobble angle θmax = 44.5° reaches when αmax = 22.8°. d, The anticorrelation between the normalized bacterial swimming speed V/V0 and the wobble angle θ. Symbols are from our experiments and the solid line is our model prediction. Figure 3b shows the same results using the normalized wobble angle. In the model, we set V = 1.8 V0 at θ = 24° based on experimental observation. Inset: V/V0 versus the misaligned angle α from our model.
Extended Data Fig. 5 Pitch and radius of bacterial helical trajectories.
a, The pitch P and the radius Rw of bacterial helical trajectories as a function of the wobble angle θ. See the definition of P and Rw in Extended Data Fig. 4a. P is shown to the left axis and Rw is shown to the right axis. The limiting pitch of 64 μm can be reached in our model. In comparison, the limiting pitch predicted by the previous minimal model of bacterial wobbling is 4 μm for bacteria with single flagellar bundle24. b, Probability distribution function (PDF) of the pitch of bacterial helical trajectories. Black squares are experimental data extracted from Fig. 2a of ref. 24. The red solid line is our model prediction.
Extended Data Fig. 6 Schematics showing the 3D configuration of a swimming bacterium and its helical trajectory.
a, The top view of the configuration. The left-handed helical trajectory encloses a cylindrical space of radius Rw (the grey region). The angular velocity of the body and the flagellar bundle ωb and ωt are shown. The velocity of the bacterium tangential to the helical trajectory Vb is indicated. Note that ωb and Vb tilt above the paper (the solid arrows), whereas ωt tilts into the paper (the dashed arrow). The average swimming speed of bacteria measured in experiments V is normal to and points out of the paper. The ω plane is normal to the paper as indicated by the purple dashed line. The cross-section of the plane with the helical cylinder has a rectangular shape with the width w < Rw. The viewpoint of b is indicated at the lower right corner. b, The side view of the ω plane. The axis of the helical trajectory is indicated by the vertical dashed line, which is in front of the ω plane above the paper. The projection of Vb along the direction of –ωcm gives V, whereas the projection of Vb along the direction of the bacterial flagellar bundle (–ωt) gives Vbz. The angles α, β and θ are the same as those defined in Extended Data Fig. 4b. The coordinate defined in Extended Data Fig. 4b is reproduced on the lower right. As colloids are depleted from the cylindrical space due to the long-range hydrodynamics (Extended Data Fig. 7), colloids that exert torque on the bacterium via the short-range lubrication interaction are outside the cylindrical space. The effect breaks the symmetric role of colloids around the bacterium, which preferably reduces α and therefore suppresses bacterial wobbling.
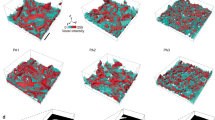
Extended Data Fig. 7 Depletion of colloids from the cylindrical space enclosed by the helical trajectory of a wobbling bacteria.
The radius of colloids is 100 nm (top row), 500 nm (middle row) and 1.5 μm (bottom row). Whereas the 100 nm and 500 nm colloids are free, the large 1.5 μm colloids are pinned in the lab frame to match the condition of the optical tweezer experiments. The small discs indicate the initial positions of colloids when the bacterium is far away from the colloids, where the distance between the bacterium and the colloids along the axis of the helical trajectory –ωcm is lbc ≡ |(rb – rc)·ωcm|/|ωcm| = 6.5 μm. Here, rb and rc are the centre of bacterial body and the centre of the colloids, respectively. The large discs indicate the positions of the corresponding colloids when lbc = 0. The lines connecting the small and large discs represent the projection of the 3D trajectories of colloids on the plane normal to –ωcm in the reference frame of the bacterium, where the bacterial body sits at the origin (0, 0) and orientates in the ω plane indicated by the black dashed lines. The large black circles mark the boundary of the cylindrical space. The radius Rw and the pitch P of the helical trajectory of the bacterium are 1.3 μm and 6.5 μm, respectively. The radius of bacterial body is 1 μm. All the colloids initially located inside the cylindrical space are depleted out of the space as the bacterium approaches the colloids. The results are obtained from numerical simulations with hydrodynamics interactions35, where the interaction between one single colloid and one wobbling bacterium is simulated in each simulation run.
Extended Data Fig. 8 The volume fraction at which the motility enhancement peaks, ϕmax.
\({\varphi }_{{\max }}\) as a function of the radius of colloids or the hydrodynamic radius of polymer coils R. R is normalized by the characteristic radius of bacterial body rb = 1 μm. Except for the datum from ref. 19, all the \({\varphi }_{{\max }}\) are in the dilute regime.
Supplementary information
Supplementary Information
Supplementary text presenting a summary of our experimental finding and a detailed description and derivation of our model.
41586_2022_4509_MOESM3_ESM.mp4
Supplementary Video The swimming of a bacterium near a colloid held in an optical trap. The radius of the colloid is 1.5 μm. The data points on the right show the swimming speed of bacteria (red squares) and the orientation of the bacterium with respect to the local tangential direction of the bacterial trajectory (black squares). Playing at 0.5× speed, the video shows the simultaneous increase of bacterial swimming speed and reduction of bacterial wobbling when the bacterium swims near to the colloid (the green region).
Rights and permissions
About this article
Cite this article
Kamdar, S., Shin, S., Leishangthem, P. et al. The colloidal nature of complex fluids enhances bacterial motility. Nature 603, 819–823 (2022). https://doi.org/10.1038/s41586-022-04509-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04509-3
This article is cited by
-
Chiral active particles are sensitive reporters to environmental geometry
Nature Communications (2024)
-
Emergence of synchronised rotations in dense active matter with disorder
Communications Physics (2023)
-
Orientational dynamics and rheology of active suspensions in weakly viscoelastic flows
Communications Physics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.