Abstract
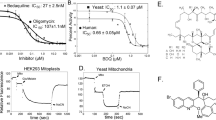
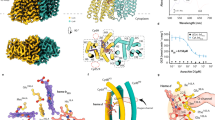
Tuberculosis—the world’s leading cause of death by infectious disease—is increasingly resistant to current first-line antibiotics1. The bacterium Mycobacterium tuberculosis (which causes tuberculosis) can survive low-energy conditions, allowing infections to remain dormant and decreasing their susceptibility to many antibiotics2. Bedaquiline was developed in 2005 from a lead compound identified in a phenotypic screen against Mycobacterium smegmatis3. This drug can sterilize even latent M. tuberculosis infections4 and has become a cornerstone of treatment for multidrug-resistant and extensively drug-resistant tuberculosis1,5,6. Bedaquiline targets the mycobacterial ATP synthase3, which is an essential enzyme in the obligate aerobic Mycobacterium genus3,7, but how it binds the intact enzyme is unknown. Here we determined cryo-electron microscopy structures of M. smegmatis ATP synthase alone and in complex with bedaquiline. The drug-free structure suggests that hook-like extensions from the α-subunits prevent the enzyme from running in reverse, inhibiting ATP hydrolysis and preserving energy in hypoxic conditions. Bedaquiline binding induces large conformational changes in the ATP synthase, creating tight binding pockets at the interface of subunits a and c that explain the potency of this drug as an antibiotic for tuberculosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession numbers EMD-22311 to EMD-22322. Atomic models have been deposited in the Protein Data Bank under accession numbers 7JG5, 7JG6, 7JG7, 7JG8, 7JG9, 7JGA, 7JGB and 7JGC. Strains and plasmids are available from the corresponding author upon reasonable request.
References
World Health Organization. Global Tuberculosis Report 2019 (World Health Organization, 2019).
Lu, P., Lill, H. & Bald, D. ATP synthase in mycobacteria: special features and implications for a function as drug target. Biochim. Biophys. Acta 1837, 1208–1218 (2014).
Andries, K. et al. A diarylquinoline drug active on the ATP synthase of Mycobacterium tuberculosis. Science 307, 223–227 (2005).
Koul, A. et al. Diarylquinolines are bactericidal for dormant mycobacteria as a result of disturbed ATP homeostasis. J. Biol. Chem. 283, 25273–25280 (2008).
World Health Organization. The Selection and Use of Essential Medicines (World Health Organization, 2019).
Médecins Sans Frontières. MSF press release, https://www.msf.org.za/stories-news/press-releases/fix-patent-laws-campaign-launches-global-campaign-urging-johnson-johnson (2019).
McNeil, M. B., Ryburn, H. W. K., Harold, L. K., Tirados, J. F. & Cook, G. M. Transcriptional inhibition of the F1F0-type ATP synthase has bactericidal consequences on the viability of mycobacteria. Antimicrob. Agents Chemother. 64, e00492-20 (2020).
Zhao, J., Benlekbir, S. & Rubinstein, J. L. Electron cryomicroscopy observation of rotational states in a eukaryotic V-ATPase. Nature 521, 241–245 (2015).
Guo, H., Suzuki, T. & Rubinstein, J. L. Structure of a bacterial ATP synthase. eLife 8, e43128 (2019).
Preiss, L. et al. Structure of the mycobacterial ATP synthase Fo rotor ring in complex with the anti-TB drug bedaquiline. Sci. Adv. 1, e1500106 (2015).
Zhou, A. et al. Structure and conformational states of the bovine mitochondrial ATP synthase by cryo-EM. eLife 4, e10180 (2015).
Guo, H., Bueler, S. A. & Rubinstein, J. L. Atomic model for the dimeric FO region of mitochondrial ATP synthase. Science 358, 936–940 (2017).
Hahn, A., Vonck, J., Mills, D. J., Meier, T. & Kuhlbrandt, W. Structure, mechanism, and regulation of the chloroplast ATP synthase. Science 360, eaat4318 (2018).
Sobti, M. et al. Cryo-EM structures provide insight into how E. coli F1Fo ATP synthase accommodates symmetry mismatch. Nat. Commun. 11, 2615 (2020).
Gajadeera, C. S. & Weber, J. Escherichia coli F1Fo-ATP synthase with a b/δ fusion protein allows analysis of the function of the individual b subunits. J. Biol. Chem. 288, 26441–26447 (2013).
Sobti, M. et al. Cryo-EM structures of the autoinhibited E. coli ATP synthase in three rotational states. eLife 5, e21598 (2016).
Abrahams, J. P., Leslie, A. G., Lutter, R. & Walker, J. E. Structure at 2.8 Å resolution of F1-ATPase from bovine heart mitochondria. Nature 370, 621–628 (1994).
Shirakihara, Y. et al. Structure of a thermophilic F1-ATPase inhibited by an ε-subunit: deeper insight into the ε-inhibition mechanism. FEBS J. 282, 2895–2913 (2015).
Cingolani, G. & Duncan, T. M. Structure of the ATP synthase catalytic complex (F1) from Escherichia coli in an autoinhibited conformation. Nat. Struct. Mol. Biol. 18, 701–707 (2011).
Zhang, A. T., Montgomery, M. G., Leslie, A. G. W., Cook, G. M. & Walker, J. E. The structure of the catalytic domain of the ATP synthase from Mycobacterium smegmatis is a target for developing antitubercular drugs. Proc. Natl Acad. Sci. USA 116, 4206–4211 (2019).
Ragunathan, P. et al. The uniqueness of subunit α of mycobacterial F-ATP synthases: an evolutionary variant for niche adaptation. J. Biol. Chem. 292, 11262–11279 (2017).
Lutter, R. et al. F1Fo-ATP synthase from bovine heart mitochondria: development of the purification of a monodisperse oligomycin-sensitive ATPase. Biochem. J. 295, 799–806 (1993).
Koul, A. et al. Diarylquinolines target subunit c of mycobacterial ATP synthase. Nat. Chem. Biol. 3, 323–324 (2007).
Hards, K. et al. Bactericidal mode of action of bedaquiline. J. Antimicrob. Chemother. 70, 2028–2037 (2015).
Hards, K. et al. Ionophoric effects of the antitubercular drug bedaquiline. Proc. Natl Acad. Sci. USA 115, 7326–7331 (2018).
Haagsma, A. C. et al. Probing the interaction of the diarylquinoline TMC207 with its target mycobacterial ATP synthase. PLoS ONE 6, e23575 (2011).
Kundu, S., Biukovic, G., Grüber, G. & Dick, T. Bedaquiline targets the ε subunit of mycobacterial F-ATP synthase. Antimicrob. Agents Chemother. 60, 6977–6979 (2016).
Lounis, N. et al. R207910 (TMC207): un nouvel antibiotique pour le traitement de la tuberculose. Med. Mal. Infect. 40, 383–390 (2010).
Guillemont, J., Meyer, C., Poncelet, A., Bourdrez, X. & Andries, K. Diarylquinolines, synthesis pathways and quantitative structure–activity relationship studies leading to the discovery of TMC207. Future Med. Chem. 3, 1345–1360 (2011).
Sutherland, H. S. et al. 3,5-Dialkoxypyridine analogues of bedaquiline are potent antituberculosis agents with minimal inhibition of the hERG channel. Bioorg. Med. Chem. 27, 1292–1307 (2019).
Haagsma, A. C. et al. Selectivity of TMC207 towards mycobacterial ATP synthase compared with that towards the eukaryotic homologue. Antimicrob. Agents Chemother. 53, 1290–1292 (2009).
Murphy, K. C. et al. ORBIT: a new paradigm for genetic engineering of mycobacterial chromosomes. MBio 9, e01467-18 (2018).
Vasanthakumar, T. et al. Structural comparison of the vacuolar and Golgi V-ATPases from Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 116, 7272–7277 (2019).
Kornberg, A. & Pricer, W. E. Jr. Enzymatic phosphorylation of adenosine and 2,6-diaminopurine riboside. J. Biol. Chem. 193, 481–495 (1951).
Russo, C. J. & Passmore, L. A. Ultrastable gold substrates for electron cryomicroscopy. Science 346, 1377–1380 (2014).
Meyerson, J. R. et al. Self-assembled monolayers improve protein distribution on holey carbon cryo-EM supports. Sci. Rep. 4, 7084 (2014).
Marr, C. R., Benlekbir, S. & Rubinstein, J. L. Fabrication of carbon films with ∼500 nm holes for cryo-EM with a direct detector device. J. Struct. Biol. 185, 42–47 (2014).
Tivol, W. F., Briegel, A. & Jensen, G. J. An improved cryogen for plunge freezing. Microsc. Microanal. 14, 375–379 (2008).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single particle cryo-EM reconstruction. Preprint at https://doi.org/10.1101/2019.12.15.877092 (2019).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Ramírez-Aportela, E. et al. Automatic local resolution-based sharpening of cryo-EM maps. Bioinformatics 36, 765–772 (2020).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D 74, 519–530 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Acknowledgements
We thank V. Kanelis for suggestions on the manuscript. H.G. was supported by an Ontario Graduate Scholarship for International Students. G.M.C. was supported by a W. P. Caven Memorial Fellowship. J.L.R. was supported by the Canada Research Chairs programme. This work was supported by Canadian Institutes of Health Research grant PJT162186 (J.L.R.) and PJT156261 (J.L.). Cryo-EM data were collected at the Toronto High-Resolution High-Throughput cryo-EM facility, supported by the Canada Foundation for Innovation and Ontario Research Fund.
Author information
Authors and Affiliations
Contributions
J.L.R. conceived the project and supervised the research. J.L. advised on M. smegmatis cell culture and chromosomal tagging strategies. S.A.B. prepared the 3×FLAG ORBIT payload plasmid and an M. smegmatis strain used in the initial stages of the project, with help from J.M. G.M.C. prepared the M. smegmatis strains used for structure determination and enzyme assays. G.M.C. and H.G. purified the protein, H.G. performed the cryo-EM and image analysis, and G.M.C. and H.G. constructed the atomic models. J.L.R., H.G. and G.M.C. wrote the manuscript and prepared figures with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Clifton Barry, Damian Ekiert and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Workflow for cryo-EM image analysis for BDQ-free, BDQ-saturated and -washed M. smegmatis ATP synthase.
a, b, Example micrograph (a) and 2D class average images (b) for BDQ-saturated M. smegmatis ATP synthase. Example particles are circled in white. We collected 7,691 movies (including 1,435,679 particle images) for the drug-free condition, 7,589 movies (including 1,825,672 particle images) for the BDQ-saturated condition and 4,962 movies (including 1,865,961 particle images) for the BDQ-washed condition. c, d, Workflow for obtaining maps of the three rotational states (c) and FO region (d).
Extended Data Fig. 2 Cryo-EM map validation.
a–d, Corrected Fourier shell correlation curves after gold-standard refinement, orientation distribution plots and local resolution maps are shown for maps of the entire complex from the BDQ-free dataset (a), BDQ-saturated dataset (b) and BDQ-washed dataset (c), as well as FO-region maps from focused refinement (d).
Extended Data Fig. 3 Examples of model-in-map fit to demonstrate map quality.
a, b, Examples are shown from each subunit in the BDQ-free model (a) and BDQ-saturated model (b).
Extended Data Fig. 4 Comparison of subunits a and b–δ from different organisms.
a, The Bacillus PS3 subunit a is in grey and the M. smegmatis subunit a is in green. b, The E. coli subunit a is in purple and the M. smegmatis subunit a is in green. c, The N-terminal domain of the δ region of the b–δ fusion protein (right side, top) is substantially larger than the N-terminal domain of Bacillus PS3 δ-subunit (right side, middle), being composed of about 270 residues instead of about 170 residues. The extra sequence contributes 5 additional α-helices to the 6 found in the Bacillus PS3 δ-subunit, giving a total of 11 α-helices in the N-terminal domain of b–δ (Fig. 1b, bottom). Mycobacterium smegmatis b–δ and Bacillus PS3 δ also form different interactions with the N-terminal sequences of the α-subunits that help to attach the peripheral stalk to F1 region (right side, red).
Extended Data Fig. 5 Two conformations of the α-extension are seen in rotational state 2.
Density for the α-extension in its predominant conformation is coloured pink, and the alternative conformation is coloured red.
Extended Data Fig. 6 The BDQ-washed structures are in the BDQ-saturated conformation.
Fit of the BDQ-saturated protein models in the BDQ-washed maps shows that the two preparations adopt the same conformation.
Supplementary information
Supplementary Table 1
Cryo-EM data collection, refinement and validation statistics. This file contains Supplementary Tables 2-4 and Supplementary Figures 1-2.
Video 1
Release of the inhibitory α extension during ATP synthesis. The video shows interpolation between the different rotational states of the enzyme, including the alternative conformation of rotational state 2, which suggests the sequence of events that occur during release of the α-extension during ATP synthesis and binding of the α-extension to subunit γ to inhibit ATP hydrolysis. The video may be viewed as a loop.
Video 2
Binding of BDQ to the mycobacterial ATP synthase. The video shows interpolation between the conformation of the ATP synthase in the BDQ-free and BDQ-bound states. BDQ in the leading, c-only, and lagging binding sites are coloured pink, yellow, and blue, respectively. Phe69 in subunit c and Phe221 in subunit a, which undergo large conformational changes upon drug binding, are shown as space-filling models. The video may be viewed as a loop.
Rights and permissions
About this article
Cite this article
Guo, H., Courbon, G.M., Bueler, S.A. et al. Structure of mycobacterial ATP synthase bound to the tuberculosis drug bedaquiline. Nature 589, 143–147 (2021). https://doi.org/10.1038/s41586-020-3004-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-3004-3
This article is cited by
-
Mitochondrial dysfunction induced by bedaquiline as an anti-Toxoplasma alternative
Veterinary Research (2023)
-
Changes within the central stalk of E. coli F1Fo ATP synthase observed after addition of ATP
Communications Biology (2023)
-
Molecular mechanism on forcible ejection of ATPase inhibitory factor 1 from mitochondrial ATP synthase
Nature Communications (2023)
-
Site-directed mutagenesis of Mycobacterium tuberculosis and functional validation to investigate potential bedaquiline resistance-causing mutations
Scientific Reports (2023)
-
Coordinated conformational changes in the V1 complex during V-ATPase reversible dissociation
Nature Structural & Molecular Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.