Abstract
A fundamental challenge in developing treatments for autism spectrum disorders is the heterogeneity of the condition. More than one hundred genetic mutations confer high risk for autism, with each individual mutation accounting for only a small fraction of cases1,2,3. Subsets of risk genes can be grouped into functionally related pathways, most prominently those involving synaptic proteins, translational regulation, and chromatin modifications. To attempt to minimize this genetic complexity, recent therapeutic strategies have focused on the neuropeptides oxytocin and vasopressin4,5,6, which regulate aspects of social behaviour in mammals7. However, it is unclear whether genetic risk factors predispose individuals to autism as a result of modifications to oxytocinergic signalling. Here we report that an autism-associated mutation in the synaptic adhesion molecule Nlgn3 results in impaired oxytocin signalling in dopaminergic neurons and in altered behavioural responses to social novelty tests in mice. Notably, loss of Nlgn3 is accompanied by a disruption of translation homeostasis in the ventral tegmental area. Treatment of Nlgn3-knockout mice with a new, highly specific, brain-penetrant inhibitor of MAP kinase-interacting kinases resets the translation of mRNA and restores oxytocin signalling and social novelty responses. Thus, this work identifies a convergence between the genetic autism risk factor Nlgn3, regulation of translation, and oxytocinergic signalling. Focusing on such common core plasticity elements might provide a pragmatic approach to overcoming the heterogeneity of autism. Ultimately, this would enable mechanism-based stratification of patient populations to increase the success of therapeutic interventions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw mass spectrometry data associated with the manuscript have been deposited in to the ProteomeXchange Consortium via the PRIDE60 partner repository with the dataset identifier PXD018808 and 10.6019/PXD018808. All renewable reagents and detailed protocols will be made available on request. Source data are provided with this paper.
References
de la Torre-Ubieta, L., Won, H., Stein, J. L. & Geschwind, D. H. Advancing the understanding of autism disease mechanisms through genetics. Nat. Med. 22, 345–361 (2016).
O’Roak, B. J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012).
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012).
Yamasue, H., Aran, A. & Berry-Kravis, E. Emerging pharmacological therapies in fragile X syndrome and autism. Curr. Opin. Neurol. 32, 635–640 (2019).
Bolognani, F. et al. A phase 2 clinical trial of a vasopressin V1a receptor antagonist shows improved adaptive behaviors in men with autism spectrum disorder. Sci. Transl. Med. 11, eaat7838 (2019).
Parker, K. J. et al. A randomized placebo-controlled pilot trial shows that intranasal vasopressin improves social deficits in children with autism. Sci. Transl. Med. 11, eaau7356 (2019).
Walum, H. & Young, L. J. The neural mechanisms and circuitry of the pair bond. Nat. Rev. Neurosci. 19, 643–654 (2018).
Dölen, G., Darvishzadeh, A., Huang, K. W. & Malenka, R. C. Social reward requires coordinated activity of nucleus accumbens oxytocin and serotonin. Nature 501, 179–184 (2013).
Cataldo, I., Azhari, A. & Esposito, G. A review of oxytocin and arginine-vasopressin receptors and their modulation of autism spectrum disorder. Front. Mol. Neurosci. 11, 27 (2018).
Ferguson, J. N. et al. Social amnesia in mice lacking the oxytocin gene. Nat. Genet. 25, 284–288 (2000).
Oettl, L. L. et al. Oxytocin enhances social recognition by modulating cortical control of early olfactory processing. Neuron 90, 609–621 (2016).
Hung, L. W. et al. Gating of social reward by oxytocin in the ventral tegmental area. Science 357, 1406–1411 (2017).
Xiao, L., Priest, M. F., Nasenbeny, J., Lu, T. & Kozorovitskiy, Y. Biased oxytocinergic modulation of midbrain dopamine systems. Neuron 95, 368–384 (2017).
Sgritta, M. et al. Mechanisms underlying microbial-mediated changes in social behavior in mouse models of autism spectrum disorder. Neuron 101, 246–259 (2019).
Peñagarikano, O. et al. Exogenous and evoked oxytocin restores social behavior in the Cntnap2 mouse model of autism. Sci. Transl. Med. 7, 271ra8 (2015).
Ebert, D. H. & Greenberg, M. E. Activity-dependent neuronal signalling and autism spectrum disorder. Nature 493, 327–337 (2013).
Sanders, S. J. et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron 70, 863–885 (2011).
Gilman, S. R. et al. Rare de novo variants associated with autism implicate a large functional network of genes involved in formation and function of synapses. Neuron 70, 898–907 (2011).
Tanaka, K. F. et al. Flexible Accelerated STOP Tetracycline Operator-knockin (FAST): a versatile and efficient new gene modulating system. Biol. Psychiatry 67, 770–773 (2010).
Ichtchenko, K., Nguyen, T. & Südhof, T. C. Structures, alternative splicing, and neurexin binding of multiple neuroligins. J. Biol. Chem. 271, 2676–2682 (1996).
Budreck, E. C. & Scheiffele, P. Neuroligin-3 is a neuronal adhesion protein at GABAergic and glutamatergic synapses. Eur. J. Neurosci. 26, 1738–1748 (2007).
Chih, B., Afridi, S. K., Clark, L. & Scheiffele, P. Disorder-associated mutations lead to functional inactivation of neuroligins. Hum. Mol. Genet. 13, 1471–1477 (2004).
Baudouin, S. J. et al. Shared synaptic pathophysiology in syndromic and nonsyndromic rodent models of autism. Science 338, 128–132 (2012).
Rothwell, P. E. et al. Autism-associated neuroligin-3 mutations commonly impair striatal circuits to boost repetitive behaviors. Cell 158, 198–212 (2014).
Tabuchi, K. et al. A neuroligin-3 mutation implicated in autism increases inhibitory synaptic transmission in mice. Science 318, 71–76 (2007).
Radyushkin, K. et al. Neuroligin-3-deficient mice: model of a monogenic heritable form of autism with an olfactory deficit. Genes Brain Behav. 8, 416–425 (2009).
Cao, W. et al. Gamma oscillation dysfunction in mPFC leads to social deficits in neuroligin 3 R451C knockin mice. Neuron 97, 1253–1260 (2018).
Modi, B. et al. Possible implication of the CA2 hippocampal circuit in social cognition deficits observed in the neuroligin 3 knock-out mouse, a non-syndromic animal model of autism. Front. Psychiatry 10, 513 (2019).
Chadman, K. K. et al. Minimal aberrant behavioral phenotypes of neuroligin-3 R451C knockin mice. Autism Res. 1, 147–158 (2008).
Jaramillo, T. C. et al. Genetic background effects in Neuroligin-3 mutant mice: Minimal behavioral abnormalities on C57 background. Autism Res. 11, 234–244 (2017).
Bariselli, S. et al. Role of VTA dopamine neurons and neuroligin 3 in sociability traits related to nonfamiliar conspecific interaction. Nat. Commun. 9, 3173 (2018).
Hitti, F. L. & Siegelbaum, S. A. The hippocampal CA2 region is essential for social memory. Nature 508, 88–92 (2014).
Gunaydin, L. A. et al. Natural neural projection dynamics underlying social behavior. Cell 157, 1535–1551 (2014).
Xu, J. et al. Neuroligin 3 regulates dendritic outgrowth by modulating Akt/mTOR signaling. Front. Cell. Neurosci. 13, 518 (2019).
Dieterich, D. C. et al. In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons. Nat. Neurosci. 13, 897–905 (2010).
Gkogkas, C. G. et al. Autism-related deficits via dysregulated eIF4E-dependent translational control. Nature 493, 371–377 (2013).
Barnes, S. A. et al. Convergence of hippocampal pathophysiology in Syngap +/− and Fmr1 −/y mice. J. Neurosci. 35, 15073–15081 (2015).
Auerbach, B. D., Osterweil, E. K. & Bear, M. F. Mutations causing syndromic autism define an axis of synaptic pathophysiology. Nature 480, 63–68 (2011).
Darnell, J. C. & Klann, E. The translation of translational control by FMRP: therapeutic targets for FXS. Nat. Neurosci. 16, 1530–1536 (2013).
Asiminas, A. et al. Sustained correction of associative learning deficits after brief, early treatment in a rat model of Fragile X Syndrome. Sci. Transl. Med. 11, eaao0498 (2019).
Gkogkas, C. G. et al. Pharmacogenetic inhibition of eIF4E-dependent Mmp9 mRNA translation reverses fragile X syndrome-like phenotypes. Cell Rep. 9, 1742–1755 (2014).
Richter, J. D., Bassell, G. J. & Klann, E. Dysregulation and restoration of translational homeostasis in fragile X syndrome. Nat. Rev. Neurosci. 16, 595–605 (2015).
Bramham, C. R., Jensen, K. B. & Proud, C. G. Tuning specific translation in cancer metastasis and synaptic memory: control at the MNK-eIF4E axis. Trends Biochem. Sci. 41, 847–858 (2016).
Yang, H. et al. Optimization of selective mitogen-activated protein kinase interacting kinases 1 and 2 inhibitors for the treatment of blast crisis leukemia. J. Med. Chem. 61, 4348–4369 (2018).
Lavallée-Adam, M., Rauniyar, N., McClatchy, D. B. & Yates, J. R. III. PSEA-Quant: a protein set enrichment analysis on label-free and label-based protein quantification data. J. Proteome Res. 13, 5496–5509 (2014).
Ueda, T. et al. Combined deficiency for MAP kinase-interacting kinase 1 and 2 (Mnk1 and Mnk2) delays tumor development. Proc. Natl Acad. Sci. USA 107, 13984–13990 (2010).
Xu, Y. et al. Translation control of the immune checkpoint in cancer and its therapeutic targeting. Nat. Med. 25, 301–311 (2019).
The Dutch-Belgian Fragile X Consortium. Fmr1 knockout mice: a model to study fragile X mental retardation. Cell 78, 23–33 (1994).
El-Brolosy, M. A. et al. Genetic compensation triggered by mutant mRNA degradation. Nature 568, 193–197 (2019).
Turiault, M. et al. Analysis of dopamine transporter gene expression pattern — generation of DAT-iCre transgenic mice. FEBS J. 274, 3568–3577 (2007).
Weigelt, S., Koldewyn, K. & Kanwisher, N. Face identity recognition in autism spectrum disorders: a review of behavioral studies. Neurosci. Biobehav. Rev. 36, 1060–1084 (2012).
McPartland, J. C., Webb, S. J., Keehn, B. & Dawson, G. Patterns of visual attention to faces and objects in autism spectrum disorder. J. Autism Dev. Disord. 41, 148–157 (2011).
Dantzer, R., Bluthe, R. M., Koob, G. F. & Le Moal, M. Modulation of social memory in male rats by neurohypophyseal peptides. Psychopharmacology (Berl.) 91, 363–368 (1987).
Mitre, M., Minder, J., Morina, E. X., Chao, M. V. & Froemke, R. C. Oxytocin Modulation of Neural Circuits. Curr. Top. Behav. Neurosci. 35, 31–53 (2018).
Tom Dieck, S. et al. Metabolic labeling with noncanonical amino acids and visualization by chemoselective fluorescent tagging. Curr. Protoc. Cell Biol. Chapter 7, Unit7.11 (2012).
Bowling, H. et al. BONLAC: A combinatorial proteomic technique to measure stimulus-induced translational profiles in brain slices. Neuropharmacology 100, 76–89 (2016).
Peterson, A. C., Russell, J. D., Bailey, D. J., Westphall, M. S. & Coon, J. J. Parallel reaction monitoring for high resolution and high mass accuracy quantitative, targeted proteomics. Mol. Cell. Proteomics 11, 1475–1488 (2012).
Ahrné, E. et al. Evaluation and improvement of quantification accuracy in isobaric mass tag-based protein quantification experiments. J. Proteome Res. 15, 2537–2547 (2016).
Wang, Y. et al. Reversed-phase chromatography with multiple fraction concatenation strategy for proteome profiling of human MCF10A cells. Proteomics 11, 2019–2026 (2011).
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44 (D1), D447–D456 (2016).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47 (D1), D607–D613 (2019).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One 6, e21800 (2011).
DiBenedictis, B. T., Nussbaum, E. R., Cheung, H. K. & Veenema, A. H. Quantitative mapping reveals age and sex differences in vasopressin, but not oxytocin, immunoreactivity in the rat social behavior neural network. J. Comp. Neurol. 525, 2549–2570 (2017).
Acknowledgements
We are grateful to A. Gomez, K. Tan, L. Traunmüller and F. Filice for comments on the manuscript and to members of the Scheiffele laboratory for discussions. We thank B. Boury Jamot for performing tests on Fmr1KO mice, F. Filice for experimental support, and A. Schmidt and the Biozentrum Proteomics Core Facility for conducting proteomics analysis. Funding: H.H. was supported by a Long-term Fellowship from the Human Frontiers Science Program. This work was supported by funds to P.S. from the Swiss National Science Foundation, a European Research Council Advanced Grant (SPLICECODE), EU-AIMS and AIMS-2-TRIALS which are supported by the Innovative Medicines Initiatives from the European Commission. The results leading to this publication has received funding from the Innovative Medicines Initiative 2 Joint Undertaking under grant agreement no. 777394. This Joint Undertaking receives support from the European Union’s Horizon 2020 research and innovation programme and EFPIA and AUTISM SPEAKS, Autistica, SFARI. The Scheiffele Laboratory is an associate member of the Swiss National Science Foundation’s National Competence Centre for Research (NCCR) RNA and Disease. F.M. was supported by NCCR SYNAPSY. Discovery of ETC-168 was financially supported by Biomedical Sciences Institutes (BMSI) and Joint Council Office (JCO Project 11 03 FG 07 05), Agency for Science, Technology and Research (A*STAR), Singapore.
Author information
Authors and Affiliations
Contributions
This work was jointly conceived by H.H. and P.S. and built on initial findings by S.B. and P.S. MNK inhibitors were developed by A.M. and K.N.; behavioural assays were developed by H.H. and F.M.; experimental procedures were performed by H.H., D.S., F.M., L.H.-B., S.B. and E.P.-G.; and data analysis was conducted by H.H., D.S., E.P.-G., F.M., P.S. and E.P.-V. The manuscript was jointly written by H.H. and P.S., with editing provided by E.-P.G., E.P.-V., S.B. and K.N.
Corresponding author
Ethics declarations
Competing interests
S.B. P.S. A.M. and K.N. have filed patents on the use of MNK inhibitors for treatment of neurodevelopmental disorders. A.M. and K.N. are current or former employees of the Experimental Drug Development Centre Singapore which has a commercial interest in the development of MNK1/2 inhibitors.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Loss of social recognition in Nlgn3KO mice.
a, b, Mean social interaction time and data for individual mice in the social habituation/recognition test plotted for wild-type (n = 11) (a) and Nlgn3KO (b) mice (n = 12). c, d, Mean social interaction time and data for individual mice plotted for DAT-cre control mice (n = 10) (c) and DAT-cre::Nlgn3KO mice (n = 11) (d). e, Example for validation of targeted gene knockdown (n = 8 mice) from AAV2-DIO-miRNlgn3-GFP viruses (green) in TH-positive cells (red) in the VTA of DAT-cre mice. f, Quantification of percentage of TH-positive cells in VTA and SNc of DAT-cre mice that express GFP from the AAV2-DIO-miR-GFP vector (n = 8). g, h, Mean social interaction time and data for individual mice plotted for control mice (g, VTA::DA-miR, n = 10) and VTA DA-specific Nlgn3 loss-of-function (h, VTA::DA-NL3, n = 8) in the social habituation/recognition test. g, Mean social interaction time and data for individual mice plotted for mice treated with vehicle (n = 12) (i) and OXTR-A (n = 11) (j). All error bars are s.e.m. Repeated-measures one-way ANOVA followed by Bonferroni’s post hoc test for planned multiple comparison (a–c, g–j) or Friedman test followed by Dunn’s post hoc test for planned multiple comparison (d). See Supplementary information for additional statistics.
Extended Data Fig. 2 Properties of NAc-projecting VTA DA neurons in wild-type and Nlgn3KO mice.
a, Representative Ih currents recorded from wild-type (black) and Nlgn3KO (blue) neurons evoked by consecutive hyperpolarizing voltage steps of −10 mV from −50 to −130 mV (bottom). At the end of each voltage step, the voltage command was returned to −130 mV to evoke tail currents (Ih-tail, depicted with an arrowhead). Red lines show fit of a single exponential function used to assess the Ih activation kinetics. b, Averaged Ih amplitudes were plotted against the voltage step. Ih current amplitudes were measured at the steady state (indicated with a filled circle in a) and the leak current values, as defined as the amplitude of the instantaneous currents at the onset of voltage steps (indicated with an asterisk in a), subtracted. c, Voltage-dependency of Ih-tail currents. Ih-tail amplitudes were normalized relative to Ih-tail at −50 mV and −130 mV. Solid lines show fits with a Boltzmann function for least square fit. The P value shows the difference in V50 between datasets. d, Activation kinetics of the Ih as determined by the τ values obtained from the exponential fitting, as a function of the voltage commands voltage. Only the values obtained from commands between −130 mV to −90 mV were evaluated. e, Comparison between groups of the resting membrane potential as assessed with current clamp recordings. f. Membrane capacitance (Cm) of wild-type and Nlgn3KO DA neurons. g, Input resistant values (Ri) for wild-type and Nlgn3KO mice. Cm and Ri values were obtained in voltage clamp mode by applying a −5 mV (200 ms) voltage command from a holding potential set at −50 mV. All error bars are s.e.m. The P value represents genotype differences. n = 5 mice per genotype, numbers on graphs represent cells. P values determined by repeated-measures two-way ANOVA (b), Boltzmann sigmoidal test (c), mixed-effects model (d), unpaired two-sided t-test (e, f), or two-sided Mann–Whitney test (g). See Supplementary Information for additional statistics.
Extended Data Fig. 3 Oxytocinergic innervation to VTA and Avpr1a mRNA levels are not affected in Nlgn3KO mice.
a, Representative images from 3 mice per genotype of neurophysin 1 (green), a cleavage product of the oxytocin neuropeptide precursor that is transported in vesicles together with oxytocin63, and TH (red) immunofluorescence in the VTA of wild-type and Nlgn3KO mice. Note that oxytocinergic axons arise from multiple hypothalamic nuclei, including the paraventricular nucleus. b, c, Mean VTA area coverage (b) and puncta fluorescence in the VTA (c) from wild-type and Nlgn3KO mice. n = 3 mice per genotype. Numbers in brackets represent sections. d, Quantification of mean Th intensity per TH+ cell. e, Quantification of Nlgn3 puncta per 100 μm2 TH+ cell. f, Quantification of Oxtr puncta per 100 μm2 TH+ cell. n = WT: 280 cells from 4 mice; Nlgn3KO: 265 cells from 3 mice (d–f). g, Targeted proteomic (PRM) measurements for oxytocin receptor (OXTR; left) and AVPR1A (right) proteins in VTA. Numbers on bars indicate mice. h, Representative images of FISH labelling of Avrp1a (cyan), Th (red) and Nlgn3 (green) in the VTA from wild-type and Nlgn3KO mice. Experiment was repeated independently twice. i, Quantification of mean Avpr1a intensity per TH+ cell. j, Quantification of Avpr1a puncta per 100 μm2 TH+ cell from wild-type and Nlgn3KO VTA. n = wild type: 169 cells from 4 animals; Nlgn3KO: 200 cells from 3 mice (i, j). All error bars are s.e.m. P values determined by unpaired two-sided t-test (b, c, g), two-sided Mann–Whitney U test (d–f, i, j). See Supplementary Information for additional statistics.
Extended Data Fig. 4 Ribosomal proteins and translation processes are altered in Nlgn3KO mice.
a, TMT proteomics: graphs plotting abundance of dopamine markers and synaptic proteins from VTA, cortex and hippocampus. Dopaminergic markers are strongly enriched in VTA samples. n = 5 mice per brain region. b, c, Proteomic analysis of wild-type and Nlgn3KO VTA, n = 5 mice per genotype. b, Enrichment of GO terms for biological processes for proteins significantly altered (P < 0.05) in Nlgn3KO mice compared to wild-type mice. c, Network-based analysis of proteins altered in Nlgn3KO VTA (P < 0.01). Blue nodes indicate downregulated proteins, red nodes upregulated proteins, light blue lines indicate interactions known from database and purple lines interactions experimentally determined. Disconnected nodes and nodes containing less than six proteins are not shown. See methods for additional information of statistics and analysis parameters. d, Mean puromycin incorporation in acute cortical slices from adult wild-type and Nlgn3KO mice. All error bars are s.e.m. P values determined by two-sided Mann–Whitney U test. See Supplementary information for additional statistics.
Extended Data Fig. 5 Pharmacological profile of novel MNK1/2 inhibitor ETC-168.
a, b, Quantification of p-ERK1/2 (a) and p-eIF4G (b) levels compared to non-phosphorylated protein in cortical neurons at DIV14 treated with ETC-168. n = 8 replicates from 3 independent experiments. c, d, Quantification of eIF4E (c), ERK1/2 (d) and eIF4G (e) levels normalized to calnexin in cortical neurons at DIV14 treated with ETC-168. P values determined by one-way ANOVA. n = 8 replicates from 3 independent experiments. f, Representative western blot of cerebellar lysate from wild-type mice treated with vehicle or ETC-168 and quantification of p-eIF4E levels compared to eIF4E. n = vehicle: 4; 1 mg kg−1: 3; 5 mg kg−1: 6. g, Representative western blot of VTA lysate from wild-type and Nlgn3KO mice (n = 7 mice per genotype). h, Quantification of p-eIF4E compared with eIF4E levels in VTA of wild-type mice and Nlgn3KO mice. n = 7 mice per genotype. i, Normalized p-eIF4E AlphaLisa counts from wild-type and Nlgn3KO VTA lysate. n = 7 mice per genotype. j, k, Representative western blot (j) and quantification (k) of p-MNK1 and MNK1 levels in VTA lysate from wild-type mice and Nlgn3KO mice. n = 7 mice per genotype. l, Normalized p-eIF4E AlphaLisa counts from VTA from wild-type treated with 5 mg kg−1 ETC-168 for 24 h +2 h. n = 5 mice per genotype. All error bars are s.e.m. P values determined by one-way ANOVA (a, c–f), Kruskal–Wallis test (b), unpaired two-sided t-test (h, k), or two-sided Mann–Whitney test (i, l). See Supplementary Information for additional statistics.
Extended Data Fig. 6 ETC-168 treatment restores cognitive rigidity in Fmr1KO mice.
a, Schematics of the place-independent cue discrimination and reversal task. This task was chosen given that phenotypes in cognitive rigidity tasks have been replicated in several studies on this model. b, Mean consecutive correct responses plotted for Fmr1WT/y and Fmr1KO/y mice. c, Treatment schedule of Fmr1WT/y and Fmr1KO/y mice. Mice were treated daily with vehicle during the learning phase and with 5 mg kg−1 ETC-168 during the reversal phase 2 h before the start of the test. d, Mean conscutive correct responses plotted for vehicle treated Fmr1WT/y and vehicle or ETC-168 Fmr1KO/y mice. Numbers in brackets indicate mice. Error bars denote s.e.m. P values determined by repeated-measures two-way ANOVA followed by Bonferrroni’s post hoc test. See Supplementary Information for additional statistics.
Extended Data Fig. 7 Effect of ETC-168 treatment on protein abundance in wild-type and Nlgn3KO mice.
a, Experimental outline. b, Representative western blot and quantification of p-eIF4E compared to eIF4E levels in VTA lysate from wild-type mice treated with vehicle or 5 mg kg−1 ETC-168 for 7 consecutive days. n = vehicle: 6; ETC-168: 7. c, Normalized p-eIF4E AlphaLisa counts from wild-type mice VTA treated with vehicle or 5 mg kg−1 ETC-168 for 7 consecutive days. n = vehicle: 6; ETC-168: 7. d, Graphs plotting TMT proteomic-normalized protein expression of dopaminergic markers in VTA from wild-type and Nlgn3KO mice treated with vehicle or ETC-168. Mice were treated for 7 days. n = 4 mice per genotype and treatment. e, f, Graphical representation of molecular function GO terms enriched in Nlgn3KO versus wild-type mice treated with vehicle (e), and Nlgn3KO mice treated with ETC-168 versus wild-type mice treated with vehicle (f). GO terms were summarized using REVIGO and only terms with Q < 0.01 are represented. g, h, TMT proteomic comparison of VTA from Nlgn3KO versus wild-type mice treated with vehicle (g) and Nlgn3KO mice treated with ETC-168 versus wild-type mice treated with vehicle (h). Relative frequency of log2-transformed fold change in core proteasome abundance (Nlgn3KO/wild type) is plotted. n = 4 mice per genotype and treatment. All error bars are s.e.m. P values determined by unpaired two-sided t-test (b), two-sided Mann–Whitney U test (c) or Kolmogorov–Smirnov test (g, h). See Methods and Supplementary Information for additional statistics.
Extended Data Fig. 8 Effect of short-term ETC-168 treatment on social recognition in wild-type and Nlgn3KO mice.
a–d, Time course of time interacting in the social habituation/recognition test for mice after short-term treatment with ETC-168. a, Wild-type mice treated with vehicle (n = 12). b, Wild-type mice treated with ETC-168 (n = 10). c, Nlgn3KO mice treated with vehicle (n = 9). d, Nlgn3KO mice treated with ETC-168 (n = 11). Error bars report s.e.m. P values determined by Friedman’s test followed by Dunn’s post hoc test for planned multiple comparison (a, c, d) or repeated-measures one-way ANOVA followed by Bonferroni’s post hoc test for planned multiple comparison (b). See Supplementary Information for additional statistics.
Extended Data Fig. 9 Effect of ETC-168 treatment is dependent on the oxytocin receptor.
a, Experimental outline. b, Mean social interaction time in Nlgn3KO mice treated with 5 mg kg−1 ETC-168 and either vehicle or 10 mg kg−1 OXTR-A. Numbers in brackets indicate mice. c, Social recognition index for Nlgn3KO mice treated with ETC-168 and vehicle, or ETC-168 and OXTR-A. Numbers on graph indicate mice. d, e, Individual values and mean of time interacting in the social habituation/recognition test after treatment with ETC-168 plus vehicle (n = 9) (d), or ETC-168 plus OXTR-A (n = 8) (e). Error bars report s.e.m. P values determined by repeated-measures two-way ANOVA followed by Bonferroni’s post hoc test (b), unpaired two-sided t-test (c), repeated-measures one-way ANOVA followed by Bonferroni’s post hoc test for planned multiple comparison (d, e). See Supplementary Information for additional statistics.
Extended Data Fig. 10 Effect of long-term ETC-168 treatment on behaviour in wild-type and Nlgn3KO mice.
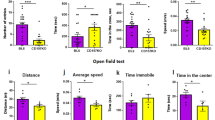
a, Experimental schematics of chronic ETC-168 treatment and behaviour schedule. Number of animals per treatment conditions for all behaviours in b–p: wild-type vehicle = 9, wild-type ETC-168 = 10, Nlgn3KO vehicle = 8, Nlgn3KO ETC-168 = 9. b, c, Mean social interaction time in wild-type (b) and Nlgn3KO (c) mice treated for 8 days with vehicle or 5 mg kg−1 ETC-168. d, Social recognition index for wild-type and Nlgn3KO mice treated with vehicle or 5 mg kg−1 ETC-168. Numbers in brackets indicate mice. e, f, Individual values and mean time interacting in the social habituation/recognition test after chronic treatment with ETC-168 for wild-type vehicle (n = 9) (e), wild-type ETC-168 (n = 10) (f), Nlgn3KO vehicle (n = 8) (g), and Nlgn3KO ETC-168 (n = 9) (h). i, Experimental schematics of object habituation/recognition test in juvenile mice. j–l, Mean object interaction time plotted for wild-type (j) and Nlgn3KO (k) mice. P value above graphs report trial. l, Object recognition index. m, Mean velocity (cm s−1) in an open field arena during 7 min. n, Time spend in centre of the open field arena. o, Number of marbles buried during a 30 min marble burying test. p, Percentage weight gain in wild-type and Nlgn3KO mice treated with ETC-168 or vehicle. P value for treatment is displayed on graphs. Error bars report s.e.m. Repeated measures two-way ANOVA followed by Bonferroni’s post hoc test for genotype and treatment (b, c, j, k); repeated-measures two-way ANOVA followed by Bonferroni’s post hoc test for genotype and treatment (d, l, o), Friedman test followed by Dunn’s post hoc test for planned multiple comparison (e, f, h), one-way ANOVA followed by Bonferroni’s post hoc test for planned multiple comparison (g), or repeated-measures two-way ANOVA (m, n, p). See Supplementary Information for additional statistics.
Supplementary information
Supplementary Information
This file contains additional information on statistics and uncropped western blots.
Supplementary Table
Supplementary Table 1: Peptides used in targeted proteomic analysis (Excel file). Peptides used to detect OXTR and V1aR in targeted proteomic analysis of VTA from wild type and Nlgn3KO mice. The table is available as a separate excel file.
Supplementary Table
Supplementary Table 2: Detailed data from shotgun proteomics (Excel file). LC–MS shotgun proteomic analysis of VTA from wild type and Nlgn3KO mice (postnatal day 31-34) treated with vehicle or ETC-168 for 10 days (n=5 mice per genotype and treatment). Sheet 1 lists protein name, gene name, number of identified peptides, normalized protein expression (n: mean of 5 per genotype and treatment condition), Log2 fold-change and P-values compared to vehicle-treated wild type. P-value was calculated using a two-sided Bayes moderated t-Test, adjusted for multiple testing using the Benjamini-Hochberg method. Sheet 2 contains GO terms for Biological Processes enriched in Nlgn3KO vehicle VTA compared to wild type vehicle VTA, and lists GO term, percentage of proteins, P-value, Fold Enrichment, Benjamini test and genes. Please see methods for additional information on statistics and data processing. The table is available as a separate excel file.
Supplementary Table
Supplementary Table 3: Summary data from tandem mass tag proteomic experiments (Excel file). Tandem mass Tag (TMT) proteomic analysis of VTA from wild type and Nlgn3KO mice (postnatal day 28-32) treated with vehicle or ETC-168 for 7days. Sheet 1 lists protein name, gene name, number of identified peptides, normalized protein expression (n: mean of 4 per genotype and treatment condition), Log2 fold-change and P-values compared to vehicle-treated wild type. P-value was calculated using a two-sided Bayes moderated t-Test, adjusted for multiple testing using the Benjamini-Hochberg method. Sheet 2-4 contains proteins used to analyze the distribution of ribosomal, mitochondrial ribosomal, and proteasome proteins, respectively. Sheet 5 and 6 contains PSEA-Quant gene enrichment analysis of GO terms with an adjusted P-value of Q<0.01. Please see methods for additional information on statistics and data processing. The table is available as a separate excel file.
Source data
Rights and permissions
About this article
Cite this article
Hörnberg, H., Pérez-Garci, E., Schreiner, D. et al. Rescue of oxytocin response and social behaviour in a mouse model of autism. Nature 584, 252–256 (2020). https://doi.org/10.1038/s41586-020-2563-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2563-7
This article is cited by
-
Differential contribution of canonical and noncanonical NLGN3 pathways to early social development and memory performance
Molecular Brain (2024)
-
Shank3 deficiency elicits autistic-like behaviors by activating p38α in hypothalamic AgRP neurons
Molecular Autism (2024)
-
Whole-brain in vivo base editing reverses behavioral changes in Mef2c-mutant mice
Nature Neuroscience (2024)
-
Social circuits and their dysfunction in autism spectrum disorder
Molecular Psychiatry (2023)
-
SCGN deficiency is a risk factor for autism spectrum disorder
Signal Transduction and Targeted Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.