Abstract
Multicellular organisms use cell-surface receptor kinases to sense and process extracellular signals. Many plant receptor kinases are activated by the formation of ligand-induced complexes with shape-complementary co-receptors1. The best-characterized co-receptor is BRASSINOSTEROID INSENSITIVE 1-ASSOCIATED KINASE 1 (BAK1), which associates with numerous leucine-rich repeat receptor kinases (LRR-RKs) to control immunity, growth and development2. Here we report key regulatory events that control the function of BAK1 and, more generally, LRR-RKs. Through a combination of phosphoproteomics and targeted mutagenesis, we identified conserved phosphosites that are required for the immune function of BAK1 in Arabidopsis thaliana. Notably, these phosphosites are not required for BAK1-dependent brassinosteroid-regulated growth. In addition to revealing a critical role for the phosphorylation of the BAK1 C-terminal tail, we identified a conserved tyrosine phosphosite that may be required for the function of the majority of Arabidopsis LRR-RKs, and which separates them into two distinct functional classes based on the presence or absence of this tyrosine. Our results suggest a phosphocode-based dichotomy of BAK1 function in plant signalling, and provide insights into receptor kinase activation that have broad implications for our understanding of how plants respond to their changing environment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information files. Source Data (gels and graphs) for Figs. 1–4 and Extended Data Figs. 1–7 and 9 are provided with the paper.
Change history
17 October 2018
In Extended Data Fig. 5d of this Letter, the blots for anti-pS612 and anti-BAK1 were inadvertently duplicated. This figure has been corrected online.
References
Hohmann, U., Lau, K. & Hothorn, M. The structural basis of ligand perception and signal activation by receptor kinases. Annu. Rev. Plant Biol. 68, 109–137 (2017).
Ma, X., Xu, G., He, P. & Shan, L. SERKing coreceptors for receptors. Trends Plant Sci. 21, 1017–1033 (2016).
Chinchilla, D. et al. A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 448, 497–500 (2007).
Heese, A. et al. The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Proc. Natl Acad. Sci. USA 104, 12217–12222 (2007).
Roux, M. et al. The Arabidopsis leucine-rich repeat receptor-like kinases BAK1/SERK3 and BKK1/SERK4 are required for innate immunity to hemibiotrophic and biotrophic pathogens. Plant Cell 23, 2440–2455 (2011).
Nam, K. H. & Li, J. BRI1/BAK1, a receptor kinase pair mediating brassinosteroid signaling. Cell 110, 203–212 (2002).
Li, J. et al. BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signaling. Cell 110, 213–222 (2002).
Sun, Y. et al. Structure reveals that BAK1 as a co-receptor recognizes the BRI1-bound brassinolide. Cell Res. 23, 1326–1329 (2013).
Wang, X. et al. Sequential transphosphorylation of the BRI1/BAK1 receptor kinase complex impacts early events in brassinosteroid signaling. Dev. Cell 15, 220–235 (2008).
Karlova, R. et al. Identification of in vitro phosphorylation sites in the Arabidopsis thaliana somatic embryogenesis receptor-like kinases. Proteomics 9, 368–379 (2009).
Yun, H. S. et al. Analysis of phosphorylation of the BRI1/BAK1 complex in Arabidopsis reveals amino acid residues critical for receptor formation and activation of BR signaling. Mol. Cells 27, 183–190 (2009).
Wu, X. et al. Transphosphorylation of E. coli proteins during production of recombinant protein kinases provides a robust system to characterize kinase specificity. Front. Plant Sci. 3, 262 (2012).
Yan, L. et al. Structural basis for the impact of phosphorylation on the activation of plant receptor-like kinase BAK1. Cell Res. 22, 1304–1308 (2012).
Ntoukakis, V., Schwessinger, B., Segonzac, C. & Zipfel, C. Cautionary notes on the use of C-terminal BAK1 fusion proteins for functional studies. Plant Cell 23, 3871–3878 (2011).
Yu, X., Feng, B., He, P. & Shan, L. From chaos to harmony: responses and signaling upon microbial pattern recognition. Annu. Rev. Phytopathol. 55, 109–137 (2017).
Schwessinger, B. et al. Phosphorylation-dependent differential regulation of plant growth, cell death, and innate immunity by the regulatory receptor-like kinase BAK1. PLoS Genet. 7, e1002046 (2011).
aan den Toorn, M., Albrecht, C. & de Vries, S. On the origin of SERKs: bioinformatics analysis of the somatic embryogenesis receptor kinases. Mol. Plant 8, 762–782 (2015).
Wu, D., Liu, Y., Xu, F. & Zhang, Y. Differential requirement of BAK1 C-terminal tail in development and immunity. J. Integr. Plant Biol. 60, 270–275 (2018).
Macho, A. P., Lozano-Durán, R. & Zipfel, C. Importance of tyrosine phosphorylation in receptor kinase complexes. Trends Plant Sci. 20, 269–272 (2015).
Wang, Y. et al. Assessment of BAK1 activity in different plant receptor-like kinase complexes by quantitative profiling of phosphorylation patterns. J. Proteomics 108, 484–493 (2014).
Macho, A. P. et al. A bacterial tyrosine phosphatase inhibits plant pattern recognition receptor activation. Science 343, 1509–1512 (2014).
Mitra, S. K. et al. An autophosphorylation site database for leucine-rich repeat receptor-like kinases in Arabidopsis thaliana. Plant J. 82, 1042–1060 (2015).
Bender, K. W. et al. Glutaredoxin AtGRXC2 catalyses inhibitory glutathionylation of Arabidopsis BRI1-associated receptor-like kinase 1 (BAK1) in vitro. Biochem. J. 467, 399–413 (2015).
Meng, X. et al. Ligand-induced receptor-like kinase complex regulates floral organ abscission in Arabidopsis. Cell Rep. 14, 1330–1338 (2016).
Meng, X. et al. Differential function of Arabidopsis SERK family receptor-like kinases in stomatal patterning. Curr. Biol. 25, 2361–2372 (2015).
Stührwohldt, N., Dahlke, R. I., Steffens, B., Johnson, A. & Sauter, M. Phytosulfokine-α controls hypocotyl length and cell expansion in Arabidopsis thaliana through phytosulfokine receptor 1. PLoS ONE 6, e21054 (2011).
Shpak, E. D., Berthiaume, C. T., Hill, E. J. & Torii, K. U. Synergistic interaction of three ERECTA-family receptor-like kinases controls Arabidopsis organ growth and flower development by promoting cell proliferation. Development 131, 1491–1501 (2004).
Suzuki, M. et al. Autophosphorylation of specific threonine and tyrosine residues in Arabidopsis CERK1 is essential for the activation of chitin-induced immune signaling. Plant Cell Physiol. 57, 2312–2322 (2016).
Zhang, G. et al. Mass spectrometry mapping of epidermal growth factor receptor phosphorylation related to oncogenic mutations and tyrosine kinase inhibitor sensitivity. J. Proteome Res. 10, 305–319 (2011).
Singh, V. et al. Tyrosine-610 in the receptor kinase BAK1 does not play a major role in brassinosteroid signaling or innate immunity. Front. Plant Sci. 8, 1273 (2017).
Robatzek, S., Chinchilla, D. & Boller, T. Ligand-induced endocytosis of the pattern recognition receptor FLS2 in Arabidopsis. Genes Dev. 20, 537–542 (2006).
Nekrasov, V. et al. Control of the pattern-recognition receptor EFR by an ER protein complex in plant immunity. EMBO J. 28, 3428–3438 (2009).
Lee, J. S. et al. Direct interaction of ligand–receptor pairs specifying stomatal patterning. Genes Dev. 26, 126–136 (2012).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Kadota, Y. et al. Direct regulation of the NADPH oxidase RBOHD by the PRR-associated kinase BIK1 during plant immunity. Mol. Cell 54, 43–55 (2014).
Felix, G., Duran, J. D., Volko, S. & Boller, T. Plants have a sensitive perception system for the most conserved domain of bacterial flagellin. Plant J. 18, 265–276 (1999).
Huffaker, A., Pearce, G. & Ryan, C. A. An endogenous peptide signal in Arabidopsis activates components of the innate immune response. Proc. Natl Acad. Sci. USA 103, 10098–10103 (2006).
Kunze, G. et al. The N terminus of bacterial elongation factor Tu elicits innate immunity in Arabidopsis plants. Plant Cell 16, 3496–3507 (2004).
Monaghan, J. et al. The calcium-dependent protein kinase CPK28 buffers plant immunity and regulates BIK1 turnover. Cell Host Microbe 16, 605–615 (2014).
Yoo, S.-D., Cho, Y.-H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565–1572 (2007).
Sun, Y. et al. Structural basis for flg22-induced activation of the Arabidopsis FLS2-BAK1 immune complex. Science 342, 624–628 (2013).
Ladwig, F. et al. Phytosulfokine regulates growth in Arabidopsis through a response module at the plasma membrane that includes CYCLIC NUCLEOTIDE-GATED CHANNEL17, H+-ATPase, and BAK1. Plant Cell 27, 1718–1729 (2015).
Bender, K. W. et al. Autophosphorylation-based calcium (Ca2+) sensitivity priming and Ca2+/calmodulin inhibition of Arabidopsis thaliana Ca2+-dependent protein kinase 28 (CPK28). J. Biol. Chem. 292, 3988–4002 (2017).
Horst, R. J. et al. Molecular framework of a regulatory circuit initiating two-dimensional spatial patterning of stomatal lineage. PLoS Genet. 11, e1005374 (2015).
Holm, L. & Laakso, L. M. Dali server update. Nucleic Acids Res. 44, W351–W355 (2016).
Lehti-Shiu, M. D. & Shiu, S.-H. Diversity, classification and function of the plant protein kinase superfamily. Philos. Trans. R. Soc. Lond. B 367, 2619–2639 (2012).
Shiu, S. H. & Bleecker, A. B. Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc. Natl Acad. Sci. USA 98, 10763–10768 (2001).
Acknowledgements
We thank K. Morehouse, M. Smoker, J. Taylor and the John Innes Centre Horticultural Services for technical help; V. Ntoukakis, M. Stegmann and J. Dang for technical advice; S. Huber, K. Bender and R. Zielinski for providing materials; Y. Belkhadir and S. Huber for reading the manuscript; and all members of the Zipfel laboratory for discussion. This work was supported by the Gatsby Charitable Foundation and the European Research Council (grant ‘PHOSPHinnATE’) (C.Z.); the Gordon and Betty Moore Foundation (grant GBMF3035) and the Howard Hughes Medical Institute (K.U.T.); the European Molecular Biology Organization (EMBO-LTFs 100-2017 to T.A.D. and 225-2015 to S.J.); the RIKEN Special Postdoctoral Research Fellowship, the JSPS Excellent Young Researcher Overseas Visit Program, and the Uehara Memorial Foundation (Y.K.); and the JIC/TSL PhD Rotation Program (B.S.).
Reviewer information
Nature thanks S. Clouse and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.P. designed and conceived experiments, collected most of the data and wrote the manuscript. T.A.D. developed and collected data in many of the biochemical assays and participated in the preparation of the final manuscript. P.D. and J.S. performed proteomics-based analysis, under the supervision of F.L.H.M. J.A. and X.Q. designed and performed ERECTA-related experiments under the supervision of K.U.T. D.S. performed the protoplast swelling experiments. B.S., Y.K. and A.P.M. performed preliminary work on BAK1 phosphorylation. L.S., S.J. and D.C. provided technical help with experiments. C.Z. designed and conceived experiments, supervised the study and wrote the manuscript. All authors commented and agreed on the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification of BAK1 phosphosites.
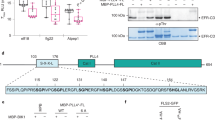
a, Representative CBB-stained SDS–PAGE gel showing proteins that are enriched upon GFP immunoprecipitation. b, Western blot of BAK1 co-immunoprecipitated with FLS2–GFP and EFR–GFP from a using anti-GFP and anti-BAK1 antibodies. For a and b, experiments were independently repeated three times. For gel and blot source data, see Supplementary Fig. 1. c, Summary of BAK1 in vivo phosphosites identified by FLS2–GFP and EFR–GFP co-immunoprecipitation followed by LC–MS/MS analysis. Spectra of identified sites are presented in Supplementary Fig. 2.
Extended Data Fig. 2 Individual or double BAK1(S602A), BAK1(T603A) or BAK1(S604A) mutations do not affect elf18-induced ROS production.
a, e, Total ROS production of bak1-4 mesophyll protoplasts transiently expressing the indicated BAK1 mutants, after treatment with 100 nM elf18 for 60 min. n = 12 biological independent suspensions. b, d, f, Western blots with anti-BAK1 antibodies. For blot source data, see Supplementary Fig. 1. c, Total ROS production after treatment of leaf discs with 100 nM flg22 for 40 min. Individual data points are shown as circles. n = 8 biologically independent leaf discs. In a, c and e, measurements are plotted as box plots displaying the first and third quartiles, split by the median; whiskers extend to a maximum of 1.5× interquartile range beyond the box. Outliers are indicated as black dots. Statistical analysis was performed using a one-way ANOVA with Dunnett’s post hoc test compared to wild-type BAK1. For a–f, experiments were repeated at least three times.
Extended Data Fig. 3 BAK1(AAA) and BAK1(S612A) plants are not affected in flg22-induced FLS2–BAK1 complex formation.
a, b, Western blots with anti-BAK1 antibodies. c, d, Co-immunoprecipitation between FLS2 and BAK1 using anti-FLS2 and anti-BAK1 antibodies. Two-week-old seedlings were treated with water (−) or 100 nM flg22 (+) for 10 min before protein extraction. Blots were stained with CBB for loading control. For a–d, experiments were repeated at least three times with similar results. For blot source data, see Supplementary Fig. 1.
Extended Data Fig. 4 Expression of the BAK1(AAA) and BAK1(S612A) variants restores the semi-dwarf rosette phenotype of bak1-4 and bak1-4 bri1-301.
a, b, Representative images of rosettes from four- to five-week-old plants. c, d, Left, representative images of rosettes from five- to seven-week-old plants. Right, western blots with anti-BAK1 antibodies. Blots were stained with CBB for loading control. Experiments were repeated at least twice with similar results. Control plants used in b have been previously pictured30. e, BES1 dephosphorylation 60 min after treatment of ten-day-old seedlings with the indicated concentrations of epi-BL (µM), as shown by western-blot analysis using anti-BES1 antibodies. Blots were probed with anti-BAK1 antibodies, and subsequently stained with CBB as loading controls. For a–e, experiments were repeated at least three times. For blot source data, see Supplementary Fig. 1.
Extended Data Fig. 5 Conservation of BAK1 C-terminal-tail phosphosites in SERK proteins across plant species.
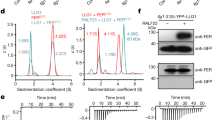
a, Clustal Omega multiple alignments were visualized using JalView v2.10.2b2. The alignment is coloured by percentage identity. Yellow, conservation of BAK1 S602, S604 and S612; green, conservation of BAK1 T603. Protein IDs used for the alignments are as follows: Populus trichocarpa PtSERK1(B9MW41), PtSERK2(B9IQM9), PtSERK3(B9HFX1), PtSERK4(B9H599), Dimocarpus longan DlSERK(B5TTV0), Medicago truncatula MtSERK1(Q8GRK2), MtSERK2(E2IXG1), MtSERK3(E2IXG8), MtSERK4(E2IXG2), MtSERK5(E2IXG3), MtSERK6(E2IXG4), Glycine max GmSERK1(C6ZGA8), GmSERK2(C6FF61), Citrus unshiu CuSERK (Q6BE26), Citrus sinensis CsSERK(C3V9W0), Vitis vinifera VvSERK1(D7TXV2), VvSERK2(A5BIY4), VvSERK3(D7STF6), Cyclamen persicum CpSERK1(A7L5U3), CpSERK2(E5D6S9), Gossypium hirsutum GhSERK1(E5Q8K6), GhSERK2(F5BZU9), GhSERK3(F6MF11), Solanum tuberosum StSERK(A3R789), Solanum peruvianum SpSERK(A6N8J2), Solanum lycopersicum SlSERK1(G0XZA3), SlSERK3A(G0XZA5), SlSERK3B(G0XZA6), Daucus carota DcSERK(O23921), Arabidopsis thaliana AtSERK1(Q94AG2), AtSERK2(Q9XIC7), AtSERK3(Q94F62), AtSERK4(Q9SKG5), AtSERK5(Q8LPS5), Nicotiana benthamiana NbSERK3A(E3VXE6), NbSERK3B(E3VXE7), Oryza sativa subsp. Indica OsbiSERK(Q6S7F1), Oryza sativa OsSERK(Q5Y8C8), OsSERKlike1(Q67X31), OsSERKlike2(Q6K4T4), Sorghum bicolor SbSERK1(C5YHV3), SbSERK2(C5Y9S6), SbSERK3(C5XVP5), Ananas comosus AcSERK1(H6SU43), AcSERK2(H6UP78), AcSERK3(H6UP79), Cyrtochilum loxense ClSERK(G2XLB1), Cocos nucifera CnSERK(Q5S1N9), Zea mays ZmSERK1(Q93W70), ZmSERK2(Q94IJ5), ZmSERK3(B4G007), Triticum aestivum TaSERK1(G4XGX1), TaSERK2(G4XGX2), TaSERKlike3(G4XGX3), Selaginella moellendorffii SmSERK1(D8SBB8), SmSERK2(D8S0N3), SmSERK3(D8S4M4), SmSERK4(D8R6C9), Marchantia polymorpha MpSERK(A7VM18), Physcomitrella patens PpSERK1(A9STU8), PpSERK2(A9SMW5), PpSERK3(A9RY79), Poa pratensis PoapSERKlike1(Q659J0), PoapSERKlike2(Q659J1), Closterium ehrenbergii CeSERK(A7VM46). b, WebLogo representation of alignment in a. c, Detection of BAK1 S612 phosphorylation using anti-pS612 specific antibodies on affinity-purified recombinant BAK1 cytoplasmic domain (BAK1CD) following an in vitro kinase assay with cold ATP. Membranes were immunoblotted with anti-pS612 and anti-MBP antibodies. d, BAK1 immunoprecipitation and detection of BAK1 S612 phosphorylation in vivo using anti-pS612 antibodies. Two-week-old seedlings were treated with water (−) or 100 nM flg22 (+) for 10 min. The same membrane was stripped and blotted again with anti-BAK1 antibodies for loading control. e, Western-blot analysis using anti-BAK1 and anti-pS612 antibodies after SDS–PAGE of crude protein extracts from two-week-old seedlings treated with 100 nM flg22 for 10 min. Blots were stained with CBB for loading control. For a–e, all experiments were repeated at least twice with similar results. For blot source data, see Supplementary Fig. 1.
Extended Data Fig. 6 Mutational screen of tyrosine residues present in the cytoplasmic domain of BAK1.
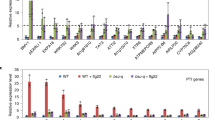
a, Total ROS production after treatment of bak1-4 mesophyll protoplasts with 100 nM elf18 for 30 min. The RLU values are normalized relative to protoplasts expressing wild-type (WT) BAK1 and are expressed as aligned dot blots showing mean ± s.e.m. n = 4 independent biological experiments. b, Total ROS production of bak1-4 mesophyll protoplasts that transiently express the indicated BAK1 mutants, after treatment with 100 nM elf18 for 60 min. n = 8 biologically independent suspensions. Measurements are plotted as box plots displaying the first and third quartiles, split by the median; whiskers extend to a maximum of 1.5× interquartile range beyond the box. Individual data points are shown as circles. c, Western blot with anti-BAK1 antibodies. For blot source data, see Supplementary Fig. 1. In a and b, statistical analysis was performed using a one-way ANOVA with Dunnett’s post hoc test compared to the wild-type control. For b and c, experiments were repeated twice with similar results.
Extended Data Fig. 7 BAK1 Y403 plants are not affected by flg22-induced FLS2–BAK1 complex formation and brassinosteroid-mediated rosette growth.
a, Western blot with anti-BAK1 antibodies. Blots were stained with CBB for loading control. b, Co-immunoprecipitation experiments between FLS2 and BAK1 using anti-FLS2 and anti-BAK1 antibodies. Two-week-old seedlings were treated with water (−) or 100 nM flg22 (+) for 10 min before protein extraction. Blots were stained with CBB for loading control. c, Representative images of rosettes from four- to five-week-old plants. d, Representative images of rosettes from five-week-old plants. The same control plants used in d are also presented in Extended Data Fig. 4c. e, Western blot with anti-BAK1 antibodies. Blots were stained with CBB for loading control. For a–e, experiments were repeated three times with similar results. For blot source data, see Supplementary Fig. 1.
Extended Data Fig. 8 The conserved Tyr-VIa residue BAK1 Y403 is in close proximity to the catalytic loop and to C408.
a, In silico representation of the structure of BAK1CD (PDB ID: 3UIM). The activation-segment region is shown in green and the catalytic loop in purple. b, Conservation of BAK1 Y403 in SERK proteins across plant species. Clustal Omega multiple alignments were visualized using JalView v2.10.2b2. The alignment is coloured by percentage identity. Magenta, conservation of BAK1 Y403. Protein IDs used for the alignments are as follows: PpSERK1(B9MW41), PpSERK2(B9IQM9), PpSERK3(B9HFX1), PpSERK4(B9H599), DlSERK(B5TTV0), MtSERK1(Q8GRK2), MtSERK2(E2IXG1), MtSERK3(E2IXG8), MtSERK4(E2IXG2), MtSERK5(E2IXG3), MtSERK6(E2IXG4), GmSERK1(C6ZGA8), GmSERK2(C6FF61), CuSERK (Q6BE26), CsSERK(C3V9W0), VvSERK1(D7TXV2), VvSERK2(A5BIY4), VvSERK3(D7STF6), CpSERK1(A7L5U3), CpSERK2(E5D6S9), GhSERK1(E5Q8K6), GhSERK2(F5BZU9), GhSERK3(F6MF11), StSERK(A3R789), SpSERK(A6N8J2), SlSERK1(G0XZA3), SlSERK3A(G0XZA5), SlSERK3B(G0XZA6), DcSERK(O23921), AtSERK1(Q94AG2), AtSERK2(Q9XIC7), AtSERK3(Q94F62), AtSERK4(Q9SKG5), AtSERK5(Q8LPS5), NbSERK3A(E3VXE6), NbSERK3B(E3VXE7), OsbiSERK(Q6S7F1), OsSERK(Q5Y8C8), OsSERKlike1(Q67X31), OsSERKlike2(Q6K4T4), SbSERK1(C5YHV3), SbSERK2(C5Y9S6), SbSERK3(C5XVP5), AcSERK1(H6SU43), AcSERK2(H6UP78), AcSERK3(H6UP79), ClSERK(G2XLB1), CnSERK(Q5S1N9), ZmSERK1(Q93W70), ZmSERK2(Q94IJ5), ZmSERK3(B4G007), TaSERK1(G4XGX1), TaSERK2(G4XGX2), TaSERKlike3(G4XGX3), SmSERK1(D8SBB8), SmSERK2(D8S0N3), SmSERK3(D8S4M4), SmSERK4(D8R6C9), MpSERK(A7VM18), PpSERK1(A9STU8), PpSERK2(A9SMW5), PpSERK3(A9RY79), PoapSERKlike1(Q659J0), PoapSERKlike2(Q659J1), CeSERK(A7VM46).
Extended Data Fig. 9 Analysis of phosphosite-specific kinase activities of BAK1.
a, Specific detection of BAK1 Y403 phosphorylation on affinity-purified recombinant BAK1CD wild-type but not Y403F or kinase-dead (D416N) proteins, using anti-pY403 antibodies and following an in vitro kinase assay with cold ATP. Blots were probed with anti-BAK1 antibodies for loading control. b, Detection of BAK1 Y403, S612, threonine and tyrosine phosphorylation using phosphorylation-specific antibodies on affinity-purified recombinant BAK1CD following an in vitro kinase assay with cold ATP. Membranes were immunoblotted with BAK1 antibodies for loading control. c, [32P]γ-ATP kinase assays showing autophosphorylation of the indicated MBP-BAK1CD phosphosite mutants. d, trans-autophosphorylation of the kinase-dead BAK1(D416N) (6×His-BAK1*CD) by the indicated BAK1 phosphosite mutants (MBP–BAK1CD). e, Trans-phosphorylation activity of the indicated affinity purified recombinant BAK1CD mutants against a kinase-dead (K105E) BIK1 substrate (GST–BIK1*). f, Trans-autophosphorylation of kinase-dead (D416N) BAK1 substrate (6×His-BAK1*CD) at S612 and Y403 by wild-type BAK1 (MBP–BAK1CD). BAK1 Y403 or S612 phosphorylation was detected using phosphorylation-specific antibodies following an in vitro incubation of the purified proteins in the presence or in the absence of cold ATP. Membranes were immunoblotted with BAK1 antibodies for loading control. In c–e, numbers indicate autoradiograph band intensity relative to the wild type. Protein loading control was determined with CBB staining. For a–f, all experiments were repeated independently at least three times. For blot source data, see Supplementary Fig. 1.
Extended Data Fig. 10 The conservation of Tyr-VIa in the A. thaliana LRR-RK cytoplasmic domain and in representative members of the 20 groups of animal-receptor tyrosine kinases.
a, b, Clustal Omega multiple alignments of A. thaliana LRR-RK cytoplasmic domain, visualized using JalView v2.10.2b2, illustrate the conservation (a) and non-conservation (b) of Tyr-VIa. The alignment is coloured by percentage identity. Magenta, Tyr-VIa. Protein IDs of the sequences used for the alignment are in Supplementary Table 1. Arrows indicate the LRR-RKs reported in this study. c, The conservation of Tyr-VIa in representative members of the 20 groups of animal receptor tyrosine kinases. Clustal Omega multiple alignments were visualized using JalView v2.10.2b2. The alignment is coloured by percentage identity. Red, position analogous to BAK1 Y403. Protein IDs used for the alignments: EGFR (P00533), AXL (P30530), DDR15 (Q08345), EphA1 (P21709), FGFR2 (P21802), HGFR (P08581), INSR (P06213), PTK7 (Q13308), LTK (P29376), MUSK (O15146), PGFRB (P09619), RET (P07949), RYK (P34925), TIE1 (P35590), NTRK1 (P04629), VGFR1 (P17948), ROR1 (Q01973), ROS (P08922), LMR1 (Q6ZMQ8), STYK1 (Q6J9G0). d, In silico alignments of the structures of BAK1CD and EGFRCD. A selected overlapping region of the BAK1 (PDB ID: 3UIM) and EGFR (PDB ID: 2JIT) cytoplasmic domain structures is presented, highlighting BAK1 Y403 and EGFR Y827.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1: Source data for images for gels and blots. Original source images for all data obtained by SDS-PAGE, western blots, autoradiography scans and Coomassie Blue stained blots and gels, Supplementary Figure 2: Identification of in vivo phosphorylation sites of BAK1. Orbitrap MS2 spectra obtained for the identified pT446, pS602, pT603, pS604 and pS612 and Supplementary Table 2: List of primers used in this study. This table lists all the primers used in this study.
Supplementary Table
This file contains Supplementary Table 1: List of Arabidopsis thaliana LRR-RKs. This table lists all the Arabidopsis thaliana LRR-RKs used for the alignments in this study.
Rights and permissions
About this article
Cite this article
Perraki, A., DeFalco, T.A., Derbyshire, P. et al. Phosphocode-dependent functional dichotomy of a common co-receptor in plant signalling. Nature 561, 248–252 (2018). https://doi.org/10.1038/s41586-018-0471-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0471-x
Keywords
This article is cited by
-
The COG1-OsSERL2 complex senses cold to trigger signaling network for chilling tolerance in japonica rice
Nature Communications (2023)
-
Perception of a divergent family of phytocytokines by the Arabidopsis receptor kinase MIK2
Nature Communications (2021)
-
Hydrogen peroxide sensor HPCA1 is an LRR receptor kinase in Arabidopsis
Nature (2020)
-
MAMP-triggered plant immunity mediated by the LysM-receptor kinase CERK1
Journal of General Plant Pathology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.