Abstract
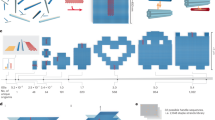
DNA nanotechnology, in particular DNA origami, enables the bottom-up self-assembly of micrometre-scale, three-dimensional structures with nanometre-precise features1,2,3,4,5,6,7,8,9,10,11,12. These structures are customizable in that they can be site-specifically functionalized13 or constructed to exhibit machine-like14,15 or logic-gating behaviour16. Their use has been limited to applications that require only small amounts of material (of the order of micrograms), owing to the limitations of current production methods. But many proposed applications, for example as therapeutic agents or in complex materials3,16,17,18,19,20,21,22, could be realized if more material could be used. In DNA origami, a nanostructure is assembled from a very long single-stranded scaffold molecule held in place by many short single-stranded staple oligonucleotides. Only the bacteriophage-derived scaffold molecules are amenable to scalable and efficient mass production23; the shorter staple strands are obtained through costly solid-phase synthesis24 or enzymatic processes25. Here we show that single strands of DNA of virtually arbitrary length and with virtually arbitrary sequences can be produced in a scalable and cost-efficient manner by using bacteriophages to generate single-stranded precursor DNA that contains target strand sequences interleaved with self-excising ‘cassettes’, with each cassette comprising two Zn2+-dependent DNA-cleaving DNA enzymes. We produce all of the necessary single strands of DNA for several DNA origami using shaker-flask cultures, and demonstrate end-to-end production of macroscopic amounts of a DNA origami nanorod in a litre-scale stirred-tank bioreactor. Our method is compatible with existing DNA origami design frameworks and retains the modularity and addressability of DNA origami objects that are necessary for implementing custom modifications using functional groups. With all of the production and purification steps amenable to scaling, we expect that our method will expand the scope of DNA nanotechnology in many areas of science and technology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rothemund, P. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006)
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009)
Hong, F., Zhang, F., Liu, Y. & Yan, H. DNA origami: scaffolds for creating higher order structures. Chem. Rev. 117, 12584–12640 (2017)
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009)
Veneziano, R. et al. Designer nanoscale DNA assemblies programmed from the top down. Science 352, 1534 (2016)
Benson, E. et al. DNA rendering of polyhedral meshes at the nanoscale. Nature 523, 441–444 (2015)
Gerling, T., Wagenbauer, K. F., Neuner, A. M. & Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non-base pairing 3D components. Science 347, 1446–1452 (2015)
Funke, J. J. & Dietz, H. Placing molecules with Bohr radius resolution using DNA origami. Nat. Nanotechnol. 11, 47–52 (2016)
Bai, X. C., Martin, T. G., Scheres, S. H. & Dietz, H. Cryo-EM structure of a 3D DNA-origami object. Proc. Natl Acad. Sci. USA 109, 20012–20017 (2012)
Zhang, F. et al. Complex wireframe DNA origami nanostructures with multi-arm junction vertices. Nat. Nanotechnol. 10, 779–784 (2015)
Wei, B., Dai, M. & Yin, P. Complex shapes self-assembled from single-stranded DNA tiles. Nature 485, 623–626 (2012)
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012)
Knudsen, J. B. et al. Routing of individual polymers in designed patterns. Nat. Nanotechnol. 10, 892–898 (2015)
Ketterer, P., Willner, E. M. & Dietz, H. Nanoscale rotary apparatus formed from tight-fitting 3D DNA components. Sci. Adv. 2, e1501209 (2016)
Marras, A. E., Zhou, L., Su, H. J. & Castro, C. E. Programmable motion of DNA origami mechanisms. Proc. Natl Acad. Sci. USA 112, 713–718 (2015)
Douglas, S. M., Bachelet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012)
Douglas, S. M., Chou, J. J. & Shih, W. M. DNA-nanotube-induced alignment of membrane proteins for NMR structure determination. Proc. Natl Acad. Sci. USA 104, 6644–6648 (2007)
Siavashpouri, M. et al. Molecular engineering of chiral colloidal liquid crystals using DNA origami. Nat. Mater. 16, 849–856 (2017)
Halley, P. D. et al. Daunorubicin-loaded DNA origami nanostructures circumvent drug-resistance mechanisms in a leukemia model. Small 12, 308–320 (2016)
Zhao, Y. X. et al. DNA origami delivery system for cancer therapy with tunable release properties. ACS Nano 6, 8684–8691 (2012)
Jiang, Q. et al. DNA origami as a carrier for circumvention of drug resistance. J. Am. Chem. Soc. 134, 13396–13403 (2012)
Jiang, Q. et al. A self-assembled DNA origami-gold nanorod complex for cancer theranostics. Small 11, 5134–5141 (2015)
Kick, B., Praetorius, F., Dietz, H. & Weuster-Botz, D. Efficient production of single-stranded phage DNA as scaffolds for DNA origami. Nano Lett. 15, 4672–4676 (2015)
Reese, C. B. Oligo- and poly-nucleotides: 50 years of chemical synthesis. Org. Biomol. Chem. 3, 3851–3868 (2005)
Ducani, C., Kaul, C., Moche, M., Shih, W. M. & Högberg, B. Enzymatic production of ‘monoclonal stoichiometric’ single-stranded DNA oligonucleotides. Nat. Methods 10, 647–652 (2013)
Gu, H., Furukawa, K., Weinberg, Z., Berenson, D. F. & Breaker, R. R. Small, highly active DNAs that hydrolyze DNA. J. Am. Chem. Soc. 135, 9121–9129 (2013)
Gu, H. & Breaker, R. R. Production of single-stranded DNAs by self-cleavage of rolling-circle amplification products. Biotechniques 54, 337–343 (2013)
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009)
Sobczak, J. P., Martin, T. G., Gerling, T. & Dietz, H. Rapid folding of DNA into nanoscale shapes at constant temperature. Science 338, 1458–1461 (2012)
Vieira, J. & Messing, J. Production of single-stranded plasmid DNA. Methods Enzymol. 153, 3–11 (1987)
Castro, C. E. et al. A primer to scaffolded DNA origami. Nat. Methods 8, 221–229 (2011)
Kim, D. N., Kilchherr, F., Dietz, H. & Bathe, M. Quantitative prediction of 3D solution shape and flexibility of nucleic acid nanostructures. Nucleic Acids Res. 40, 2862–2868 (2012)
Shih, W. M., Quispe, J. D. & Joyce, G. F. A 1.7-kilobase single-stranded DNA that folds into a nanoscale octahedron. Nature 427, 618–621 (2004)
Engler, C., Kandzia, R. & Marillonnet, S. A one pot, one step, precision cloning method with high throughput capability. PLoS One 3, e3647 (2008)
Marchi, A. N., Saaem, I., Vogen, B. N., Brown, S. & LaBean, T. H. Toward larger DNA origami. Nano Lett. 14, 5740–5747 (2014)
Chasteen, L., Ayriss, J., Pavlik, P. & Bradbury, A. R. Eliminating helper phage from phage display. Nucleic Acids Res. 34, e145 (2006)
Clokie, M. R. & Kropinski, A. M. Bacteriophages Ch. 8, 77–80 (Springer, 2008)
Kick, B., Hensler, S., Praetorius, F., Dietz, H. & Weuster-Botz, D. Specific growth rate and multiplicity of infection affect high-cell-density fermentation with bacteriophage M13 for ssDNA production. Biotechnol. Bioeng. 114, 777–784 (2017)
Stahl, E., Martin, T. G., Praetorius, F. & Dietz, H. Facile and scalable preparation of pure and dense DNA origami solutions. Angew. Chem. Int. Ed. 53, 12735–12740 (2014)
Sambrook, J. & Russell, D. Molecular Cloning: A Laboratory Manual 3rd edn Vol. 1, Ch. 5 (Cold Spring Harbor, 2001)
Scheres, S. H., Núñez-Ramírez, R., Sorzano, C. O., Carazo, J. M. & Marabini, R. Image processing for electron microscopy single-particle analysis using XMIPP. Nat. Protocols 3, 977–990 (2008)
Niekamp, S. et al. Folding complex DNA nanostructures from limited sets of reusable sequences. Nucleic Acids Res. 44, e102 (2016)
Acknowledgements
We thank D. Maslak (TUM Research Center for Industrial Biotechnology, Technical University of Munich) for cost estimation in pilot-scale production at the TUM Pilot Plant for Industrial Biotechnology. We thank T. Gerling, E. Meier, M. Schickinger and J. Funke for discussions. This project was supported by European Research Council starting grant 256270, the Deutsche Forschungsgemeinschaft through grants provided within TUM IGSSE (Biomat 05 PSN), the Gottfried–Wilhelm–Leibniz Program, the SFB863, and the Excellence Cluster CIPSM (Center for Integrated Protein Science Munich), the ERASynBio project ‘BioOrigami’, funded by Bundesministerium für Bildung und Forschung grant 031 A 458.
Author information
Authors and Affiliations
Contributions
F.P. performed the research. H.D. designed the research. B.K., K.L.B. and D.W.-B. developed and performed the high-cell-density fermentation and large-scale folding procedures. M.N.H. contributed to DNAzyme optimization. F.P. and H.D. wrote the manuscript and prepared the figures. All authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
A provisional patent has been filed that lists F.P. and H.D. as authors.
Additional information
Reviewer Information Nature thanks M. Famulok and B. Högberg for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 DNAzyme cassette architectures with only one DNAzyme or with one class II and one class I enzyme are not suitable for staple production.
a, Schematic representation of a pseudogene in which adjacent staples are separated by a single DNAzyme, yielding staples with a 40-base-long DNAzyme overhang at their 3′ ends. b, Schematic representation of a pseudogene in which staples are separated by one class II and one class I DNAzyme, yielding staples without 3′ overhangs. c, d, Agarose-gel-electrophoretic characterization of the self-cleavage reaction of a pseudogene using the DNAzyme cassette architecture shown in b; c, self-cleavage at different salt concentrations at 37 °C; d, self-cleavage at different temperatures at salt concentrations of 5 mM MgCl2 and 2 mM ZnSO4. sta, reference sample containing chemically synthesized staples; DZ, reference sample containing chemically synthesized oligonucleotides with the sequence of the ‘waste’ DNAzymes.
Extended Data Figure 2 The first two base pairs of the P2 stem in class I DNAzymes are crucial for efficient self-cleavage, whereas the sequence of the P1 stem does not affect cleavage efficiency.
a, b, Left, schematic representation of DNAzyme variants as they would be used at the 5′ (a) or 3′ (b) end of a staple; black letters indicate bases that were classified as highly conserved26, red letters indicate positions that we identified to be crucial for efficient cleaving activity, P1 and P2 indicate the two stems, and black triangles indicate the cleavage sites. Right, agarose-gel-electrophoretic analysis of the reaction kinetics of the respective DNAzyme variant. The slower migrating band corresponds to the uncleaved oligonucleotide and the faster migrating band to the reaction product. The activity time constant τ is defined as the first time point at which the intensity of the product band exceeds the intensity of the uncleaved DNAzyme. c, d, Comparison of the cleavage kinetics of different sequence variations of the DNAzymes shown in a and b, respectively.
Extended Data Figure 3 The DNAzyme at the 3′ end of a staple should have a P2 stem that consists of the final 8 bases of the staple, followed by AC.
Left, schematic representation of a class I DNAzyme as it would be used at the 3′ end of a staple. Right, comparison of different sequence variants of the DNAzyme. The activity time constant τ is defined as in Extended Data Fig. 2. Sequences labelled ‘v1’ are inefficiently self-cleaving DNAzyme sequences taken from the pseudogene shown in Extended Data Fig. 4b. Sequences labelled ‘v2’ have a longer double-stranded stem; sequences labelled ‘v3’, ‘v4’ or ‘v5’ contain the optimal base pairs at the first two positions of the P2 stem. Sequence variants v5 are used in the pseudogene shown in Extended Data Fig. 4a, which is also the architecture used for all of the structures shown here.
Extended Data Figure 4 DNAzyme cassettes comprising two copies of the class I DNAzyme with GT as the first two bases in each P2 stem function robustly and independently of their respective sequences in the context of a long pseudogene.
a, b, Top, schematic representations of two phagemids that contain the same set of staple sequences but are constructed using slightly different DNAzyme architectures. Bottom, analysis of the self-cleavage kinetics of the respective phagemids using alkaline-denaturing agarose gel electrophoresis. Phagemid variant 1 (a) is based on the minimal DNAzyme sequence as identified by us. Phagemid variant 2 (b) is based on the minimal DNAzyme sequence. ref, reference sample containing chemically synthesized staple strands and ‘waste’ DNAzyme snippets. c, Auto-levelled (left) and highly oversaturated (right) scans of a denaturing urea-polyacrylamide gel. L, double-stranded DNA ladder; DZ 1 and DZ 2, chemically synthesized oligonucleotides with the sequence of excised DNAzyme cassettes (84 bases long; DZ 1 and DZ 2 denote different sequences in the P1 and P2 stem regions); chem, mixture of all chemically synthesized staple oligonucleotides needed to fold the pointer (staple lengths vary between 41 and 80 bases, thus multiple bands); bio, mixture of cleaved staple pseudogenes that contain all of the staples needed to fold the pointer, purified either by anion exchange chromatography (IEX) or by ethanol precipitation (EtOH).
Extended Data Figure 5 Staple lengths of more than 200 bases lead to decreased folding quality in the DNA origami nanorod.
a, Agarose-gel-electrophoretic analysis of self-assemly reactions of nanorods designed with staple lengths of 210 (left) or 420 (right) bases. Assembly mixtures were subjected to temperature ramps at different MgCl2 concentrations; 0 mM MgCl2 indicates reference samples that were not subjected to the temperature ramp. A well-folded structure would be expected to run faster than the scaffold in the reference sample. b, c, Negative-staining TEM micrographs of nanorods assembled using 210-base-long (b) or 420-base-long (c) staples.
Extended Data Figure 6 Backbone ssDNA can be removed efficiently using anion exchange chromatography, and biotechnological staple production removes the need for a large excess of staple strands over scaffold strands.
a, Left, agarose-gel-electrophoretic analysis of pointer self-assembly reactions containing either chemically synthesized staples (chem) with (oh) or without (no oh) the additional overhangs produced by the DNAzyme cassette or biotechnologically produced (bio) staples purified using only ethanol precipitation (EtOH) or using anion exchange chromatography (IEX). Right, similar comparison, but with samples in which excess strands were removed via PEG purification before loading on the gel. All samples have overhangs. b, Agarose-gel-electrophoretic analysis of pointer self-assembly reactions using a constant scaffold concentration of 20 nM and varying staple concentrations of 15–100 nM. sc, scaffold only; sta, biotechnologically produced staples only; chem, self-assembly reactions using chemically synthesized staples; bio, self-assembly reactions using biotechnologically produced staples. For this experiment, biotechnologically produced staples were not purified using anion exchange chromatography, which leads to the appearance of the pointer + backbone band. c, Negative-staining TEM micrographs of pointer structures assembled from cleaved phagemids containing backbone ssDNA: left, unpurified sample; right, PEG-purified sample. d, Agarose-gel-electrophoretic analysis of the different fractions collected from an anion exchange chromatography purification of the cleaved pointer phagemids at different NaCl concentrations. M, marker; ref, unpurified sample; FT, flow through. The red box indicates a section of the image that was auto-levelled individually.
Extended Data Figure 7 Site-specific modification of biotechnologically produced DNA nanostructures.
a, Individual scans of the gel shown in Fig. 3b recorded in the ethidium bromide channel (left) and the Cy5 channel (right). b, c, Schematic representation (left), negative-staining TEM micrographs (centre) and agarose-gel-electrophoretic analysis (right) of knockout-mediated oligomerization (b) or passivation (c) of the screw-nut. Oligomerization of the monomeric screw-nut variant (b) was induced by replacing passivating staples with non-passivating staples. Passivation of the oligomerizing screw-not variant (c) was achieved by replacing non-passivating staples with passivating staples. Scale bar, 100 nm.
Extended Data Figure 8 Establishing a fed-batch process for phagemid ssDNA production.
a, The titre of phagemid particles containing the target ssDNA is plotted against the time of the feeding phase for different multiplicities of infection (MOIs) of helper phage M13K07 (white triangles, MOI = 0.05 pfu cfu−1; green triangles, MOI = 0.8 pfu cfu−1; blue triangles, MOI = 1.3 pfu cfu−1; pfu, plaque-forming units; cfu, colony-forming units). The infection with helper phages was conducted after 5 h of predefined exponential feeding (growth rate of 0.15 h−1) at a cell dry weight of 22.4 g l−1 (OD600 = 54). b, Comparison of product (blue trianges) and helper (red triangles) phages over time during the feeding phase for MOI = 1.3 pfu cfu−1. c, Image of an agarose gel comparing the phagemid and helper-phage ssDNA at different processing times after helper-phage addition, millilitre-scale and litre-scale purification. d, Reproduction of the fed-batch process with helper phage M13KO7 infection for phagemid ssDNA production. Blue, grey and black symbols correspond to three independent reproductions of the fed-batch process. e, Agarose-gel-electrophoretic analysis of bioreactor-derived phagemid ssDNA in comparison to shaker flask (ref) and helper-phage ssDNA. Error bars in a, b and d represent the standard deviation of three technical replicates. Gel scans in c and e are oversaturated to enable the detection of minor side products.
Extended Data Figure 9 Cost estimate for the biotechnological production of DNA origami nanorods.
a, Comparison of the net costs for the reaction-systems shake flask (6 l) and the stirred-tank reactor (STR) with laboratory-scale (1.9 l) and pilot-scale (800 l) fermentation. b, c, Relative contributions of different expense factors to the total net cost of fermentation at the laboratory (b) or pilot (c) scale. See also Supplementary Table 1 for details about the cost analysis.
Supplementary information
Supplementary Data 1
This file contains caDNAno design diagrams of nanorod variants. a Variant with ∼100 bases per staple shown in Fig. 2a. b Variant with ∼200 bases per staple shown in Extended Data Fig. 5. c Variant with ∼400 bases per staple shown in Extended Data Fig. 5. (PDF 504 kb)
Supplementary Data 2
This file contains caDNAno design diagrams of screw-nut variants. a Variant with polythymidine passivation shown in Fig. 2b and Fig. 3c. b Variant without polythymidine passivation shown in Fig. 3d. (PDF 511 kb)
Supplementary Data 3
caDNAno design diagram of the brick shown in Fig. 2d (PDF 1060 kb)
Supplementary Table 1
Calculation of cost for material and chemicals of a 1.9-liter fermentation process in a stirred-tank bioreactor. (PDF 37 kb)
Supplementary Table 2
This file contains sequences of all phagemids used in this work, of the chemically synthesized staples used for the pointer, as well as of all oligonucleotides used in the knock-out experiments. (XLS 144 kb)
Rights and permissions
About this article
Cite this article
Praetorius, F., Kick, B., Behler, K. et al. Biotechnological mass production of DNA origami. Nature 552, 84–87 (2017). https://doi.org/10.1038/nature24650
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24650
This article is cited by
-
Expedient production of site specifically nucleobase-labelled or hypermodified RNA with engineered thermophilic DNA polymerases
Nature Communications (2024)
-
Nanobiosensors: A powerful Technology for Early Detection of Plant Parasitic Nematodes
Sensing and Imaging (2024)
-
Biodistribution and function of coupled polymer-DNA origami nanostructures
Scientific Reports (2023)
-
The harmony of form and function in DNA nanotechnology
Nature Nanotechnology (2023)
-
Isothermal self-assembly of multicomponent and evolutive DNA nanostructures
Nature Nanotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.