Abstract
Activation of the PTEN–PI3K–mTORC1 pathway consolidates metabolic programs that sustain cancer cell growth and proliferation1,2. Here we show that mechanistic target of rapamycin complex 1 (mTORC1) regulates polyamine dynamics, a metabolic route that is essential for oncogenicity. By using integrative metabolomics in a mouse model3 and human biopsies4 of prostate cancer, we identify alterations in tumours affecting the production of decarboxylated S-adenosylmethionine (dcSAM) and polyamine synthesis. Mechanistically, this metabolic rewiring stems from mTORC1-dependent regulation of S-adenosylmethionine decarboxylase 1 (AMD1) stability. This novel molecular regulation is validated in mouse and human cancer specimens. AMD1 is upregulated in human prostate cancer with activated mTORC1. Conversely, samples from a clinical trial with the mTORC1 inhibitor everolimus5 exhibit a predominant decrease in AMD1 immunoreactivity that is associated with a decrease in proliferation, in line with the requirement of dcSAM production for oncogenicity. These findings provide fundamental information about the complex regulatory landscape controlled by mTORC1 to integrate and translate growth signals into an oncogenic metabolic program.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
17 January 2018
Please see accompanying Corrigendum (http://doi.org/10.1038/nature25470). In Extended Data Figs 5e and 9h, the protein ‘NSIN1’ was corrected to ‘SIN1’. In Extended Data Fig. 8e, the extracted ion chromatogram of the TVLASPQKIEGFK peptide for phosphorylated AMD1 was added to the left panel, and the following text was added to the figure legend for Extended Data Fig. 8e: “Extracted ion chromatogram (XIC) of the TVLASPQKIEGFK peptide (left);”. In addition, the following affiliation was added to author Violeta Serra: ‘CIBERONC, Instituto de Salud Carlos III, C/ Monforte de Lemos 3-5, Pabellón 11, Planta 0, 28029 Madrid, Spain’.
References
Efeyan, A., Comb, W. C. & Sabatini, D. M. Nutrient-sensing mechanisms and pathways. Nature 517, 302–310 (2015)
Ben-Sahra, I. & Manning, B. D. mTORC1 signaling and the metabolic control of cell growth. Curr. Opin. Cell Biol. 12, 72–82 (2017)
Chen, Z. et al. Crucial role of p53-dependent cellular senescence in suppression of Pten-deficient tumorigenesis. Nature 436, 725–730 (2005)
Ugalde-Olano, A. et al. Methodological aspects of the molecular and histological study of prostate cancer: focus on PTEN. Methods 77-78, 25–30 (2015)
Tabernero, J. et al. Dose- and schedule-dependent inhibition of the mammalian target of rapamycin pathway with everolimus: a phase I tumor pharmacodynamic study in patients with advanced solid tumors. J. Clin. Oncol. 26, 1603–1610 (2008)
Carracedo, A. & Pandolfi, P. P. The PTEN-PI3K pathway: of feedbacks and cross-talks. Oncogene 27, 5527–5541 (2008)
Engelman, J. A., Luo, J. & Cantley, L. C. The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat. Rev. Genet. 7, 606–619 (2006)
Song, M. S., Salmena, L. & Pandolfi, P. P. The functions and regulation of the PTEN tumour suppressor. Nat. Rev. Mol. Cell Biol. 13, 283–296 (2012)
Pegg, A. E. S-Adenosylmethionine decarboxylase. Essays Biochem. 46, 25–45 (2009)
Xiong, H. & Pegg, A. E. Mechanistic studies of the processing of human S-adenosylmethionine decarboxylase proenzyme. Isolation of an ester intermediate. J. Biol. Chem. 274, 35059–35066 (1999)
Kryukov, G. V. et al. MTAP deletion confers enhanced dependency on the PRMT5 arginine methyltransferase in cancer cells. Science 351, 1214–1218 (2016)
Marjon, K. et al. MTAP deletions in cancer create vulnerability to targeting of the MAT2A/PRMT5/RIOK1 axis. Cell Reports 15, 574–587 (2016)
Mavrakis, K. J. et al. Disordered methionine metabolism in MTAP/CDKN2A-deleted cancers leads to dependence on PRMT5. Science 351, 1208–1213 (2016)
Shlomi, T., Fan, J., Tang, B., Kruger, W. D. & Rabinowitz, J. D. Quantitation of cellular metabolic fluxes of methionine. Anal. Chem. 86, 1583–1591 (2014)
Eskens, F. A. et al.; European Organization for Research and Treatment of Cancer Early Clinical Studies Group. Phase I and pharmacological study of weekly administration of the polyamine synthesis inhibitor SAM 486A (CGP 48 664) in patients with solid tumors. Clin. Cancer Res. 6, 1736–1743 (2000)
Paridaens, R. et al. A phase I study of a new polyamine biosynthesis inhibitor, SAM486A, in cancer patients with solid tumours. Br. J. Cancer 83, 594–601 (2000)
Regenass, U. et al. CGP 48664, a new S-adenosylmethionine decarboxylase inhibitor with broad spectrum antiproliferative and antitumor activity. Cancer Res. 54, 3210–3217 (1994)
Siu, L. L. et al. A phase I and pharmacokinetic study of SAM486A, a novel polyamine biosynthesis inhibitor, administered on a daily-times-five every-three-week schedule in patients with advanced solid malignancies. Clin. Cancer Res. 8, 2157–2166 (2002)
Serra, H. et al. PTEN mediates Notch-dependent stalk cell arrest in angiogenesis. Nat. Commun. 6, 7935 (2015)
Greenberg, N. M. et al. Prostate cancer in a transgenic mouse. Proc. Natl Acad. Sci. USA 92, 3439–3443 (1995)
Origanti, S. et al. Ornithine decarboxylase mRNA is stabilized in an mTORC1-dependent manner in Ras-transformed cells. Biochem. J. 442, 199–207 (2012)
Bale, S. & Ealick, S. E. Structural biology of S-adenosylmethionine decarboxylase. Amino Acids 38, 451–460 (2010)
Zhang, Y. et al. Coordinated regulation of protein synthesis and degradation by mTORC1. Nature 513, 440–443 (2014)
Ouyang, D. Y. et al. Autophagy is differentially induced in prostate cancer LNCaP, DU145 and PC-3 cells via distinct splicing profiles of ATG5. Autophagy 9, 20–32 (2013)
Hsu, P. P. et al. The mTOR-regulated phosphoproteome reveals a mechanism of mTORC1-mediated inhibition of growth factor signaling. Science 332, 1317–1322 (2011)
Zhao, J., Zhai, B., Gygi, S. P. & Goldberg, A. L. mTOR inhibition activates overall protein degradation by the ubiquitin proteasome system as well as by autophagy. Proc. Natl Acad. Sci. USA 112, 15790–15797 (2015)
Huang, Z. et al. Clinical efficacy of mTOR inhibitors in solid tumors: a systematic review. Future Oncol. 11, 1687–1699 (2015)
O’Reilly, K. E. et al. mTOR inhibition induces upstream receptor tyrosine kinase signaling and activates Akt. Cancer Res. 66, 1500–1508 (2006)
Carracedo, A. et al. Inhibition of mTORC1 leads to MAPK pathway activation through a PI3K-dependent feedback loop in human cancer. J. Clin. Invest. 118, 3065–3074 (2008)
Gerner, E. W. & Meyskens, F. L., Jr. Polyamines and cancer: old molecules, new understanding. Nat. Rev. Cancer 4, 781–792 (2004)
Song, M. S. et al. Nuclear PTEN regulates the APC-CDH1 tumor-suppressive complex in a phosphatase-independent manner. Cell 144, 187–199 (2011)
Chen, Z. et al. Differential p53-independent outcomes of p19(Arf) loss in oncogenesis. Sci. Signal. 2, ra44 (2009)
Motameni, A. R. et al. Delivery of the immunosuppressive antigen Salp15 to antigen-presenting cells by Salmonella enterica serovar Typhimurium aroA mutants. Infect. Immun. 72, 3638–3642 (2004)
Torrano, V. et al. The metabolic co-regulator PGC1α suppresses prostate cancer metastasis. Nat. Cell Biol. 18, 645–656 (2016)
Dowling, R. J. et al. mTORC1-mediated cell proliferation, but not cell growth, controlled by the 4E-BPs. Science 328, 1172–1176 (2010)
Wiederschain, D. et al. Single-vector inducible lentiviral RNAi system for oncology target validation. Cell Cycle 8, 498–504 (2009)
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014)
Carracedo, A. et al. A metabolic prosurvival role for PML in breast cancer. J. Clin. Investigation 122, 3088–3100 (2012)
Salazar, M. et al. Loss of Tribbles pseudokinase-3 promotes Akt-driven tumorigenesis via FOXO inactivation. Cell Death Differ. 22, 131–144 (2015)
Alimonti, A. et al. A novel type of cellular senescence that can be enhanced in mouse models and human tumor xenografts to suppress prostate tumorigenesis. J. Clin. Invest. 120, 681–693 (2010)
Revandkar, A. et al. Inhibition of Notch pathway arrests PTEN-deficient advanced prostate cancer by triggering p27-driven cellular senescence. Nat. Commun. 7, 13719 (2016)
Fuhrer, T., Heer, D., Begemann, B. & Zamboni, N. High-throughput, accurate mass metabolome profiling of cellular extracts by flow injection–time-of-flight mass spectrometry. Anal. Chem. 83, 7074–7080 (2011)
Jha, A. K. et al. Network integration of parallel metabolic and transcriptional data reveals metabolic modules that regulate macrophage polarization. Immunity 42, 419–430 (2015)
Melamud, E., Vastag, L. & Rabinowitz, J. D. Metabolomic analysis and visualization engine for LC-MS data. Anal. Chem. 82, 9818–9826 (2010)
Fumagalli, S., Ivanenkov, V. V., Teng, T. & Thomas, G. Suprainduction of p53 by disruption of 40S and 60S ribosome biogenesis leads to the activation of a novel G2/M checkpoint. Genes Dev. 26, 1028–1040 (2012)
Grasso, C. S. et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature 487, 239–243 (2012)
Lapointe, J. et al. Gene expression profiling identifies clinically relevant subtypes of prostate cancer. Proc. Natl Acad. Sci. USA 101, 811–816 (2004)
Taylor, B. S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010)
Tomlins, S. A. et al. Integrative molecular concept modeling of prostate cancer progression. Nat. Genet. 39, 41–51 (2007)
Acknowledgements
We thank N. Sonenberg for providing 4EBP1- and 4EBP2-targeting shRNAs, D. Alessi for discussions and technical advice, A.M. Cuervo and E. Arias-Perez for technical advice, the Basque biobank for research (BIOEF) for critical support with human specimens and Novartis for providing SAM486A. Funding: Ramón y Cajal award (to A.C., A.E., J.M., D.O.), Juan de la Cierva (to E.C., A.C.-M.), BFU grant (to R.B.: BFU2014-52282-P and BFU2011-25986), SAF grant (to A.C.: SAF2016-79381-R, FEDER/EU; M.L.M.-C.: SAF2014-54658-R; to J.M.F.-P.: SAF2015-66312; to J.M.M: SAF 2014-52097R; to A.E.: SAF2015-67538-R; to J.A.: SAF2015-65327R; to G.T.: SAF2011-24967) from the Spanish Ministry of Economy, Industry and Competitiveness (MINECO); European Union (to A.C.: ERC-StG-336343, PoC754627; to A.C.-M.: CIG 660191; to J.A.: 602272; to A.E.: ERC-2014-STG-638891); Basque Government Department of Health (to V.T.: 2016111109; to J.M.F.-P: 2015111149), Department of Education (to A.C.: PI2012/03 and IKERTALDE I.T.1106-16; to R.B.: PI2012/42; to M.L.M.-C.: 2013) and PhD grants (to A.A.-A. and L.V.-J.); AECC (to V.T.: 2016 JP Bizkaia; to N.M.-M.: 2011 JP Bizkaia; to M.L.M.-C.); ISCIII (to A.C.: PI10/01484, PI13/00031; to J.M.: Proteored PR.B.2 and grant PT13/0001; to R.F.: PI15/209; to V.S.: PI13/01714, CP14/00228); Ramón Areces foundation (to J.M.F.-P.); Basque Department of Industry, Tourism and Trade (Etortek) (to A.C.); FERO Foundation (to A.C., V.S.); Fundación Vasca de Innovación e Investigación Sanitarias, BIOEF (to V.T.: BIO15/CA/052); BBVA Foundation (to A.C.; P.N. team); National Institutes of Health (to C.C.-C. and M.C.: P01CA087497; to J.M.M.: R01AT001576); Fundación CRIS contra el Cáncer (to D.O. team); 2014 Stewart Rahr Young Investigator Award from the Prostate Cancer Foundation (to D.O.); FPU predoctoral fellowship (to Y.C.: 15/05126); Catalan Agency AGAUR (to V.S.: 2014 SGR 1331); Medical Research Council (to R. Bago, D. Alessi laboratory: grant number MC_UU_12016/2). The activity of CIBERONC was co-funded with FEDER funds.
Author information
Authors and Affiliations
Contributions
In vitro studies: A.Z.-L. and A.A.-A. with support from N.M.-M., S.F.-R., and L.A. In vivo studies: A.Z.-L., A.A.-A. and N.M.-M. with support for tail vein from N.B. Genotyping: P.S.-M. Metabolomics analysis: M.Clas., P.Q., S.M., A.B., G.L., and D.P., K. Marks and K. Marjon at AGIOS and D.C., S.M.V.L., and J.M.F.-P. at CIC bioGUNE. Everolimus trial samples and staining: V.S., J.J., L.P., P.N., I.T., J.T., T.M., and J.B. Human prostate specimens: A.U.-O., A.L.-I., I.L.-V., and M.U. Histochemical analysis in mice: S.F.-R., A. Rev., and A. Alim. Generation of molecular AMD1 tools: J.D.S. and R.B. Immune cell analysis and GST–AMD1 purification: J.T.-C., I.F.-D., and J.A. Phosphoproteomics: P.X.-E., A.E., and J.M. Kinase assay: R. Bago. TRAMP mice material: statistical supervision and bioinformatics, A.R.-C.; polysome profiling, A.G. and G.T. Genetically engineered mouse model pathology: M.C.-M. and C.C.-C. Technical support and discussions: B.D.M., J.M.M., M.L.M.-C., M.A., F.E., I.A., A.M.A., V.T., L.V.-J., P.Z.-G., M.P., A.C.-M., R.F., and Y.Z. K. Marjon directed the metabolomics strategy, analysed the results, and contributed to discussions and manuscript preparation and revision. A.C. directed the project, supervised data analysis, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Metabolomics characterization of mouse and human prostate cancer.
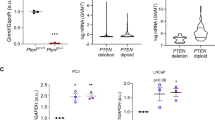
a, Incidence of pathological alterations observed in Ptenpc+/+ and Ptenpc−/− mice. Number of mice as indicated. LG-PIN, low-grade prostatic intraepithelial neoplasia; HG-PIN: high-grade prostatic intraepithelial neoplasia; Focal Adc, focal adenocarcinoma. b, Representative immunohistochemical images of prostate tissue stained with haematoxylin and eosin (H&E), Pten, AktS473, and RpS6S235/6 from mice at 3 and 6 months of age (representative of three mice per condition). c, Experimental design of the TOF-MS metabolomics analysis. AP, anterior prostate; DLP, dorsolateral prostate. d, e, Volcano plot (d) and principal component analysis (PCA, e) from altered metabolites in TOF-MS metabolomic analysis performed in Ptenpc−/− and Ptenpc+/+ mouse prostate samples at the indicated age (6 months Ptenpc+/+anterior prostate, n = 4 mice; remainder of conditions, n = 5 mice). Grey dots: not significantly altered; red dots: significantly increased in Ptenpc−/− prostate extracts; blue dots: significantly decreased in Ptenpc−/− prostate extracts. f, LC/MS analysis of methionine cycle and polyamine pathway metabolites from Ptenpc−/− versus Ptenpc+/+ mouse prostate samples at the indicated age (anterior prostate 3 months, n = 5 mice; 6 months, n = 4 mice). Median ± interquartile range. g, LC/MS analysis of methionine cycle and polyamine pathway metabolites from prostate cancer versus BPH human specimens (six prostate specimens per condition). Median ± interquartile range. *P < 0.05; **P < 0.01; ***P < 0.001. One-tailed Mann–Whitney U-test (f, g) was used for data analysis.
Extended Data Figure 2 Metabolic tracing of [13C]l-methionine in Pten-prostate specific knockout mice.
a, Plasma LC/MS analysis of the indicated metabolite concentration after intravenous injection of [U-13C5]l-methionine (100 mg kg−1) in C57BL/6 mice at 3 months of age (time 0 min, n = 7 mice; time 10 min/60 min, n = 6 mice). The unlabelled (M + 0, 12C) and major labelled (13C, M + 4 or M + 5) metabolite concentration is presented in the histogram. Error bars, s.e.m. b, Experimental design of the [U-13C5]l-methionine (100 mg kg−1) in vivo. U-13C5-Met, l-methionine labelled with13C in five carbons;1 h, prostate samples extracted after 1 h pulse with [U-13C5]l-methionine; 10 h, prostate samples extracted after 10 h pulse with [U-13C5]l-methionine; c, Summary schematic of the alterations observed in the metabolomic analysis in Ptenpc−/− versus Ptenpc+/+. Spm, spermine; spd, spermidine; ODC1, ornithine decarboxylase 1; SpdS, spermidine synthase; SpmS, spermine synthase. d, Fractional labelling of the indicated metabolites from Fig. 1c. Median ± interquartile range (1 h (top), n = 4; 10 h (bottom), n = 3). FC, fold change. One-tailed Mann–Whitney U-test (d) was used for data analysis.
Extended Data Figure 3 Genetic modulation of AMD1 in prostate cancer cells.
a, Technical setup of anti-AMD1 antibody using the indicated constructs or shRNAs in DU145 cells. AMD1S229A mutant lacks processing ability and is expressed exclusively as a pro-enzyme (representative western blot out of three independent experiments). b, Schematic representation of tagged ectopic AMD1 processing. c, Impact of ectopic Myc-AMD1–HA expression on foci number in DU145 cells in vitro (n = 3 independent experiments). d–f, Impact of ectopic Myc-AMD1–HA expression on tumour volume (d, n = 8 tumours per condition), AMD1 protein levels (e, n = 3 tumours per condition), and dcSAM abundance (f, n = 5 tumours per condition) in DU145 xenografts grown for 43 days. Data in d are represented as box-and-whisker plot. f, Dot plot with the median and the interquartile range. g–o, Effect of constitutive silencing of AMD1 (g, i, mRNA levels; h–j, protein expression) on cell number (k, n), anchorage-independent growth (l, o), and dcSAM abundance (m) with two different hairpins in DU145 (g, h, k, l, m) and PC3 (i, j, n, o) cells (n = 3 or 4 independent experiments as indicated by dots). shSC, scramble short hairpin; sh3 and sh4, two different short-hairpins targeting AMD1. Mean ± s.e.m. p, Effect of doxycycline-inducible (100 ng ml−1) AMD1 silencing on cell number in DU145 cells (sh3, n = 4; sh4 and sh5, n = 3 independent experiments as indicated by dots). q–s, Impact of AMD1-inducible silencing in tumour volume from DU145 xenografts (tumours analysed: sh3 no doxycycline, n = 12; sh3 doxycycline, n = 14; sh4 no doxycycline, n = 10; sh4 doxycycline, n = 7; sh5 no doxycycline, n = 10; sh5 doxycycline, n = 11). Box-and-whisker plot. t, Impact of AMD1-inducible silencing in dcSAM abundance in DU145 xenografts from q to s (n = 5 tumours). Median ± interquartile range. u, v, Dose-dependent effect of doxycycline on cell number in DU145 (u) and PC3 (v) cells (cell number measured at day 6) (n = 3 independent experiments as indicated by dots). *P < 0.05; **P < 0.01; ***P < 0.001. Error bars, mean ± s.e.m. Red asterisk in western blots indicates non-specific band. Dashed lines indicate cell numbers in scramble short-hairpin-transduced cells. One-tailed t-test (c, g, i, k–p, u, v), and one-tailed Mann–Whitney U-test (d, f, q–t).
Extended Data Figure 4 Genetic and pharmacological manipulation of AMD1 in prostate cancer cells.
a–c, DU145 cells carrying doxycycline-inducible shRNA against AMD1 (sh3) were transduced with empty (Mock), sh3-resistant wild type (AMD1sh3R), or processing-deficient (AMD1sh3R/S229A) AMD1 constructs. AMD1 protein (a, representative experiment out of four), dcSAM abundance (b), and cell number expression (c) in the aforementioned cells (n = 4 independent experiments as indicated by dots). Asterisks indicate significant differences compared with the corresponding DU145 cells in the absence of doxycycline, and hash symbol indicates significant differences in the indicated comparison. Mean ± s.e.m. d, Effect of MTA (25 μM) on AMD1 silencing (sh3)-elicited anti-proliferative activity. MTA was administered at day 0 and cells were analysed at day 3 (n = 3 independent experiments as indicated by dots). e, Mtap gene expression levels in Ptenpc+/+ and Ptenpc−/− mice at the indicated time points (see Extended Data Fig. 1a) (3 months, n = 3 mice; 6 months, n = 6 mice). Median ± interquartile range. f, MTAP gene expression analysis in publicly available data sets (see Methods; N, normal; number of patients analysed: ref. 48, normal n = 29, prostate cancer n = 150; ref. 46, normal n = 12, prostate cancer n = 76; ref. 47, normal, n = 9, prostate cancer, n = 17). g, h, Effect of pharmacological AMD1 inhibition with SAM486A on cell number (g and left h; DU145, n = 5; PC3, n = 4 independent experiments as indicated by dots), and anchorage-independent growth (right h, n = 3 independent experiments as indicated by dots) in PC3 or DU145 cells as indicated. Mean ± s.e.m. i, Effect of pharmacological AMD1 inhibition with SAM486A in established DU145 xenograft tumour volume (vehicle, n = 11 tumours; SAM486A, n = 10 tumours). Box-and-whisker plot. j, k, Effect of pharmacological AMD1 inhibition with SAM486A in activated T CD4 cell number (96 h (j), n = 3 independent experiments as indicated by dots) or IL-2 production (k, n = 3–6 independent experiments as indicated by dots). */#P < 0.05; **/##P < 0.01; ***P < 0.001. Dashed line indicates cell number (g, h left) or IL-2 abundance (j, k) in vehicle-treated cells. Student’s t-test (b–d, f–h, j, k) and one-tailed Mann–Whitney test (e, i).
Extended Data Figure 5 PTEN–PI3K–mTORC1-dependent regulation of AMD1 in prostate cancer.
a, b, Amd1 protein quantification from Fig. 3a (a, n = 3 mice) and mRNA expression (b, n = 5 or 6 mice as indicated by dots) in Ptenpc−/− and Ptenpc+/+ mice of the indicated age. Median ± interquartile range. c, AMD1 gene expression analysis in publicly available data sets (see Methods; number of patients analysed: ref. 48, normal n = 29, prostate cancer n = 150; ref. 46, normal n = 12, prostate cancer n = 76; ref. 47, normal n = 9, prostate cancer n = 17). d, Representative western blot showing the expression of the indicated proteins upon doxycycline-inducible expression (24 h) of YFP–PTENWT or catalytically inactive YFP–PTENC124S in PTEN-deficient LNCaP prostate cancer cells (n = 3 independent experiments). e, Schematic representation of the PI3K signalling pathway and the pharmacological/genetic tools used in this study. f, ProAMD1 and AMD1 protein quantification from Fig. 3b (sample number as indicated by dots). g, AMD1 gene expression upon treatment (24 h) with vehicle (V, DMSO), rapamycin (R, 20nM) and Torin-1 (T, 250 nM for PC3 and DU145, 125 nM for LNCaP) (LNCaP, n = 8 for Torin-1 and n = 6 for rapamycin; PC3 and DU145, n = 6 independent experiments as indicated by dots). Mean ± s.e.m. h, Representative western blot analysis of AMD1 levels upon 24-h treatment of DU145 cells with vehicle, Torin-1 (mTORC1/2 inhibitor; 250 nM), PD032901 (ERK-MAPK inhibitor, PD; 100 nM), SP600125 (JNK-SAPK inhibitor, SP; 10 μM), and SB203580 (p38-MAPK inhibitor, SB; 5 μM) (n = 3 independent experiments). i, Impact of inducible RAPTOR silencing in DU145 cells on proAMD1 protein levels (doxycycline-induced, 250 ng ml−1) (representative experiment out of n = 6). j, Impact of inducible TSC2 silencing in DU145 cells on proAMD1 protein levels (doxycycline-induced, 250 ng ml−1) (representative experiment out of n = 6). *P < 0.05; **P < 0.01; ***P < 0.001. Red asterisk in western blots indicates non-specific band. Arrows indicate specific immunoreactive bands. Student’s t-test (c, f, g) and Mann–Whitney test (a, b).
Extended Data Figure 6 mTORC1-dependent AMD1 regulation in vivo.
a, Effect of spermidine (Spd, 0.75 μM) on PC3 cell number upon rapamycin treatment (1 nM, outcome measured 72 h after treatment). Drugs were administered at day 0 and only spermidine was additionally administered at day 1 (n = 4 independent experiments as indicated by dots). b, Effect of RAD001 treatment on prostate pathological features and mTORC1 activity (haematoxylin and eosin and RpS6S235/6 staining by immunohistochemistry) (n = 3 mice). c, ProAmd1 and Amd1 protein abundance quantification from Fig. 3e (left; number of mice as indicated by dots). d, Representative immunohistochemical images of prostate tissue from wild-type or TRAMP mice (+/T, 28–32 weeks old) stained with haematoxylin and eosin (top) and RpS6S235/6 (bottom, Ptenpc+/+ and Ptenpc−/− prostate tissues are presented as a comparison of the RpS6 phosphorylation levels) (n = 3 mice). e, Evaluation of AMD1 expression by western blot in prostate tissues from wild-type or TRAMP mice (+/T, 28–32 weeks old) (n = 4 mice). *P < 0.05; **P < 0.01. Error bars, mean ± s.e.m. Arrows indicate specific immunoreactive bands. Student’s t-test (a) and Mann–Whitney test (c, e).
Extended Data Figure 7 Contribution of mTORC1 effector pathways and targets on the regulation of AMD1.
a, ODC1 gene expression upon treatment (24 h) of vehicle (DMSO), rapamycin (20 nM) and Torin-1 (250 nM for PC3 and DU145, 125 nM for LNCaP) in PC3, LNCaP, and DU145 cells (n = 5 independent experiments as indicated by dots). Mean ± s.e.m. b, Putrescine abundance upon treatment (24 h) of vehicle (DMSO), rapamycin (20 nM), and Torin-1 (250 nM) in DU145 and PC3 cells (n = 3 independent experiments as indicated by dots). Mean ± s.e.m. c, Odc1 gene expression in 3- and 6-month-old Ptenpc+/+ and Ptenpc−/− mice (n = 3–6 as indicated by dots). Median ± interquartile range. d, Putrescine abundance in 12-week-old Ptenpc−/− mice upon treatment with vehicle or RAD001 (10 mg per kg (body weight), 6 days per week) for 4 weeks (n = 5 mice). Mean ± s.e.m. e, Representative western blot (n = 3 independent experiments) depicting the changes in expression of the indicated proteins upon 24 h treatment of DU145 cells with rapamycin (20 nM) and/or DFMO (an inhibitor of ODC1, 50 μM) with the corresponding vehicles. f, Representative western blot showing LC3 lipidation in HCQ-treated (6 h) DU145 and PC3 cells, as a readout of macro-autophagy (n = 3 independent experiments). Arrow indicates LC3-II. g, For the analysis of translation initiation, polysome profiling analysis of AMD1 and L11 as positive control in DU145 cells treated with vehicle or rapamycin (20 nM, 8 h) is shown. Error bars, s.d. from technical replicates. h, Effect of pharmacological p70S6K inhibition with PF4708671 (10 μM) on AMD1 protein expression in DU145 cells (representative experiment out of five). i, Effect of 4EBP1/2 silencing on proAMD1 and AMD1 protein expression (upper panels) (representative experiment out of three). Lower panels show 4EBP1 (n = 5 independent experiments) and 4EBP2 (n = 4 independent experiments) gene expression in shRNA-transduced DU145 cells. *P < 0.05; **P < 0.01; ***P < 0.001. Red asterisks in western blots indicate non-specific band. Arrows indicate specific immunoreactive bands. Student’s t-test (a, b, d, i) and Mann–Whitney U-test (c).
Extended Data Figure 8 Regulation of proAMD1 stability by mTORC1.
a, ProAmd1 (left) and Amd1 (right) protein abundance quantification from Fig. 4a (n = 3 independent experiments as indicated by dots). Error bars, mean ± s.e.m. b, Representative western blot of DU145 cells expressing Ser-229-Ala (S229A) mutant Myc-AMD1–HA treated with vehicle or Torin-1 (250 nM, 6 h) in the presence or absence of MG132 (5 μM, 6 h) (n = 3 independent experiments). Quantification is provided in the right panel. c, Representative MS/MS spectrum of the TVLASPQKIEGFK peptide in proAMD1 and AMD1, in which phosphorylation was unambiguously assigned to the S298 residue. d, Calculated areas under the curves from extracted ion chromatogram in Fig. 4b of the TVLASPQKIEGFK peptide for phosphorylated (highlighted in red in the sequence, left) and total (right) proAMD1. e, Calculated areas under the curves from extracted ion chromatogram of the TVLASPQKIEGFK peptide for phosphorylated (highlighted in brown in the sequence, left) and total (right) AMD1. f, Schematic representation of the working hypothesis of proAMD1 regulation by mTORC1-dependent phosphorylation. g, Effect of Torin (250 nM) on proAMD1 and AMD1 protein at different time points in Myc-AMD1–HA-expressing DU145 cells (representative western blot out of three independent experiments; right, densitometric quantification). Error bars, s.e.m. h, i, Representative western blot depicting the stability of ectopic proAMD1 and AMD1 in DU145 cells challenged with vehicle or Torin-1 (250 nM, 2 h) upon CHX treatment (densitometry of proAMD1 and AMD1 levels is represented in right panels (h) and half-life reduction in proAMD1 (i) (n = 4 independent experiments). j, k, Representative western blot depicting the stability of S229A mutant Myc-AMD1-HA construct in DU145 cells challenged with vehicle or Torin-1 (250 nM, 2 h) upon CHX treatment (densitometry of proAMD1 is represented in the right panel (j) and half-life reduction in proAMD1 (k) (n = 3 independent experiments). Mean ± s.e.m. NS, not significant. *P < 0.05; **P < 0.01; ***P < 0.001. Student’s t-test.
Extended Data Figure 9 Contribution of proAMD1 phosphorylation downstream mTORC1 to the stability of the enzyme.
a, b, Representative western blot depicting the stability of wild type and S298A phospho-mutant Myc-AMD1-HA constructs in DU145 cells upon cycloheximide treatment (densitometry of proAMD1 and AMD1 is represented in right panels (a) and half-life reduction in proAMD1 (b) (n = 3 independent experiments). Mean ± s.e.m. c, Representative western blot depicting the stability of S298A phospho-mutant Myc-AMD1-HA construct in DU145 cells challenged with vehicle (V) or MG132 (5 μM) upon CHX treatment (densitometry of proAMD1 is represented in right panel, n = 3 independent experiments). Mean ± s.e.m. d, mTORC1 kinase activity (by means of RAPTOR immunoprecipitation from HEK293 cells) on GST–proAMD1S229A or GST–proAMD1S229A/S298A, using bacteria-purified proteins. GST–p70S6K is presented as positive control. AZD8055 is used as control of mTORC1 inhibition. e, Correlation analysis between p70S6KpT389 and AMD1 densitometry values in prostate cancer specimens from Fig. 4c (n = 15 patient specimens). f, Quantification of AMD1 immunoreactivity in response to everolimus in tumour biopsies, on the basis of the ΔH score (n = 14 specimen pairs). g, Box-and-whisker plot of the immunoreactivity of mTOR downstream effectors (AKTS473, RpS6pS240/244, 4EBP1/2pT70, eIF4GpS1108) in cancer patients with (ΔH score for KI67 < 0; n = 6 specimen pairs) or without (ΔH score for KI67 ≥ 0; n = 4 specimen pairs) anti-proliferative tumour response upon treatment with everolimus. h, Schematic representation of the main findings of this study. Orn, ornithine; Put, putrescine; Spm, spermine; ODC1, ornithine decarboxylase 1; PIP2, phosphatidyl inositol bisphosphate; PIP3, phosphatidyl inositol triphosphate; *P < 0.05; **P < 0.01; NS, not significant. Arrows indicate specific immunoreactive/autoradiographic bands. One-tailed Student’s t-test (a–c), two-tailed Mann–Whitney test (g), and Spearman’s correlation analysis (e).
Supplementary information
Supplementary Information
This file contains the uncropped blots. (PDF 4438 kb)
Supplementary Table 1
Complete list of detected ions in the Time-of-flight analysis. (XLSX 2095 kb)
Supplementary Table 2
Candidate pathways extracted from TOF-MS analysis. (XLSX 14 kb)
Supplementary Table 3
Pathway enrichment analysis of the 73 ions consistently altered in the mouse model analysis. (XLSX 9 kb)
Supplementary Table 4
Clinicopathological characteristics of patients with prostate pathology. pN, Lymph node positivity; Pn, Perineural invasion. (XLSX 11 kb)
Supplementary Table 5
Relevant mass isotopomer values in the 13C-U5-Methionine administration analysis by LC/MS. (XLSX 15 kb)
Supplementary Table 6
Effect of SAM486A administration (5mg/Kg; 5 days/week) in immunocompetent C57BL/6 mice. (XLSX 15 kb)
Supplementary Table 7
LC/MS analysis in prostate tissue extracts from WT and TRAMP (TRAMP+/T) mice. (XLSX 17 kb)
Supplementary Table 8
Clinic-pathological characteristics of patients enrolled in the Everolimus clinical trial. (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Zabala-Letona, A., Arruabarrena-Aristorena, A., Martín-Martín, N. et al. mTORC1-dependent AMD1 regulation sustains polyamine metabolism in prostate cancer. Nature 547, 109–113 (2017). https://doi.org/10.1038/nature22964
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22964
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.